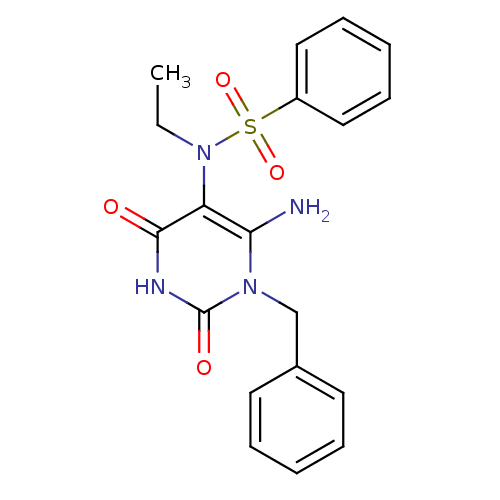
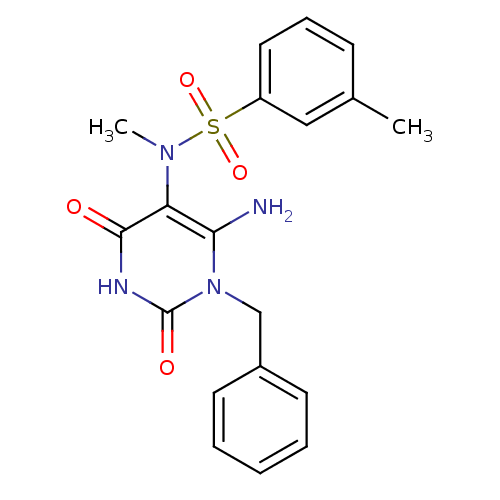
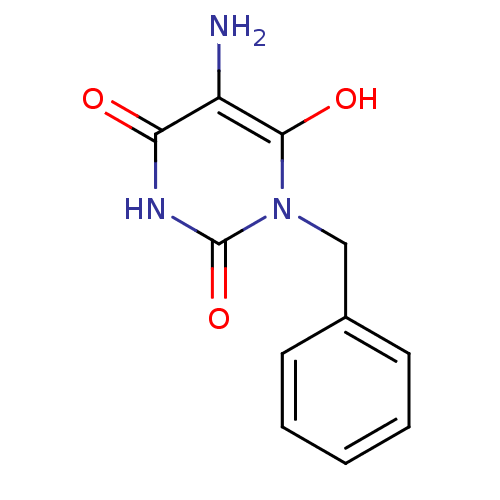
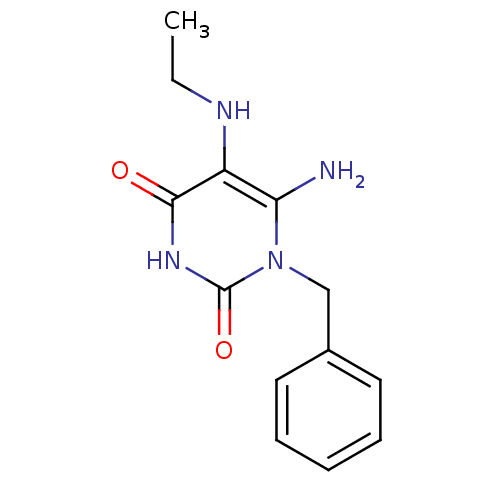
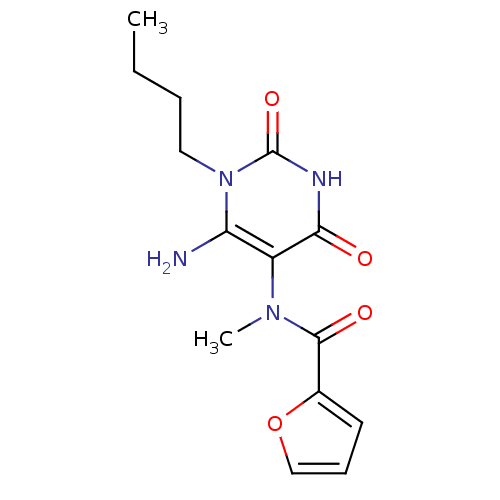
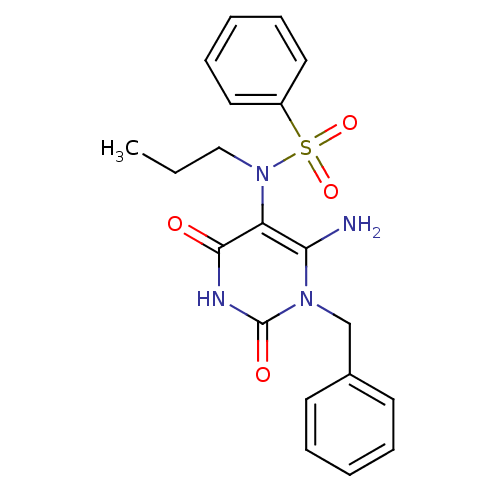
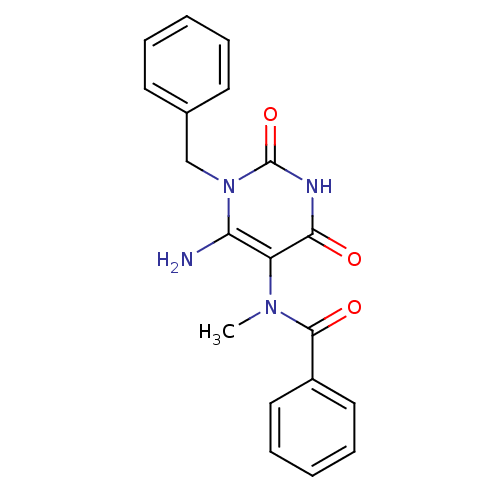
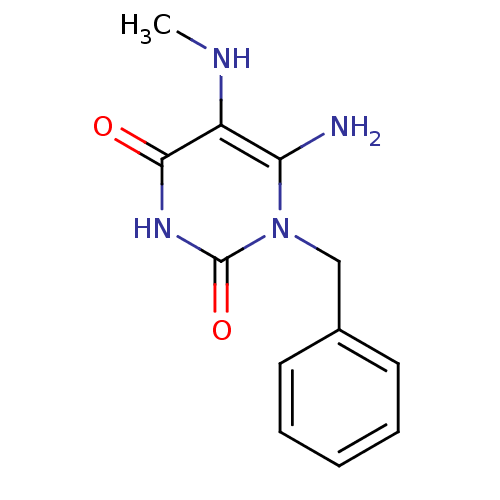
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 73nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
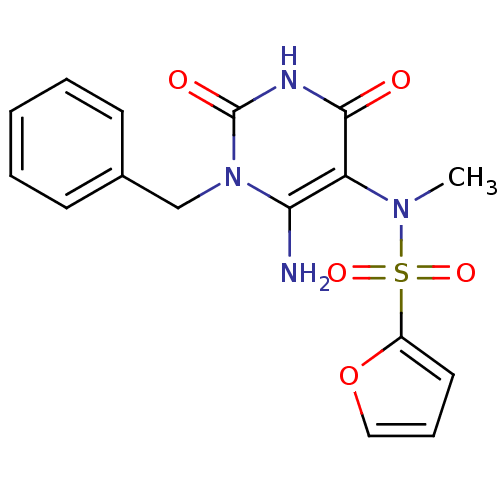
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 102nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
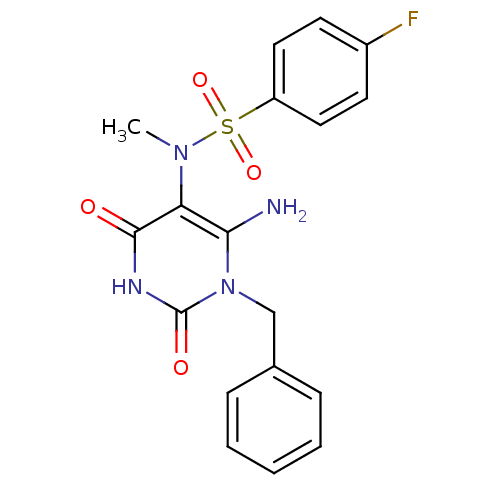
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 105nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
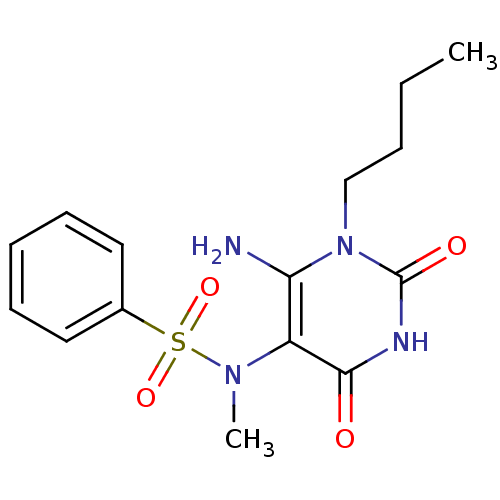
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 175nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 205nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 210nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 387nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 1.14E+3nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 1.32E+3nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 4.70E+3nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 5.90E+3nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 6.00E+4nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
TargetGlucose-1-phosphate thymidylyltransferase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
University of St. Andrews
University of St. Andrews
Affinity DataIC50: 6.00E+4nMAssay Description:Inhibition assay using glucose-1-phosphate thymidylyltransferase (RmlA).More data for this Ligand-Target Pair
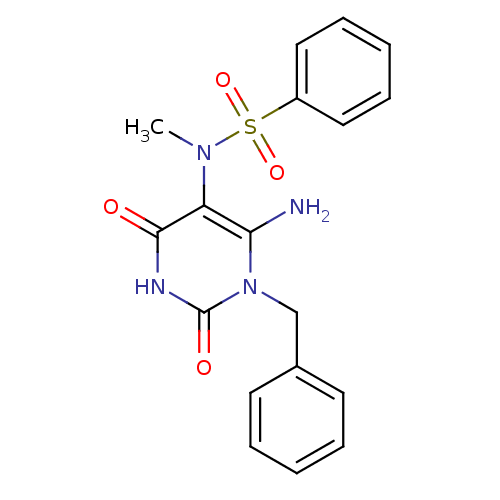
 3D Structure (crystal)
3D Structure (crystal)