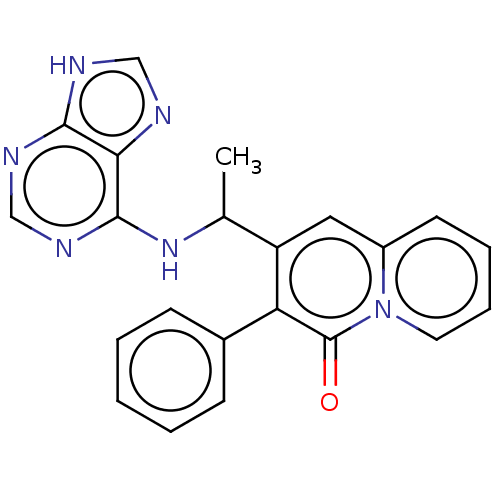
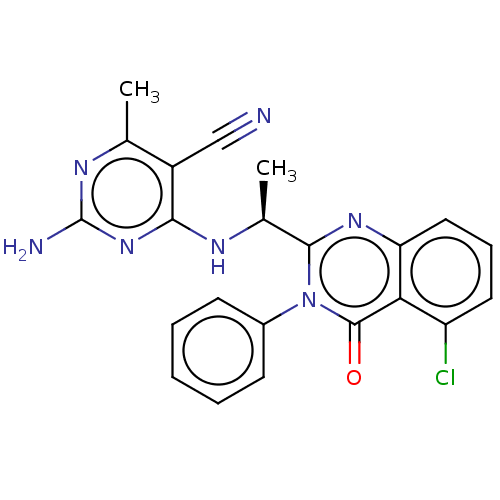
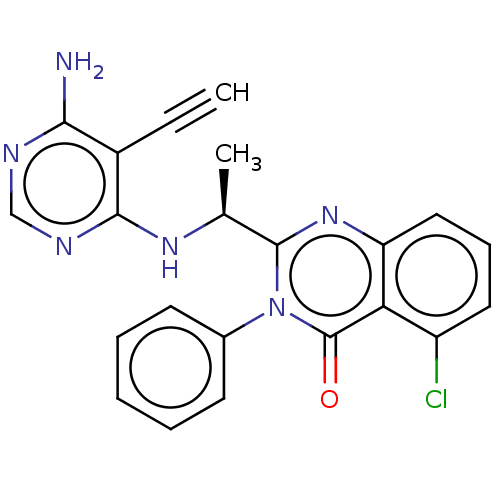
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.00100nMAssay Description:Inhibition of p110 delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.00300nMAssay Description:Inhibition of p110 delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.00300nMAssay Description:Inhibition of p110 delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.00500nMAssay Description:Inhibition of p110 delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.00500nMAssay Description:Inhibition of p110 delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.00700nMAssay Description:Inhibition of p110 delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0100nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0100nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0126nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0158nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataKi: 0.0200nMAssay Description:Inhibition of P13Kdelta (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataKi: 0.0240nMAssay Description:Binding affinity to p110 delta (unknown origin) assessed as inhibition constant in presence of ATP by TR-FRET assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0251nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0251nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0251nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0316nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0350nMpH: 7.5Assay Description:PI3K enzymatic activity was assayed by measuring the amount of product phosphatidylinositol 3,4,5-phosphate (PIP3) formed from substrate 4,5 phosphat...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0350nMpH: 7.5Assay Description:PI3K enzymatic activity was assayed by measuring the amount of product phosphatidylinositol 3,4,5-phosphate (PIP3) formed from substrate 4,5 phosphat...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0360nMAssay Description:Inhibition of p110 delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0398nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of PI3Kdelta (unknown origin) preincubated for 30 mins followed by insulin stimulation for 5 mins by Western blot analysisMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0430nMAssay Description:Inhibition of p110 delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0430nMAssay Description:Enzymatic activity of the class I PI3K isoforms in the presence of the compounds of Table 1 above was measured using a time-resolved fluorescence res...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0430nMAssay Description:The TR-FRET assay can monitor formation of the product 3,4,5-inositol triphosphate molecule (PIP3) as it competed with fluorescently labeled PIP3 for...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0480nMpH: 7.5Assay Description:PI3K enzymatic activity was assayed by measuring the amount of product phosphatidylinositol 3,4,5-phosphate (PIP3) formed from substrate 4,5 phosphat...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0501nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0501nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of PI3Kdelta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0600nMAssay Description:Inhibition of human p110delta/p85alpha using PIP2 as substrate in presence of ATP measured by HTRF assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0631nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0700nMAssay Description:Inhibition of human p110delta/p85alpha using PIP2 as substrate in presence of ATP measured by HTRF assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataKi: 0.0790nMAssay Description:PI3K Binding assays are intended for determining the biochemical potency of small molecule PI3K inhibitors. The PI3K lipid kinase reaction is perform...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataKi: 0.0794nMAssay Description:Inhibition of PI3Kdelta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataKi: 0.0794nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2 as substrate preincubated for 15 mins followed by substrate addition measured after 60 mins by HT...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0794nMAssay Description:Inhibition of recombinant human N-terminal His6-tagged full length P110delta/full length untagged human p85alpha expressed in baculovirus infected Sf...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0800nMAssay Description:Inhibition of recombinant human full length P13Kdelta expressed in Sf9 cells assessed as reduction in ATP-dependent phosphorylation by chromatography...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataKi: 0.0800nMAssay Description:Inhibition of human PI3K p110delta/p85alpha by HTRF assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0860nMAssay Description:Enzymatic activity of the class I PI3K isoforms in the presence of the compounds of Table 1 above was measured using a time-resolved fluorescence res...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.0860nMAssay Description:The TR-FRET assay can monitor formation of the product 3,4,5-inositol triphosphate molecule (PIP3) as it competed with fluorescently labeled PIP3 for...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataIC50: 0.0900nMAssay Description:Inhibition of recombinant human full length His-tagged p110delta/p85alpha expressed in baculovirus expression system measured after 60 mins in presen...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of PI3Kdelta (unknown origin) after 60 mins using fluorescein-labeled kinase tracer by HTRF assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of PI3Kdelta in human SUDHL6 cellsMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of PI-3K delta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of human full length PI3K p110delta/p85alpha using phosphatidylinositol 3,4,5-trisphosphate as substrate measured after 30 mins by TR-FRET...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition human full length PI3Kdelta catalytic subunit/p85alpha assessed as formation of PIP3 after 30 mins by europium labeled GRP-based TR-FRET a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of PI3Kdelta (unknown origin)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:TR-FRET monitored the formation of 3,4,5-inositol triphosphate molecule that competed with fluorescently labeled PIP3 for binding to the GRP-1 plecks...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Glaxosmithkline R&D
Curated by ChEMBL
Glaxosmithkline R&D
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition human full length PI3Kdelta catalytic subunit/p85alpha assessed as formation of PIP3 after 30 mins by europium labeled GRP-based TR-FRET a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.102nMpH: 7.5Assay Description:PI3K enzymatic activity was assayed by measuring the amount of product phosphatidylinositol 3,4,5-phosphate (PIP3) formed from substrate 4,5 phosphat...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Cancer Chemotherapy Center
Curated by ChEMBL
Cancer Chemotherapy Center
Curated by ChEMBL
Affinity DataIC50: 0.106nMpH: 7.5Assay Description:PI3K enzymatic activity was assayed by measuring the amount of product phosphatidylinositol 3,4,5-phosphate (PIP3) formed from substrate 4,5 phosphat...More data for this Ligand-Target Pair
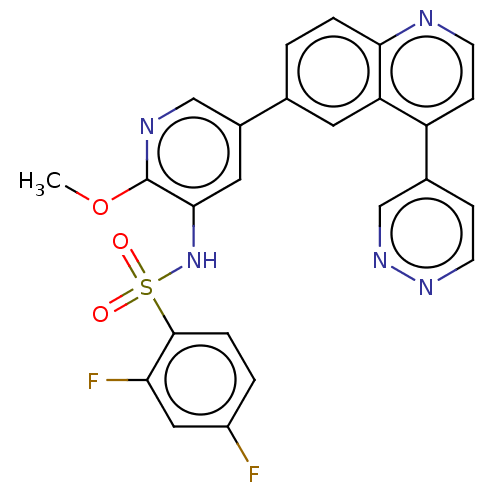



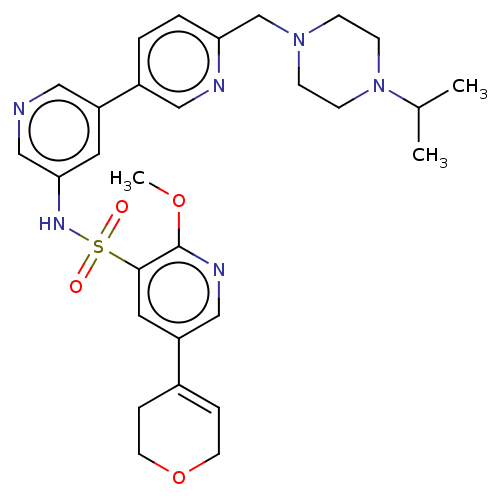
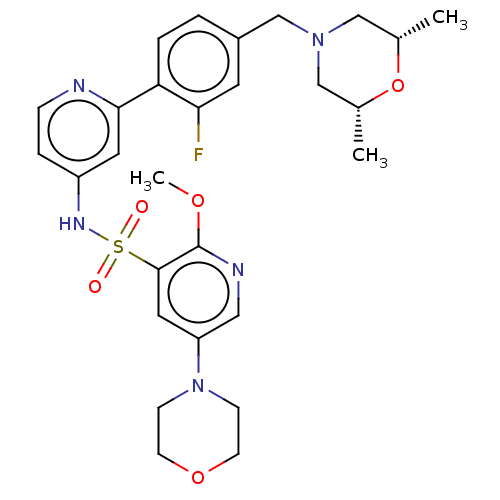


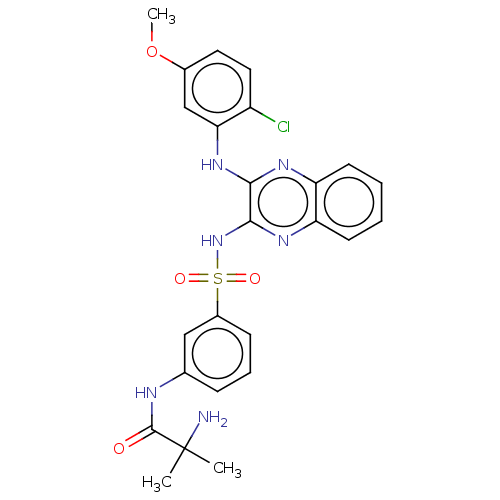
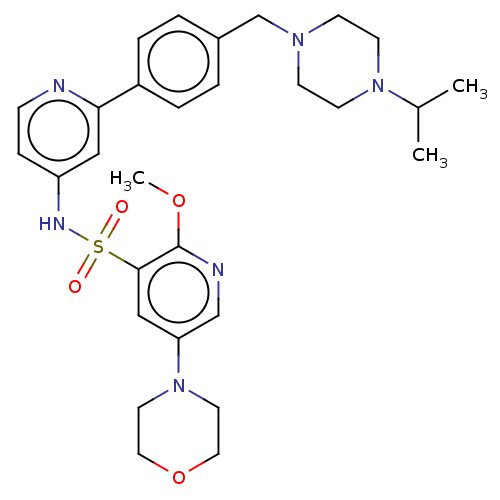


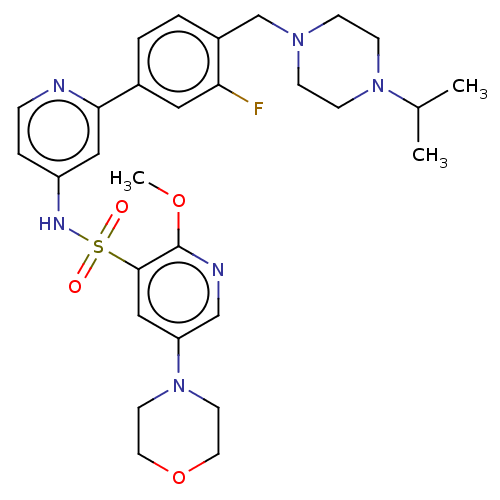

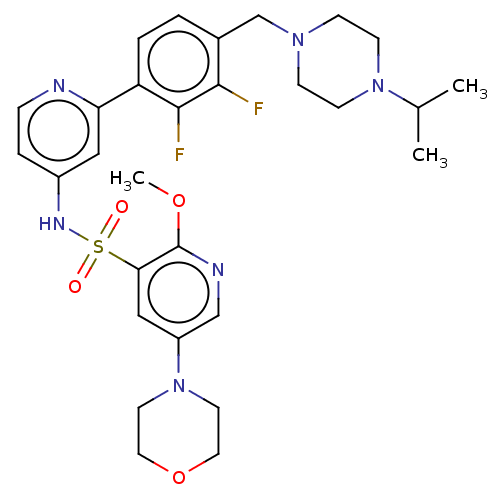
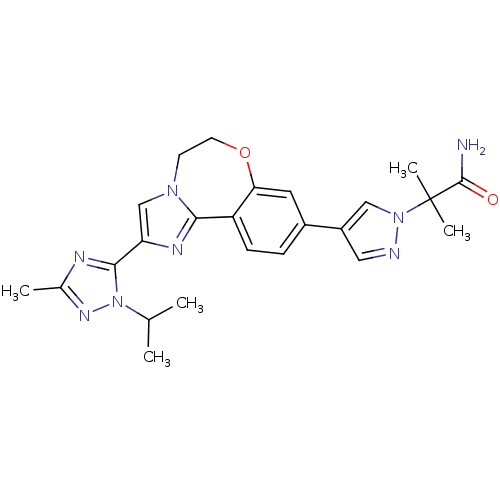
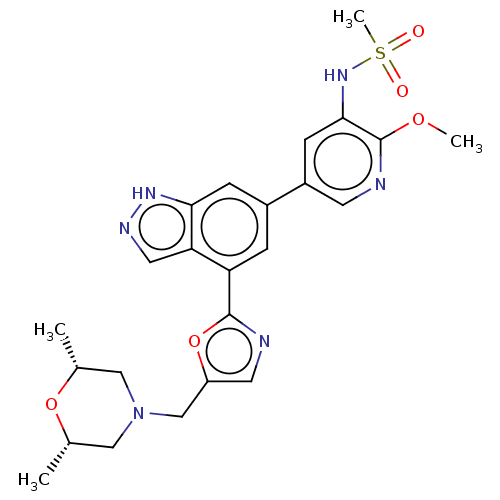


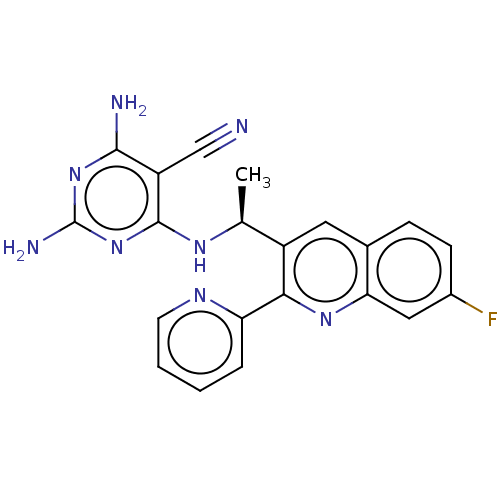
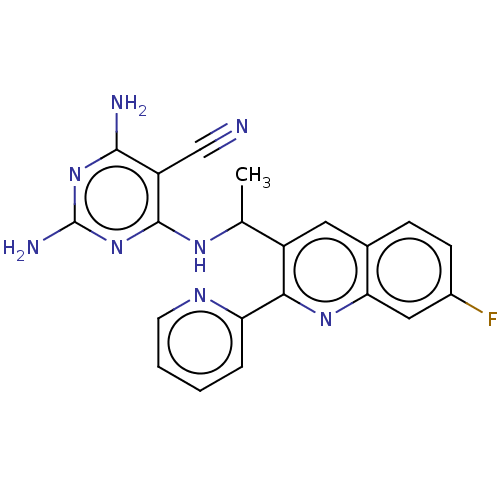
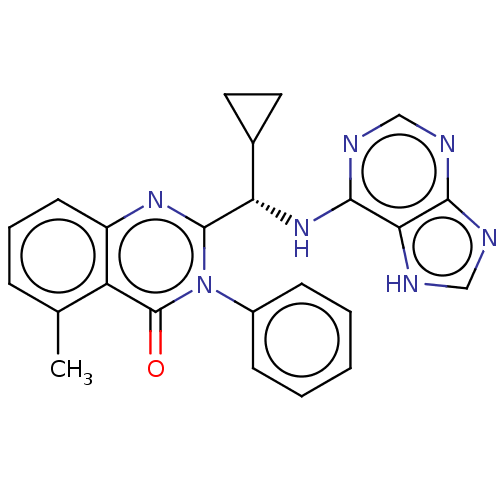
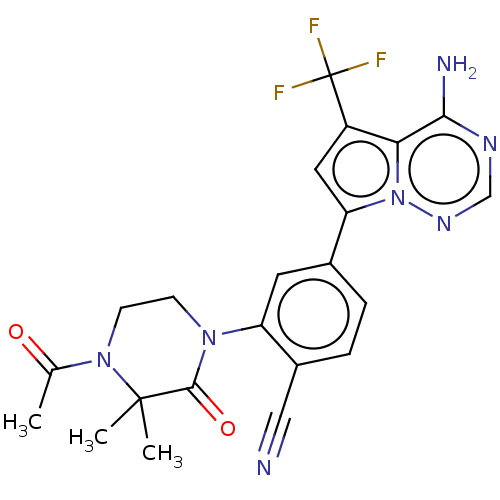

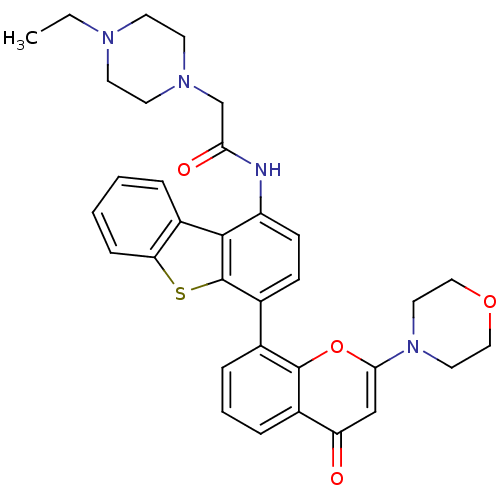
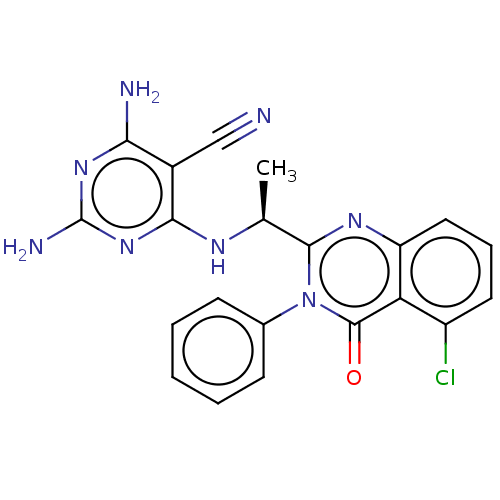
 3D Structure (crystal)
3D Structure (crystal)