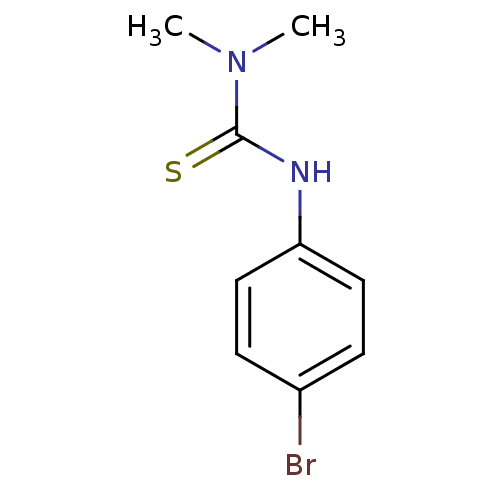
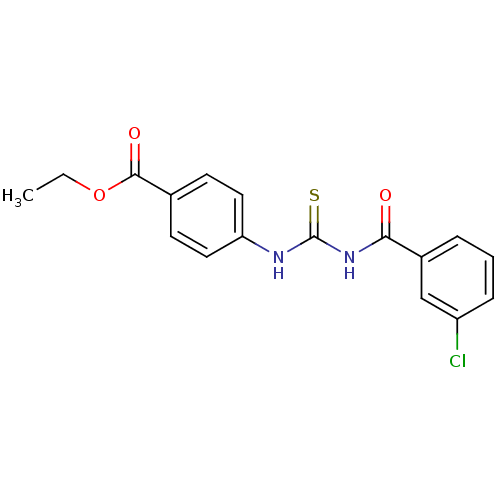
Affinity DataIC50: 2.20nMAssay Description:Inhibition of jack bean urease assessed as reduction in ammonia production using urea as substrate after 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Inhibition of jack bean ureaseMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of jack bean urease using urea as substrate pretreated for 15 mins followed by substrate addition by phenol red based spectrophotometric m...More data for this Ligand-Target Pair
Affinity DataIC50: 58nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Inhibition of jack bean urease using urea as substrate pretreated for 15 mins followed by substrate addition by phenol red based spectrophotometric m...More data for this Ligand-Target Pair
Affinity DataIC50: 64nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 72nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 77nMpH: 8.2Assay Description:The mixture of this enzyme inhibition assay was composed of 10 µL of enzyme (5 U/mL) and 10 µL of each test compound in the series (3a-3s) in 40 µL b...More data for this Ligand-Target Pair
Affinity DataIC50: 81nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 89nMpH: 8.2Assay Description:The mixture of this enzyme inhibition assay was composed of 10 µL of enzyme (5 U/mL) and 10 µL of each test compound in the series (3a-3s) in 40 µL b...More data for this Ligand-Target Pair
Affinity DataKi: 90nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataKi: 94nMAssay Description:Inhibition of jack bean ureaseMore data for this Ligand-Target Pair
Affinity DataIC50: 96nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 102nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 102nMpH: 8.2Assay Description:The mixture of this enzyme inhibition assay was composed of 10 µL of enzyme (5 U/mL) and 10 µL of each test compound in the series (3a-3s) in 40 µL b...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of jack beans urease assessed as hydrolysis of urea into ammonia preincubated for 10 mins followed by urea addition measured after 10 mins...More data for this Ligand-Target Pair
Affinity DataKi: 122nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataKi: 124nMAssay Description:Inhibition of jack bean ureaseMore data for this Ligand-Target Pair
Affinity DataKi: 127nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 127nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataKi: 127nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMpH: 8.2 T: 2°CAssay Description:The assay mixture contained 40 µL of buffer (0.01 M LiCl2, 1 mM EDTA, 0.01M K2HPO4, pH 8.2) containing 100 mM urea, 10 µL of enzyme (5U/mL)...More data for this Ligand-Target Pair
Affinity DataKi: 134nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataKi: 135nMAssay Description:Inhibition of jack bean ureaseMore data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of jack bean urease assessed as reduction in ammonia production using urea as substrate after 15 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of jack bean urease using urea as substrate pretreated for 15 mins followed by substrate addition by phenol red based spectrophotometric m...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of jack bean urease using urea as substrate pretreated for 15 mins followed by substrate addition by phenol red based spectrophotometric m...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of jack beans urease assessed as hydrolysis of urea into ammonia preincubated for 10 mins followed by urea addition measured after 10 mins...More data for this Ligand-Target Pair
Affinity DataKi: 152nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of jack beans urease assessed as hydrolysis of urea into ammonia preincubated for 10 mins followed by urea addition measured after 10 mins...More data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of jack bean urease using urea as substrate pretreated for 15 mins followed by substrate addition by phenol red based spectrophotometric m...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of jack bean urease using urea as substrate pretreated for 15 mins followed by substrate addition by phenol red based spectrophotometric m...More data for this Ligand-Target Pair
Affinity DataIC50: 177nMpH: 8.2Assay Description:The mixture of this enzyme inhibition assay was composed of 10 µL of enzyme (5 U/mL) and 10 µL of each test compound in the series (3a-3s) in 40 µL b...More data for this Ligand-Target Pair
Affinity DataIC50: 177nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 181nMpH: 8.2Assay Description:The mixture of this enzyme inhibition assay was composed of 10 µL of enzyme (5 U/mL) and 10 µL of each test compound in the series (3a-3s) in 40 µL b...More data for this Ligand-Target Pair
Affinity DataKi: 191nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Inhibition of Canavalia ensiformis (jack bean) urease assessed as release of ammonia after 30 min by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 210nMpH: 8.2 T: 2°CAssay Description:The assay mixture contained 40 µL of buffer (0.01 M LiCl2, 1 mM EDTA, 0.01M K2HPO4, pH 8.2) containing 100 mM urea, 10 µL of enzyme (5U/mL)...More data for this Ligand-Target Pair
Affinity DataIC50: 215nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of jack bean urease using urea as substrate pretreated for 15 mins followed by substrate addition by phenol red based spectrophotometric m...More data for this Ligand-Target Pair
Affinity DataIC50: 233nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair
Affinity DataKi: 255nMAssay Description:Inhibition of jack bean ureaseMore data for this Ligand-Target Pair
Affinity DataIC50: 260nMpH: 8.2 T: 2°CAssay Description:The assay mixture contained 40 µL of buffer (0.01 M LiCl2, 1 mM EDTA, 0.01M K2HPO4, pH 8.2) containing 100 mM urea, 10 µL of enzyme (5U/mL)...More data for this Ligand-Target Pair
Affinity DataIC50: 263nMAssay Description:Inhibition of jack bean urease assessed as ammonia production after 30 mins by indophenol methodMore data for this Ligand-Target Pair