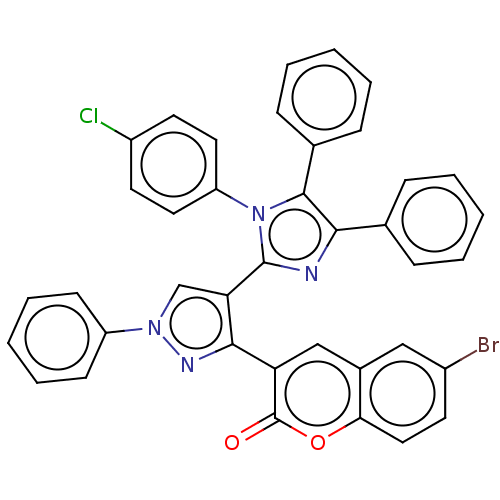
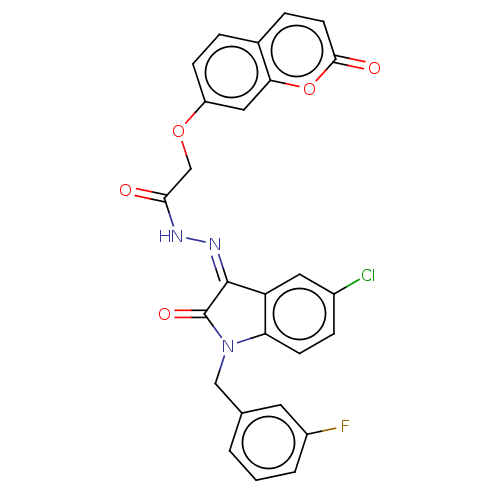
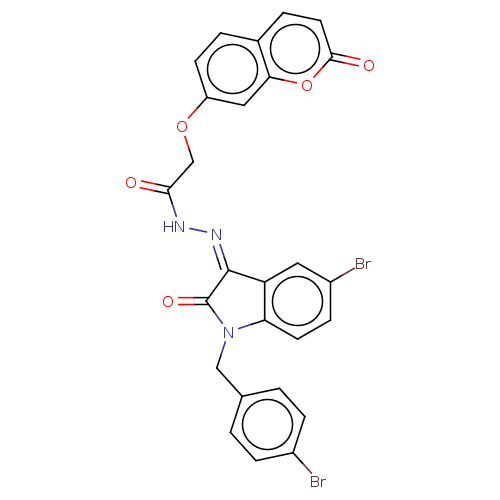
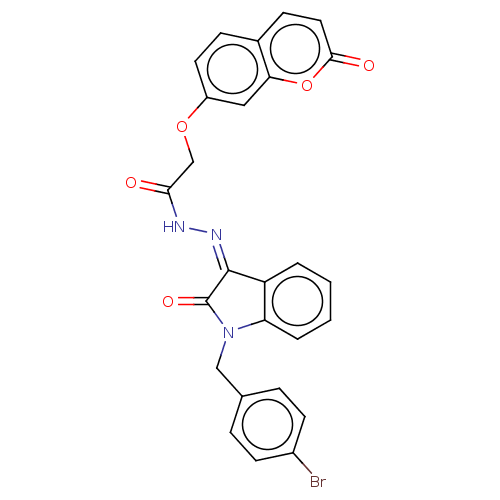
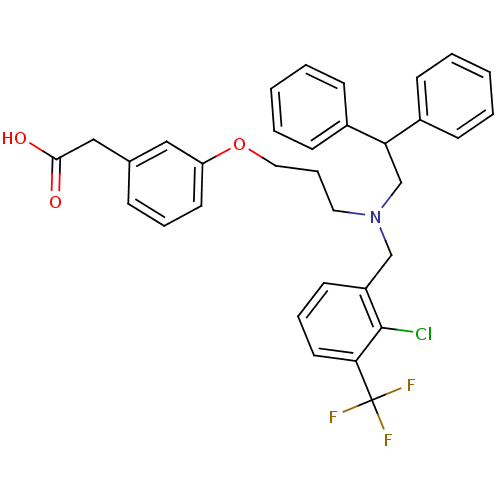
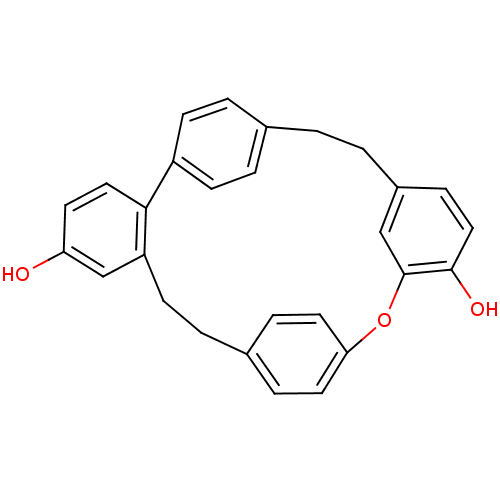
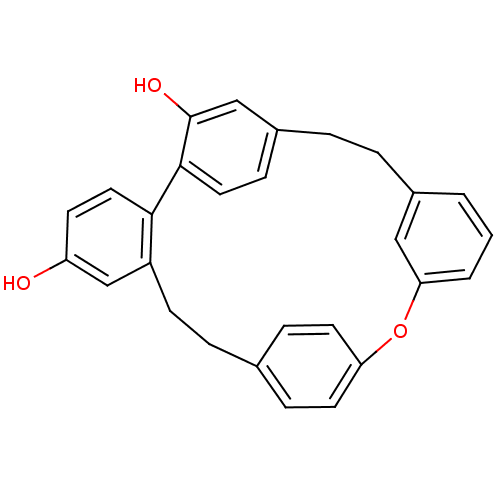
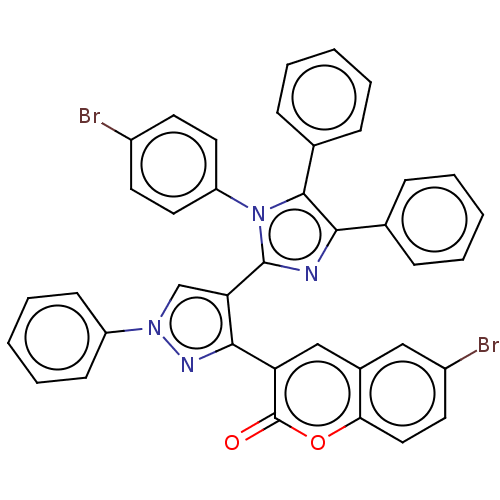
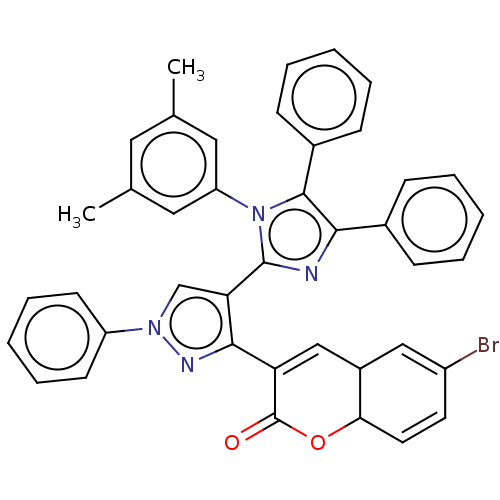
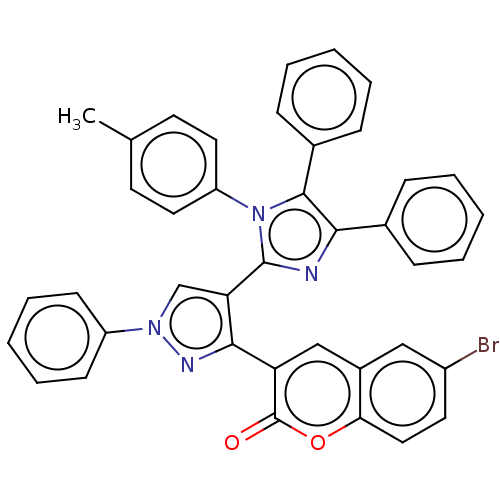
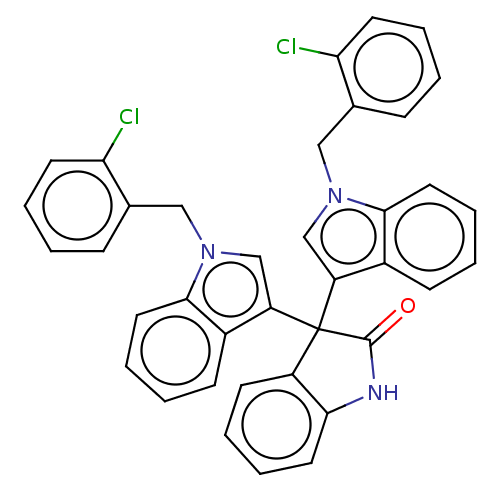
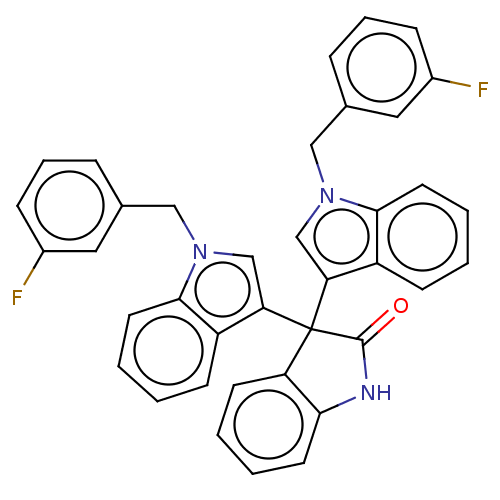
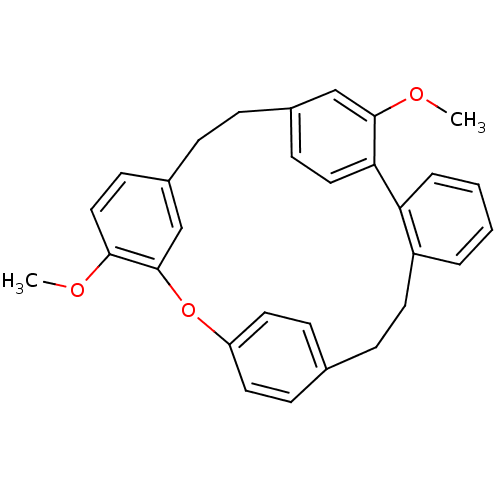
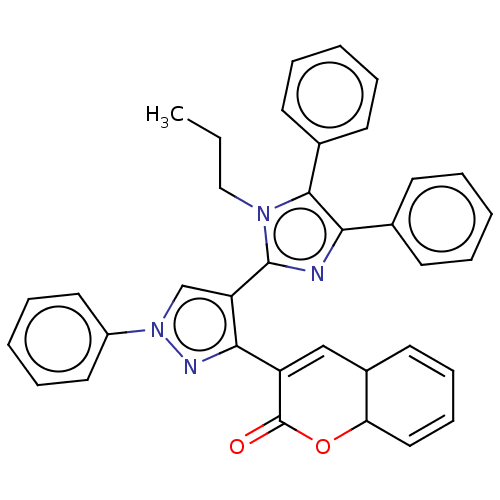
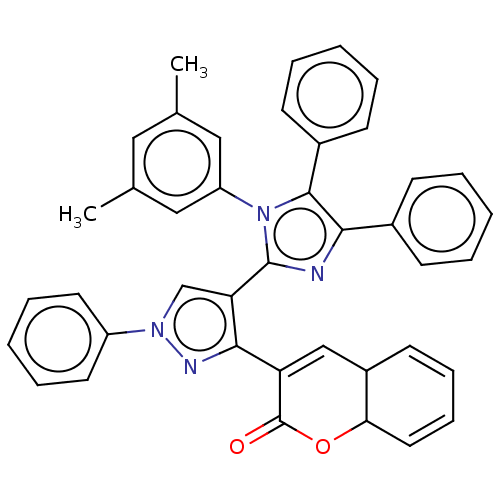
Affinity DataIC50: 2.53E+3nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 2.56E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 2.85E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 2.88E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 3.98E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 4.07E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 4.48E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+3nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+3nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+3nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 5.21E+3nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 5.64E+3nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 5.83E+3nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 5.98E+3nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 7.31E+3nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 7.31E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 7.37E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 7.92E+3nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 8.60E+3nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 9.16E+3nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 9.27E+3nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 9.33E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 9.47E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 9.83E+3nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 9.90E+3nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+4nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.27E+4nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 1.36E+4nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 1.37E+4nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 1.41E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.42E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+4nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.62E+4nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 2.05E+4nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 2.47E+4nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 2.54E+4nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 3.01E+4nMpH: 6.8 T: 2°CAssay Description:The α-glucosidase inhibition assay had been carried out using baker's yeast α-glucosidase (EC 3.2.1.20) and p-nitrophenyl α-d-gluco...More data for this Ligand-Target Pair
Affinity DataIC50: 3.44E+4nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 3.83E+4nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+4nMpH: 7.0 T: 2°CAssay Description:The alpha-glucosidase inhibitory activity of test compounds was determined in a 96-well plate format. The reaction mixture containing enzyme and chro...More data for this Ligand-Target Pair
Affinity DataIC50: 4.48E+4nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair
Affinity DataIC50: 4.64E+4nMpH: 6.8 T: 2°CAssay Description:For α-glucosidase inhibition assay, α-glucosidase enzyme (Cat No. 5003-1KU Type I; isolated from Saccharomyces cereviciae) was used. Acar...More data for this Ligand-Target Pair