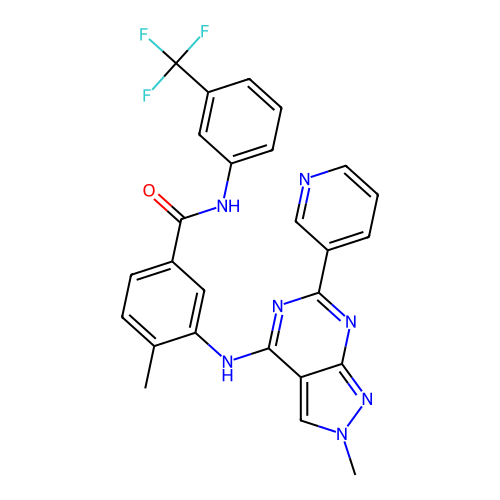
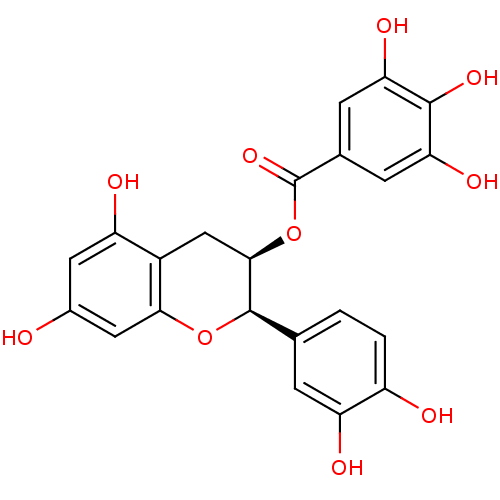
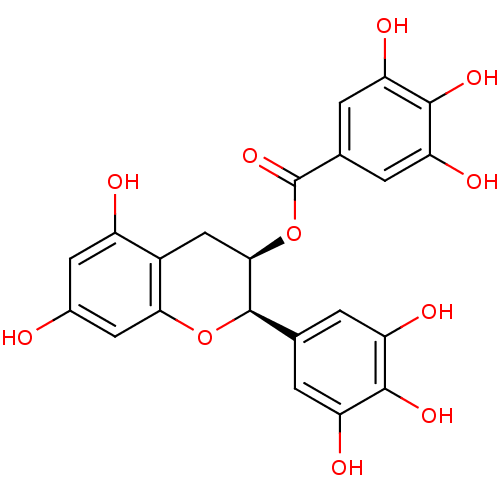
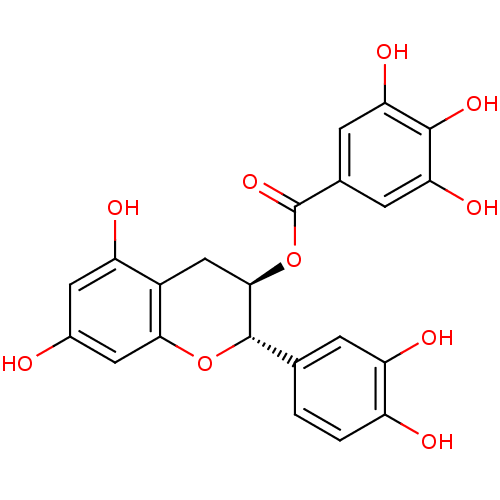
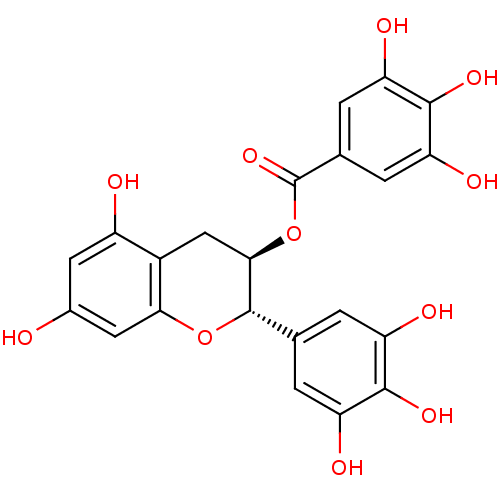
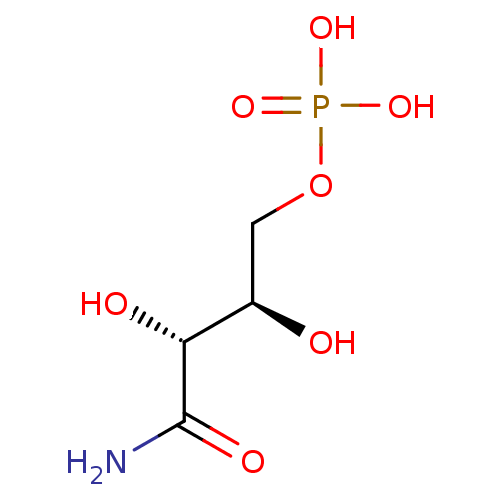
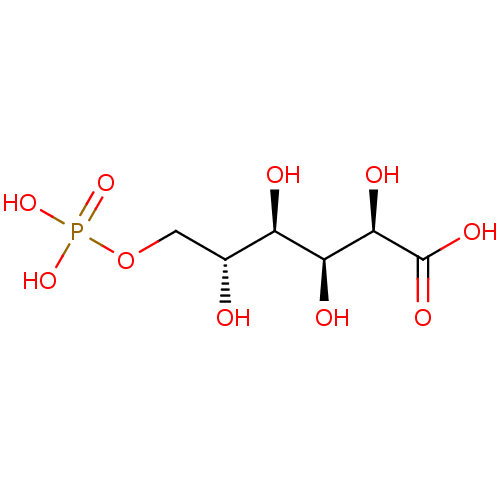
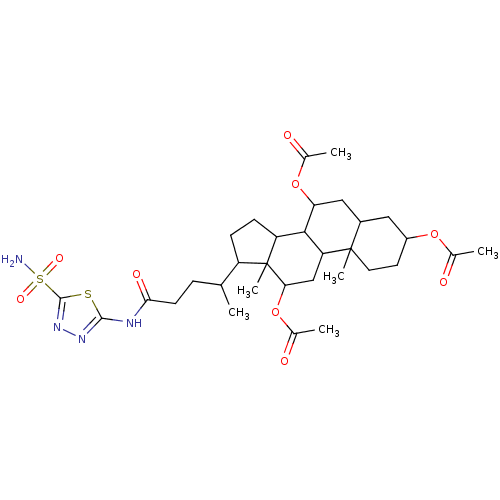
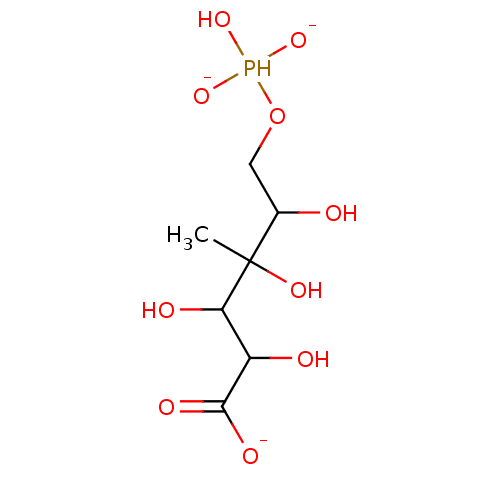
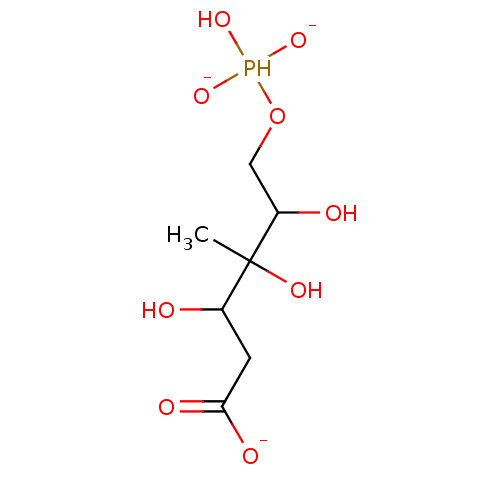
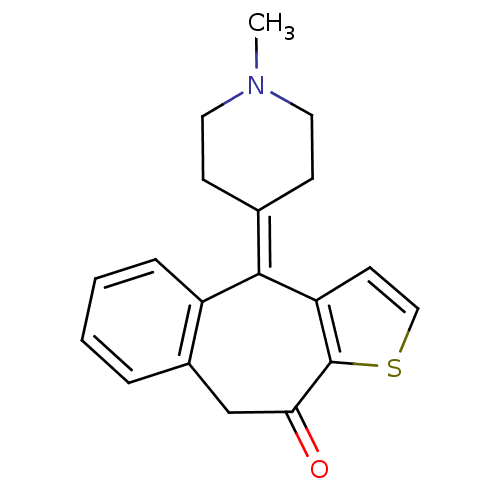
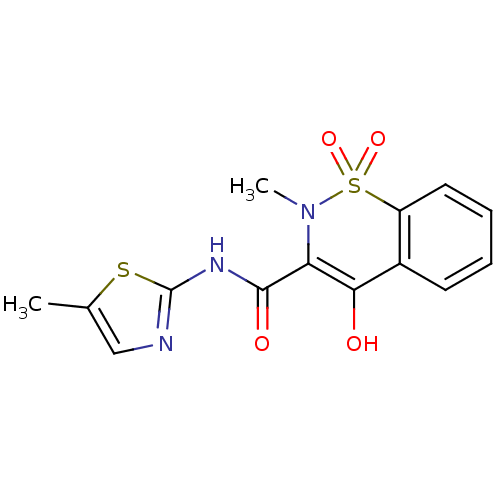
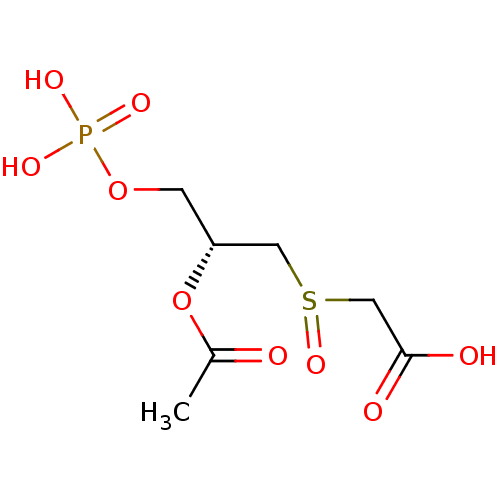
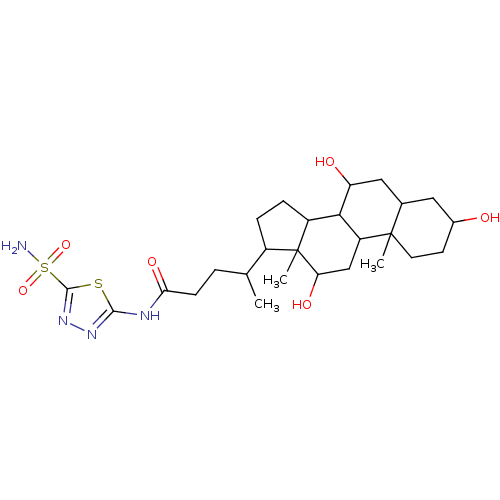
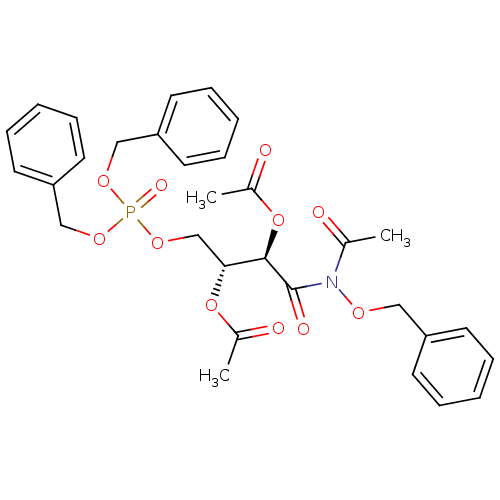
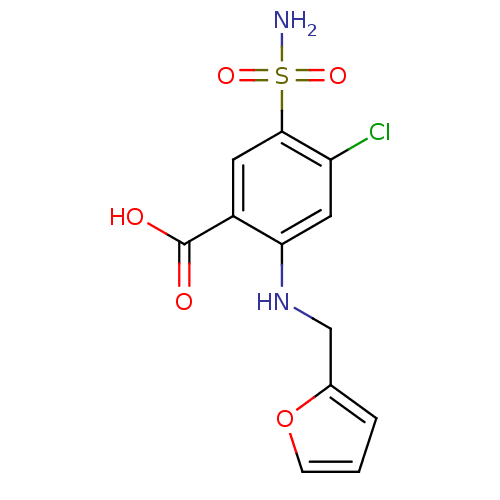
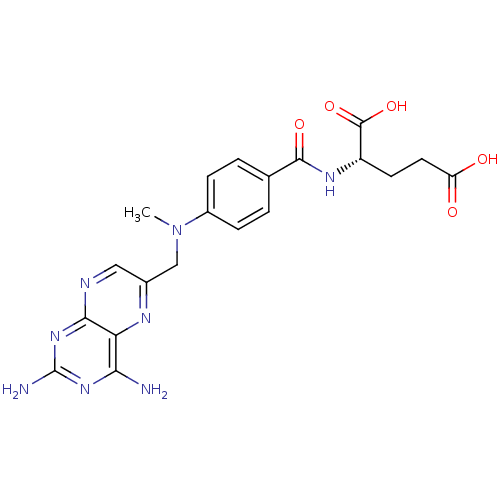
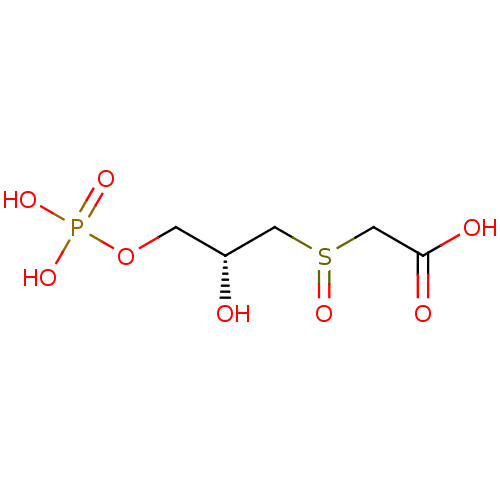
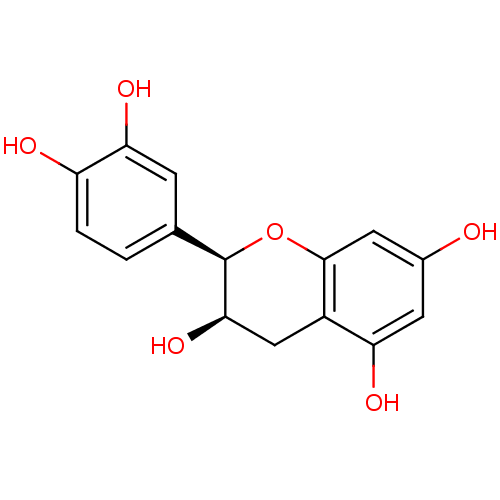
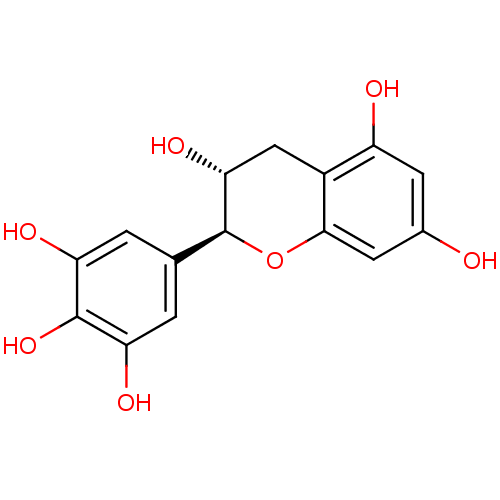
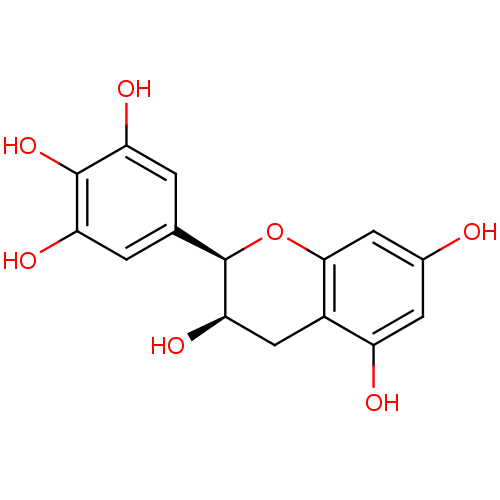
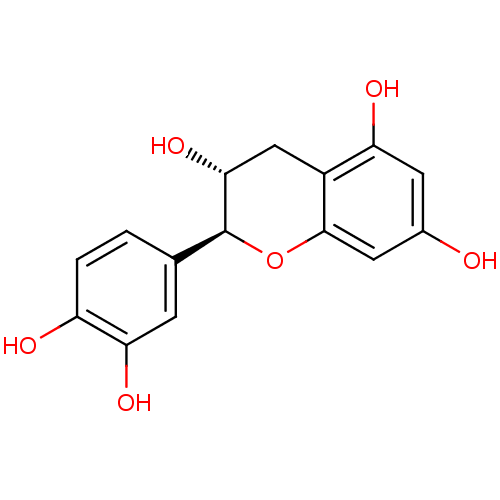
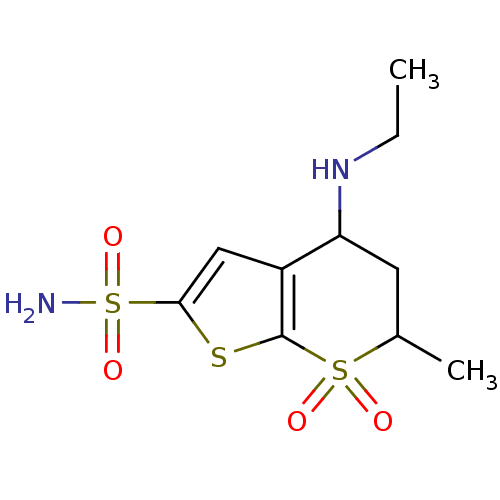
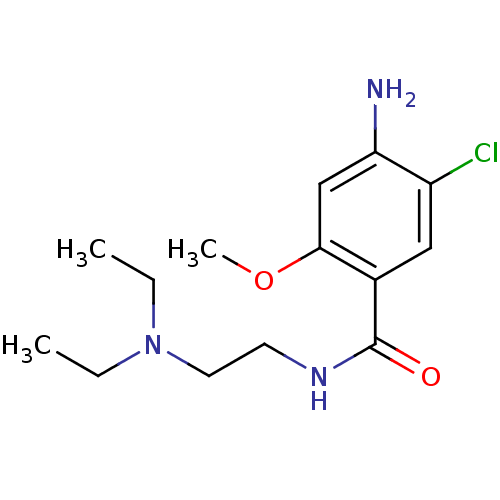
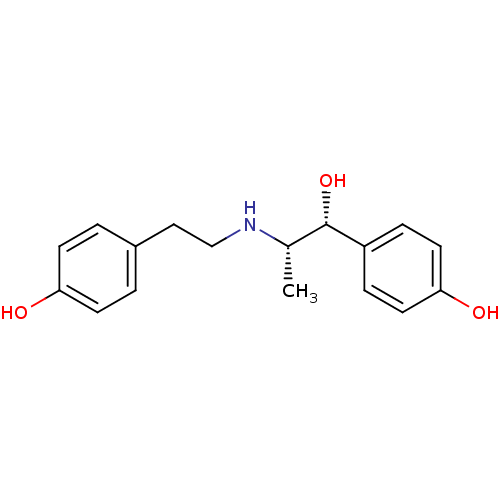
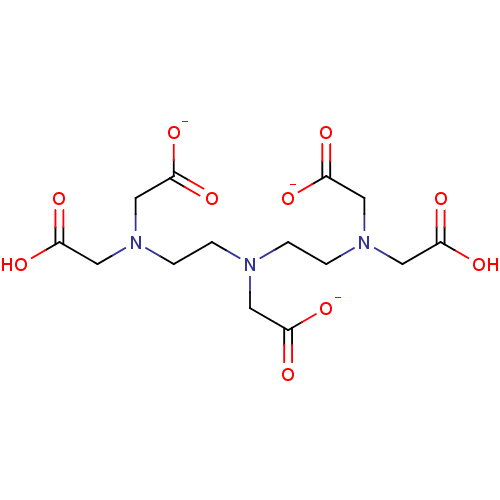
Affinity DataKi: 10nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKd: 131nMAssay Description:Binding affinity to human PGD incubated for 45 mins by Kinobead based pull down assayMore data for this Ligand-Target Pair
Affinity DataKi: 360nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+3nMAssay Description:Inhibition of 6PGDMore data for this Ligand-Target Pair
Affinity DataIC50: 1.45E+3nMAssay Description:Inhibition of 6PGDMore data for this Ligand-Target Pair
Affinity DataKi: 1.52E+3nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKd: 2.00E+3nMAssay Description:log Kd which is the binding affinity against 6-phosphogluconate dehydrogenaseMore data for this Ligand-Target Pair
Affinity DataKi: 4.20E+3nM IC50: 2.53E+3nMAssay Description:The enzyme activities in the absence of drugs or chemicals were taken as 100%. For each drug or chemical, an activity%-(drug) graph was drawn and dr...More data for this Ligand-Target Pair
Affinity DataKi: 2.54E+3nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+3nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 4.40E+3nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 8.30E+3nM IC50: 8.40E+3nMAssay Description:In order to determine the effects of some drugs on human 6PGD, concentrations of ketotifen (0.0018-0.0282 mM), dacarbazine (0.0049-0.054 mM), meloxic...More data for this Ligand-Target Pair
Affinity DataKi: 1.01E+4nM IC50: 1.20E+4nMAssay Description:In order to determine the effects of some drugs on human 6PGD, concentrations of ketotifen (0.0018-0.0282 mM), dacarbazine (0.0049-0.054 mM), meloxic...More data for this Ligand-Target Pair
Affinity DataKi: 1.07E+4nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKi: 1.60E+4nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKd: 2.85E+4nMAssay Description:Binding affinity to human PGD incubated for 45 mins by Kinobead based pull down assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.92E+4nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKi: 4.00E+4nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 5.09E+4nM IC50: 5.50E+4nMAssay Description:In order to determine the effects of some drugs on human 6PGD, concentrations of ketotifen (0.0018-0.0282 mM), dacarbazine (0.0049-0.054 mM), meloxic...More data for this Ligand-Target Pair
Affinity DataKi: 8.78E+4nM IC50: 6.01E+4nMAssay Description:The enzyme activities in the absence of drugs or chemicals were taken as 100%. For each drug or chemical, an activity%-(drug) graph was drawn and dr...More data for this Ligand-Target Pair
Affinity DataKi: 1.09E+5nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 1.27E+5nM IC50: 1.25E+5nMAssay Description:In order to determine the effects of some drugs on human 6PGD, concentrations of ketotifen (0.0018-0.0282 mM), dacarbazine (0.0049-0.054 mM), meloxic...More data for this Ligand-Target Pair
Affinity DataKi: 1.37E+5nM IC50: 1.42E+5nMAssay Description:In order to determine the effects of some drugs on human 6PGD, concentrations of ketotifen (0.0018-0.0282 mM), dacarbazine (0.0049-0.054 mM), meloxic...More data for this Ligand-Target Pair
Affinity DataKi: 1.92E+5nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKi: 1.98E+5nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 5.88E+5nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of Trypanosoma brucei expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 6.25E+5nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKi: 7.14E+5nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKi: 7.70E+5nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataKi: 7.84E+5nMAssay Description:Inhibition constant against 6-phosphogluconate dehydrogenase of sheep liverMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of 6PGDMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of 6PGDMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of 6PGDMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of 6PGDMore data for this Ligand-Target Pair
Affinity DataKi: 3.14E+6nM IC50: 1.41E+6nMAssay Description:The enzyme activities in the absence of drugs or chemicals were taken as 100%. For each drug or chemical, an activity%-(drug) graph was drawn and dr...More data for this Ligand-Target Pair
Affinity DataKi: 2.11E+6nM IC50: 3.27E+6nMAssay Description:In order to determine the effects of some drugs on human 6PGD, concentrations of ketotifen (0.0018-0.0282 mM), dacarbazine (0.0049-0.054 mM), meloxic...More data for this Ligand-Target Pair
Affinity DataKi: 6.04E+6nM IC50: 3.66E+6nMAssay Description:In order to determine the effects of some drugs on human 6PGD, concentrations of ketotifen (0.0018-0.0282 mM), dacarbazine (0.0049-0.054 mM), meloxic...More data for this Ligand-Target Pair
Affinity DataKi: 7.34E+7nM IC50: 2.09E+8nMAssay Description:In order to determine the effects of some drugs on human 6PGD, concentrations of ketotifen (0.0018-0.0282 mM), dacarbazine (0.0049-0.054 mM), meloxic...More data for this Ligand-Target Pair
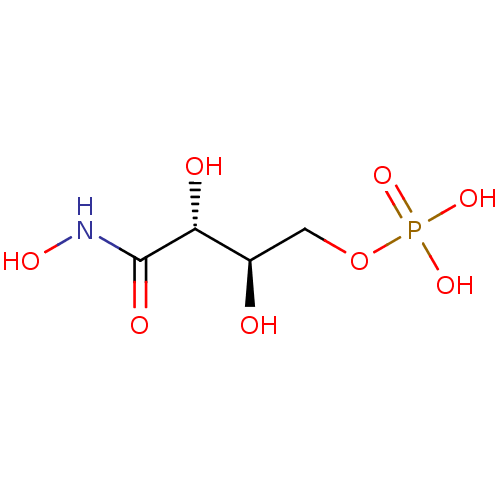
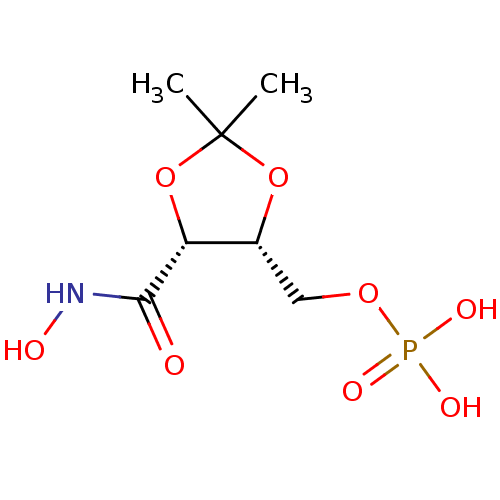
 3D Structure (crystal)
3D Structure (crystal)