Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Lysine-specific demethylase 6A
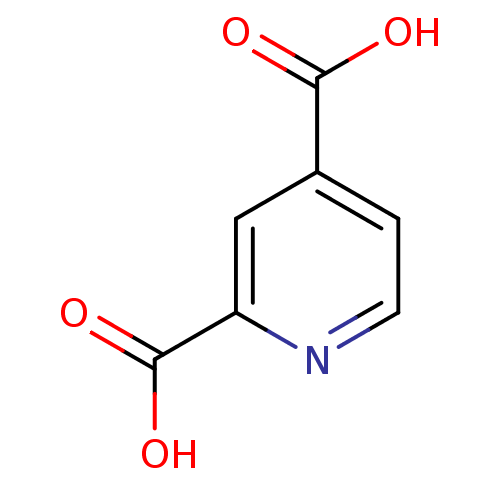
Ligand
BDBM26113
Substrate
n/a
Meas. Tech.
ChEMBL_835850 (CHEMBL2071961)
IC50
177000±n/a nM
Citation
 Leurs, U; Clausen, RP; Kristensen, JL; Lohse, B Inhibitor scaffold for the histone lysine demethylase KDM4C (JMJD2C). Bioorg Med Chem Lett 22:5811-3 (2012) [PubMed] Article
Leurs, U; Clausen, RP; Kristensen, JL; Lohse, B Inhibitor scaffold for the histone lysine demethylase KDM4C (JMJD2C). Bioorg Med Chem Lett 22:5811-3 (2012) [PubMed] Article More Info.:
Target
Name:
Lysine-specific demethylase 6A
Synonyms:
Histone demethylase UTX | KDM6A | KDM6A_HUMAN | Lysine-specific demethylase 6A | Lysine-specific demethylase 6A (KDM6A(UTX)) | Lysine-specific demethylase 6A (KDM6A) | UTX | Ubiquitously-transcribed TPR protein on the X chromosome | Ubiquitously-transcribed X chromosome tetratricopeptide repeat protein
Type:
Protein
Mol. Mass.:
154198.48
Organism:
Homo sapiens (Human)
Description:
O15550
Residue:
1401
Sequence:
MKSCGVSLATAAAAAAAFGDEEKKMAAGKASGESEEASPSLTAEEREALGGLDSRLFGFVRFHEDGARTKALLGKAVRCYESLILKAEGKVESDFFCQLGHFNLLLEDYPKALSAYQRYYSLQSDYWKNAAFLYGLGLVYFHYNAFQWAIKAFQEVLYVDPSFCRAKEIHLRLGLMFKVNTDYESSLKHFQLALVDCNPCTLSNAEIQFHIAHLYETQRKYHSAKEAYEQLLQTENLSAQVKATVLQQLGWMHHTVDLLGDKATKESYAIQYLQKSLEADPNSGQSWYFLGRCYSSIGKVQDAFISYRQSIDKSEASADTWCSIGVLYQQQNQPMDALQAYICAVQLDHGHAAAWMDLGTLYESCNQPQDAIKCYLNATRSKSCSNTSALAARIKYLQAQLCNLPQGSLQNKTKLLPSIEEAWSLPIPAELTSRQGAMNTAQQNTSDNWSGGHAVSHPPVQQQAHSWCLTPQKLQHLEQLRANRNNLNPAQKLMLEQLESQFVLMQQHQMRPTGVAQVRSTGIPNGPTADSSLPTNSVSGQQPQLALTRVPSVSQPGVRPACPGQPLANGPFSAGHVPCSTSRTLGSTDTILIGNNHITGSGSNGNVPYLQRNALTLPHNRTNLTSSAEEPWKNQLSNSTQGLHKGQSSHSAGPNGERPLSSTGPSQHLQAAGSGIQNQNGHPTLPSNSVTQGAALNHLSSHTATSGGQQGITLTKESKPSGNILTVPETSRHTGETPNSTASVEGLPNHVHQMTADAVCSPSHGDSKSPGLLSSDNPQLSALLMGKANNNVGTGTCDKVNNIHPAVHTKTDNSVASSPSSAISTATPSPKSTEQTTTNSVTSLNSPHSGLHTINGEGMEESQSPMKTDLLLVNHKPSPQIIPSMSVSIYPSSAEVLKACRNLGKNGLSNSSILLDKCPPPRPPSSPYPPLPKDKLNPPTPSIYLENKRDAFFPPLHQFCTNPNNPVTVIRGLAGALKLDLGLFSTKTLVEANNEHMVEVRTQLLQPADENWDPTGTKKIWHCESNRSHTTIAKYAQYQASSFQESLREENEKRSHHKDHSDSESTSSDNSGRRRKGPFKTIKFGTNIDLSDDKKWKLQLHELTKLPAFVRVVSAGNLLSHVGHTILGMNTVQLYMKVPGSRTPGHQENNNFCSVNINIGPGDCEWFVVPEGYWGVLNDFCEKNNLNFLMGSWWPNLEDLYEANVPVYRFIQRPGDLVWINAGTVHWVQAIGWCNNIAWNVGPLTACQYKLAVERYEWNKLQSVKSIVPMVHLSWNMARNIKVSDPKLFEMIKYCLLRTLKQCQTLREALIAAGKEIIWHGRTKEEPAHYCSICEVEVFDLLFVTNESNSRKTYIVHCQDCARKTSGNLENFVVLEQYKMEDLMQVYDQFTLAPPLPSASS