Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Misshapen-like kinase 1
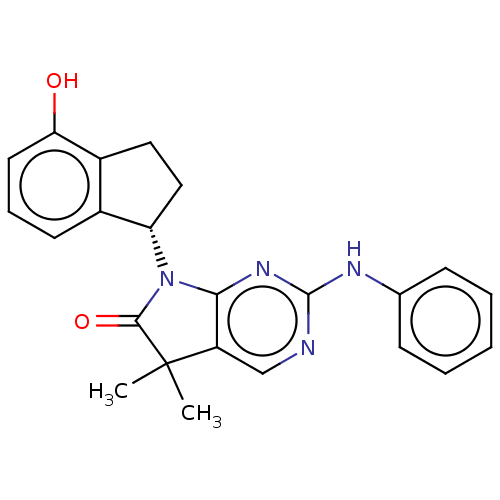
Ligand
BDBM50208911
Substrate
n/a
Meas. Tech.
ChEMBL_1633277 (CHEMBL3876069)
IC50
10000±n/a nM
Citation
 Katz, JD; Haidle, A; Childers, KK; Zabierek, AA; Jewell, JP; Hou, Y; Altman, MD; Szewczak, A; Chen, D; Harsch, A; Hayashi, M; Warren, L; Hutton, M; Nuthall, H; Su, HP; Munshi, S; Stanton, MG; Davies, IW; Munoz, B; Northrup, A Structure guided design of a series of selective pyrrolopyrimidinone MARK inhibitors. Bioorg Med Chem Lett 27:114-120 (2017) [PubMed] Article
Katz, JD; Haidle, A; Childers, KK; Zabierek, AA; Jewell, JP; Hou, Y; Altman, MD; Szewczak, A; Chen, D; Harsch, A; Hayashi, M; Warren, L; Hutton, M; Nuthall, H; Su, HP; Munshi, S; Stanton, MG; Davies, IW; Munoz, B; Northrup, A Structure guided design of a series of selective pyrrolopyrimidinone MARK inhibitors. Bioorg Med Chem Lett 27:114-120 (2017) [PubMed] Article More Info.:
Target
Name:
Misshapen-like kinase 1
Synonyms:
B55 | GCK family kinase MiNK | MAP4K6 | MAPK/ERK kinase kinase kinase 6 | MEK kinase kinase 6 | MEKKK 6 | MINK | MINK1 | MINK1_HUMAN | Misshapen-like kinase 1 (MINK) | Misshapen/NIK-related kinase | Mitogen-activated protein kinase kinase kinase kinase 6 | Voltage-gated potassium channel subunit Kv7.1/Misshapen-like kinase 1 | YSK2 | ZC3
Type:
PROTEIN
Mol. Mass.:
149838.92
Organism:
Homo sapiens (Human)
Description:
ChEMBL_793919
Residue:
1332
Sequence:
MGDPAPARSLDDIDLSALRDPAGIFELVEVVGNGTYGQVYKGRHVKTGQLAAIKVMDVTEDEEEEIKQEINMLKKYSHHRNIATYYGAFIKKSPPGNDDQLWLVMEFCGAGSVTDLVKNTKGNALKEDCIAYICREILRGLAHLHAHKVIHRDIKGQNVLLTENAEVKLVDFGVSAQLDRTVGRRNTFIGTPYWMAPEVIACDENPDATYDYRSDIWSLGITAIEMAEGAPPLCDMHPMRALFLIPRNPPPRLKSKKWSKKFIDFIDTCLIKTYLSRPPTEQLLKFPFIRDQPTERQVRIQLKDHIDRSRKKRGEKEETEYEYSGSEEEDDSHGEEGEPSSIMNVPGESTLRREFLRLQQENKSNSEALKQQQQLQQQQQRDPEAHIKHLLHQRQRRIEEQKEERRRVEEQQRREREQRKLQEKEQQRRLEDMQALRREEERRQAEREQEYKRKQLEEQRQSERLQRQLQQEHAYLKSLQQQQQQQQLQKQQQQQLLPGDRKPLYHYGRGMNPADKPAWAREVEERTRMNKQQNSPLAKSKPGSTGPEPPIPQASPGPPGPLSQTPPMQRPVEPQEGPHKSLVAHRVPLKPYAAPVPRSQSLQDQPTRNLAAFPASHDPDPAIPAPTATPSARGAVIRQNSDPTSEGPGPSPNPPAWVRPDNEAPPKVPQRTSSIATALNTSGAGGSRPAQAVRARPRSNSAWQIYLQRRAERGTPKPPGPPAQPPGPPNASSNPDLRRSDPGWERSDSVLPASHGHLPQAGSLERNRVGVSSKPDSSPVLSPGNKAKPDDHRSRPGRPADFVLLKERTLDEAPRPPKKAMDYSSSSEEVESSEDDEEEGEGGPAEGSRDTPGGRSDGDTDSVSTMVVHDVEEITGTQPPYGGGTMVVQRTPEEERNLLHADSNGYTNLPDVVQPSHSPTENSKGQSPPSKDGSGDYQSRGLVKAPGKSSFTMFVDLGIYQPGGSGDSIPITALVGGEGTRLDQLQYDVRKGSVVNVNPTNTRAHSETPEIRKYKKRFNSEILCAALWGVNLLVGTENGLMLLDRSGQGKVYGLIGRRRFQQMDVLEGLNLLITISGKRNKLRVYYLSWLRNKILHNDPEVEKKQGWTTVGDMEGCGHYRVVKYERIKFLVIALKSSVEVYAWAPKPYHKFMAFKSFADLPHRPLLVDLTVEEGQRLKVIYGSSAGFHAVDVDSGNSYDIYIPVHIQSQITPHAIIFLPNTDGMEMLLCYEDEGVYVNTYGRIIKDVVLQWGEMPTSVAYICSNQIMGWGEKAIEIRSVETGHLDGVFMHKRAQRLKFLCERNDKVFFASVRSGGSSQVYFMTLNRNCIMNW