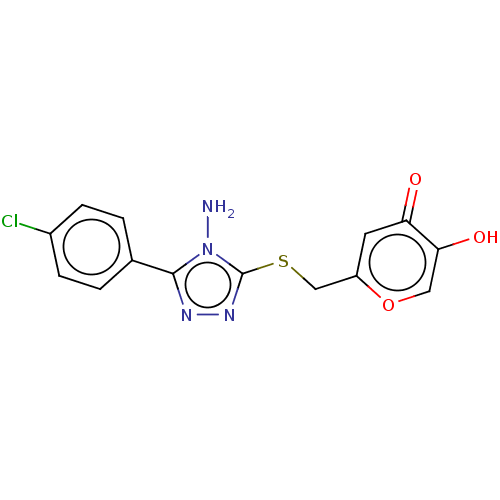
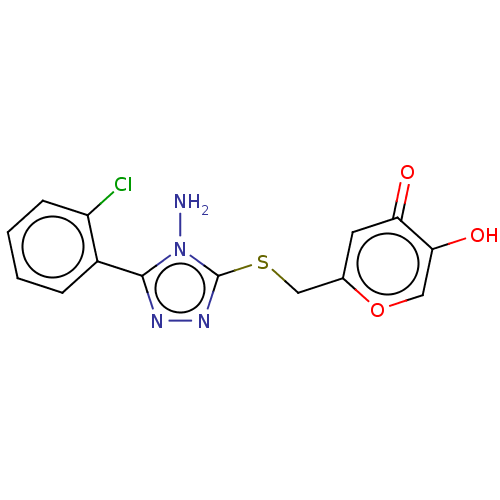
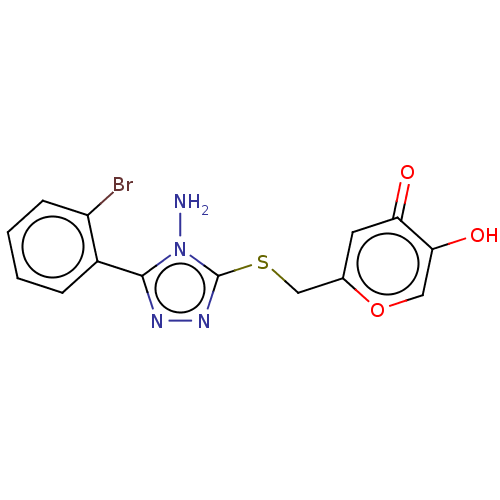
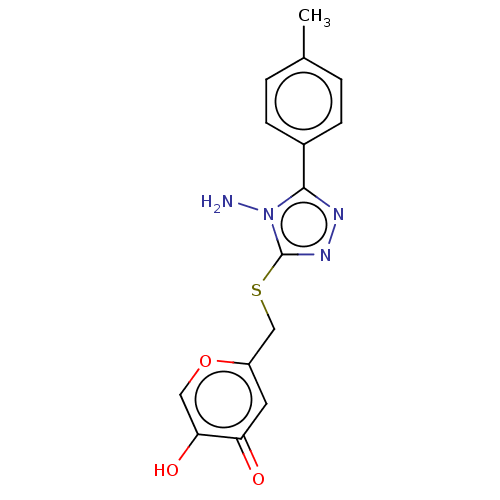
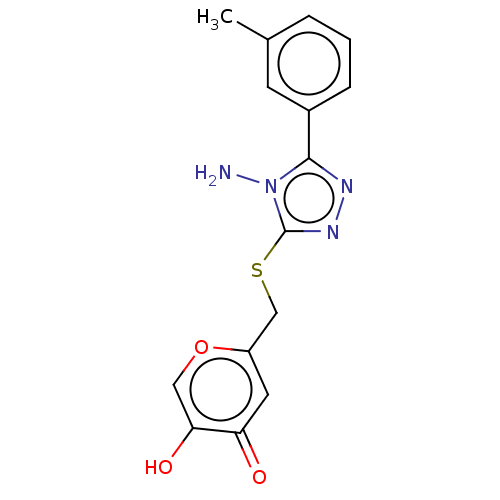
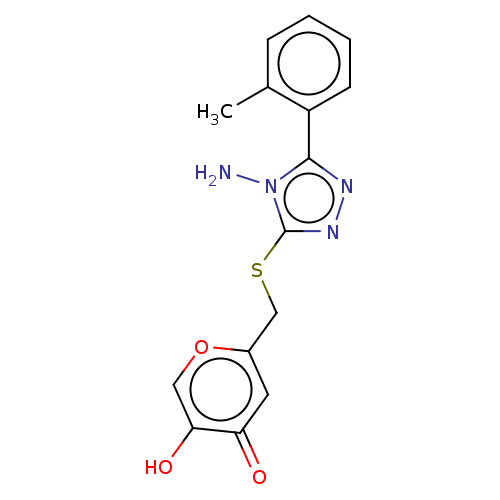
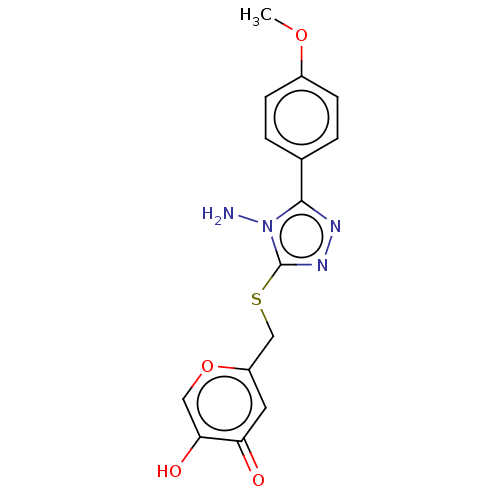
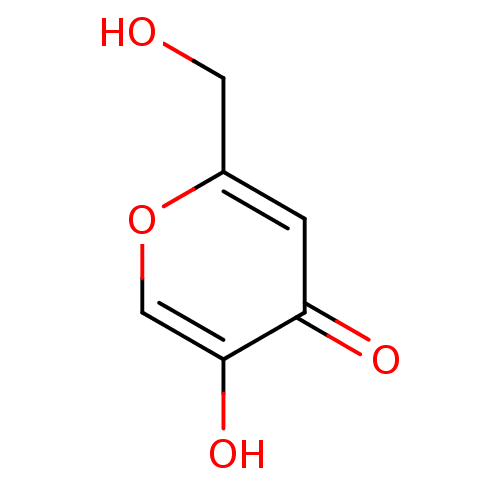
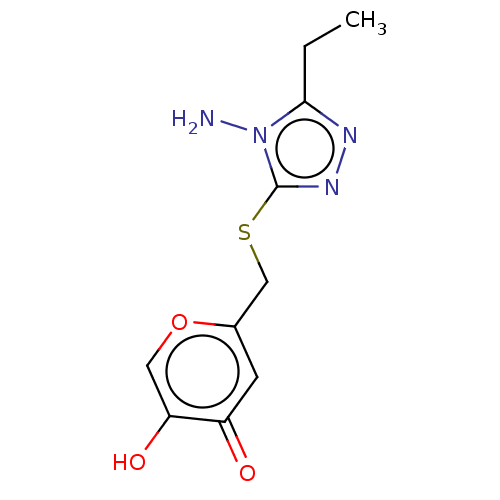
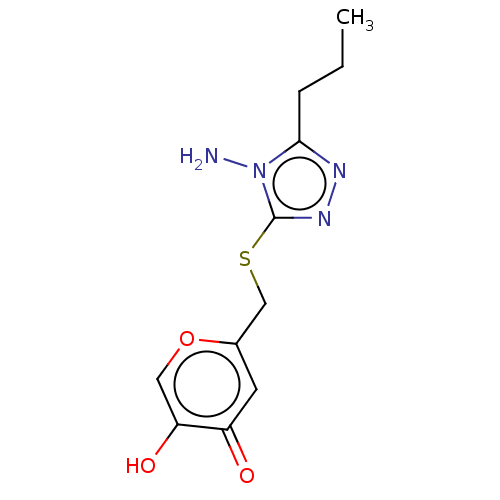
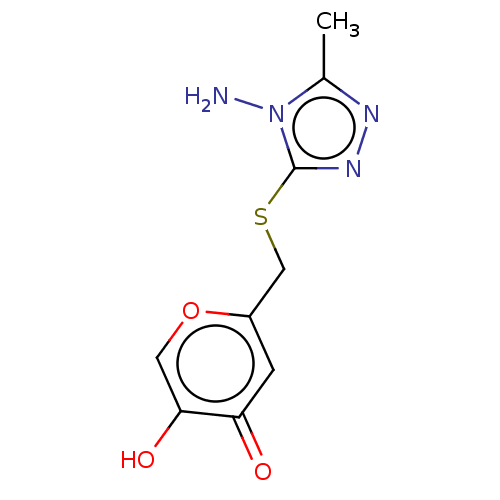
Affinity DataIC50: 4.50E+3nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.35E+3nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.98E+3nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.25E+3nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.82E+3nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.12E+3nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.45E+3nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.12E+4nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+4nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.57E+4nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.82E+4nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+4nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair