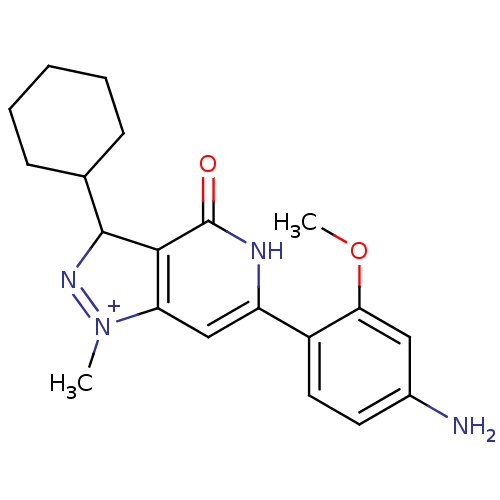
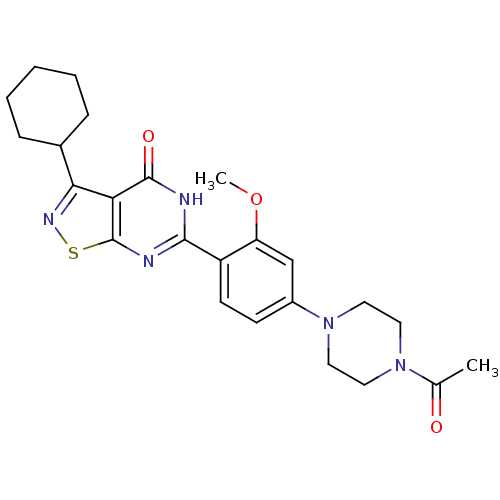
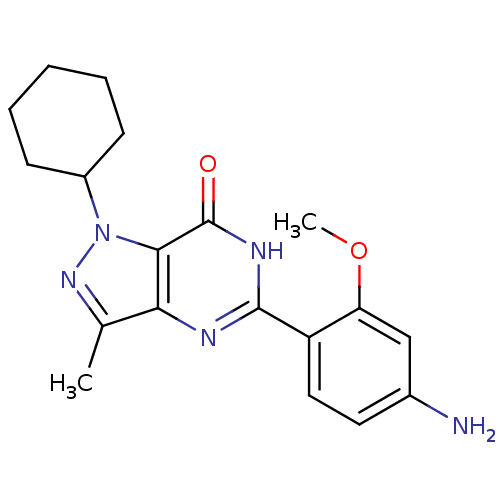
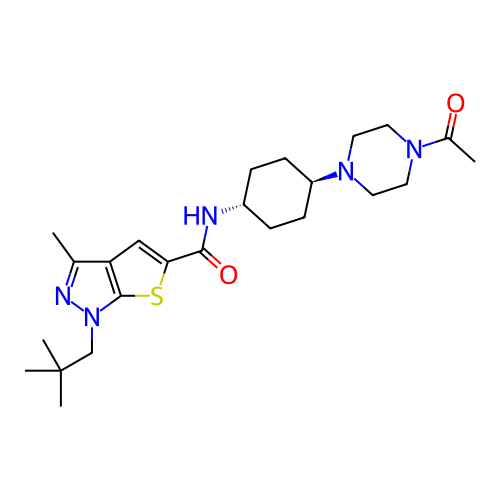
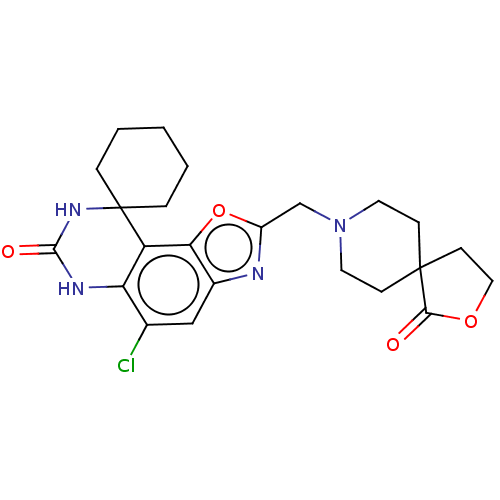
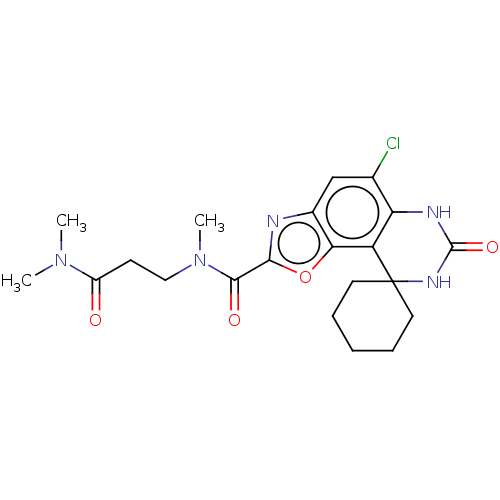
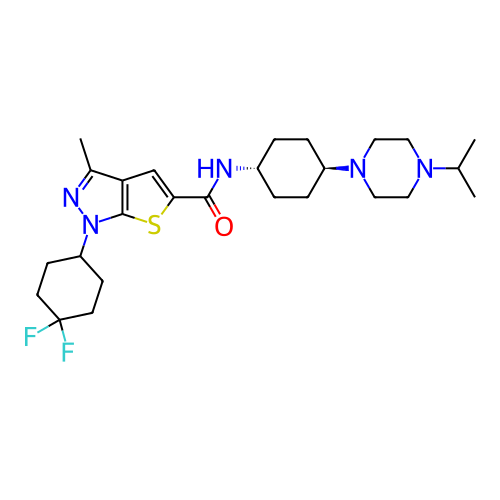
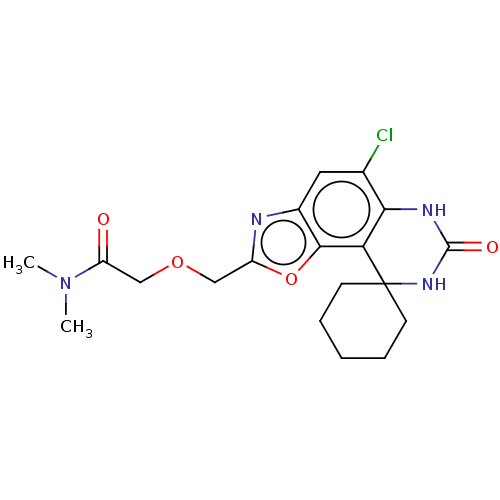
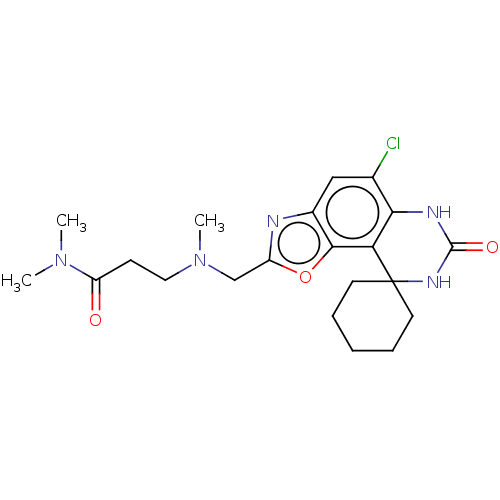
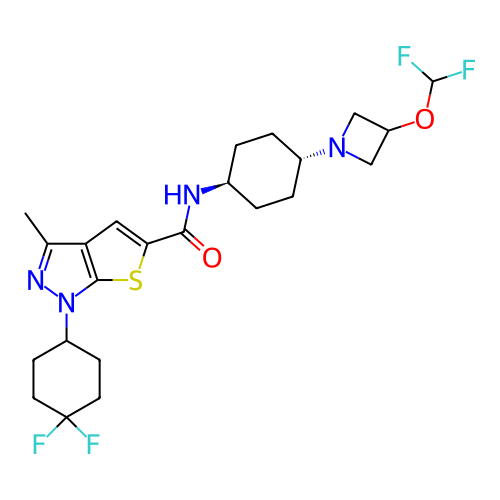
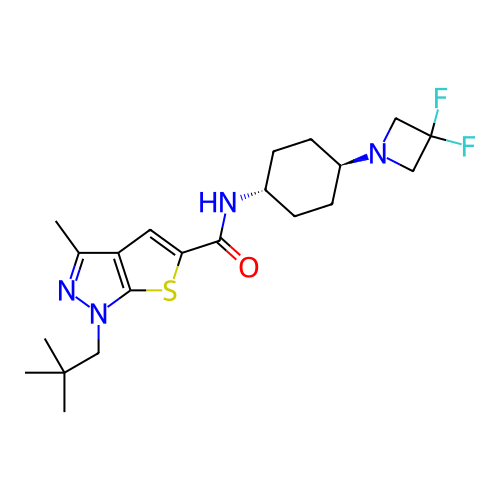
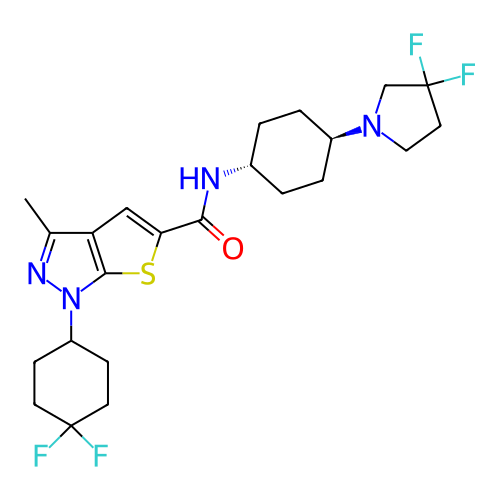
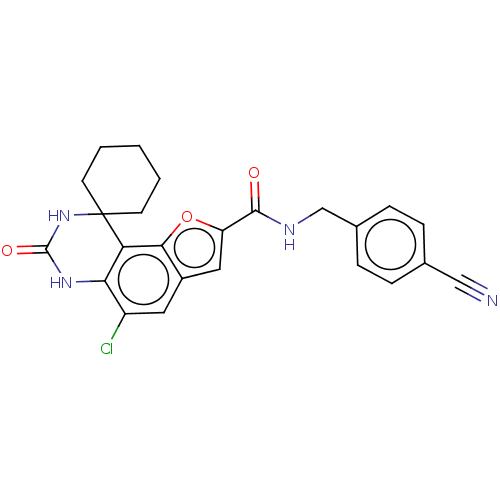
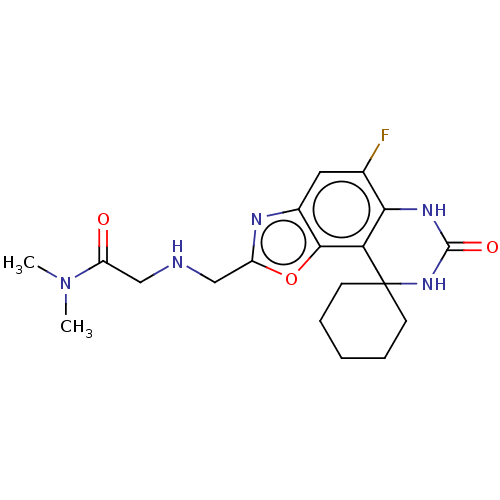
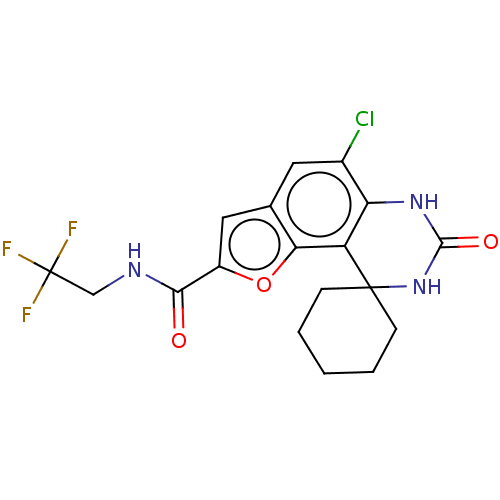
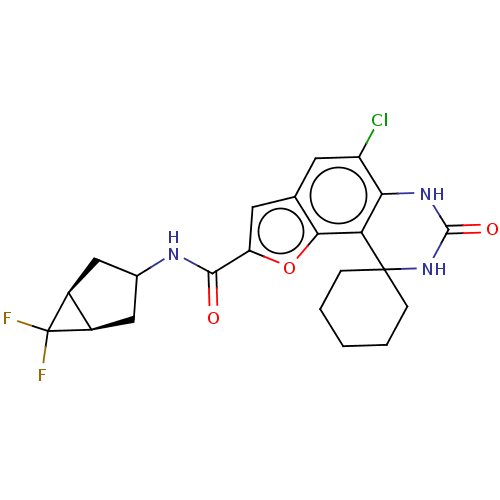
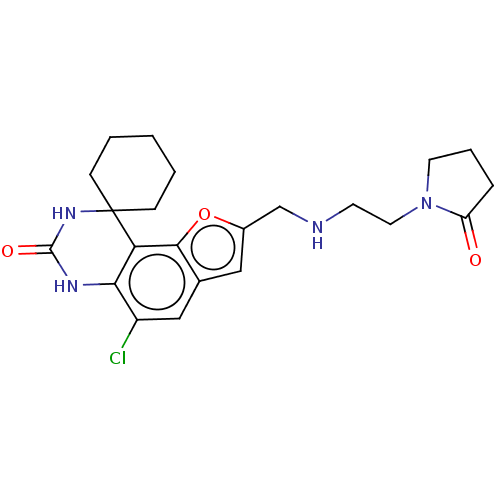
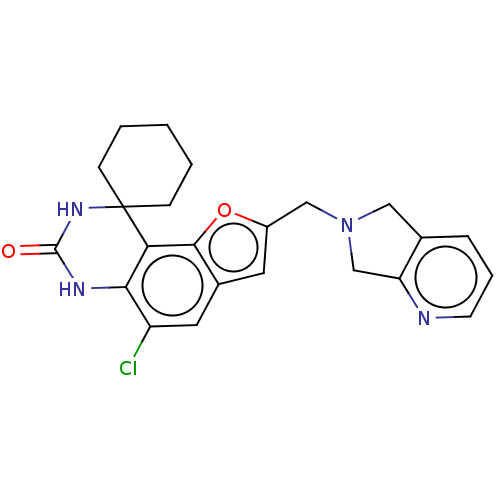
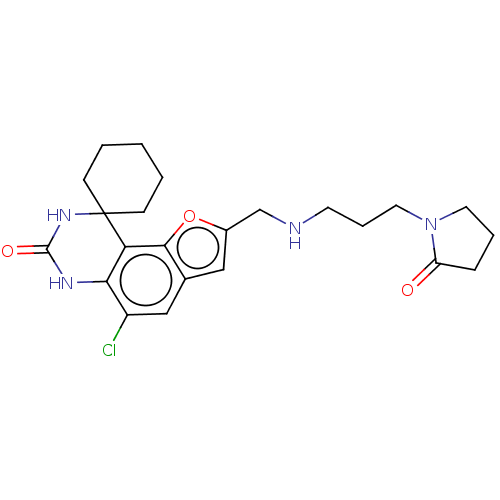
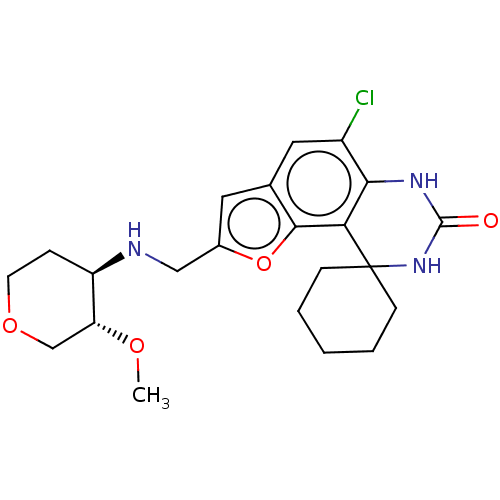
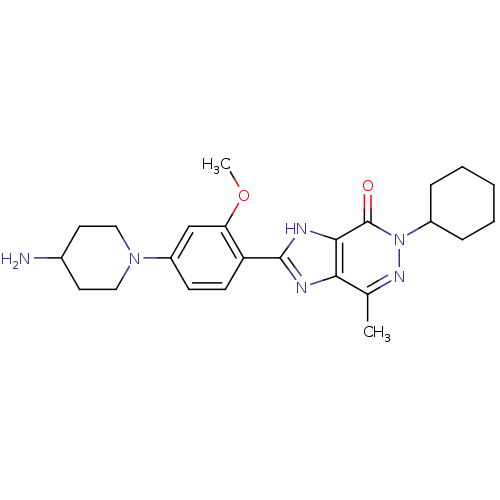
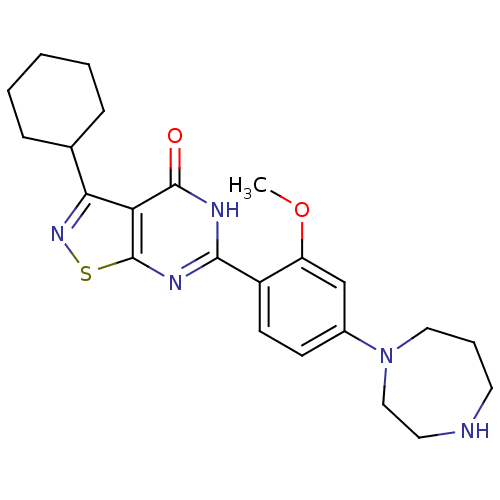
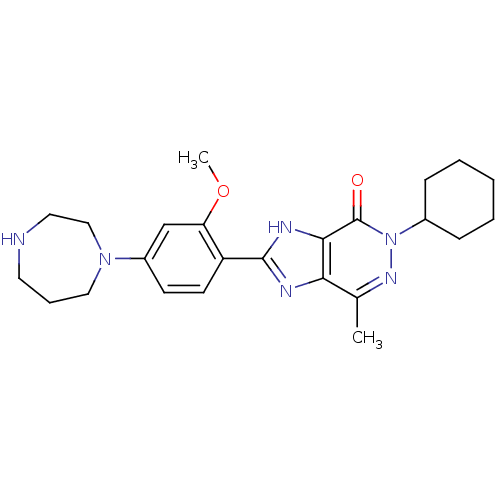
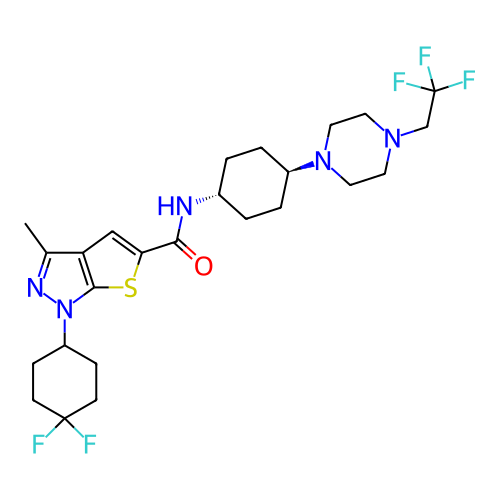
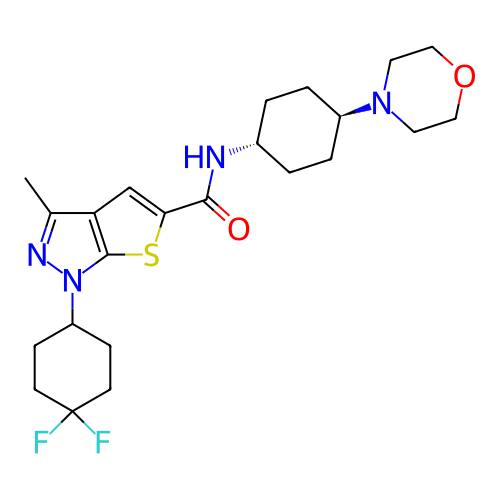
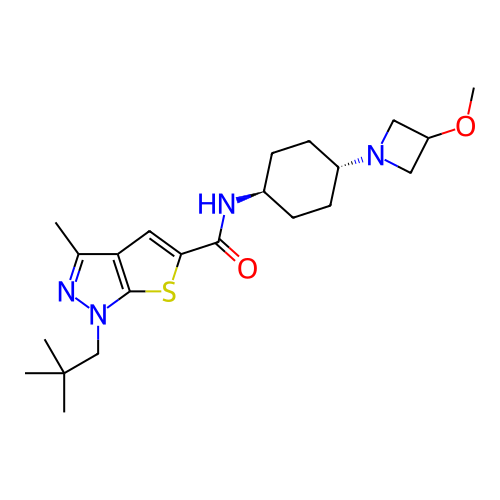
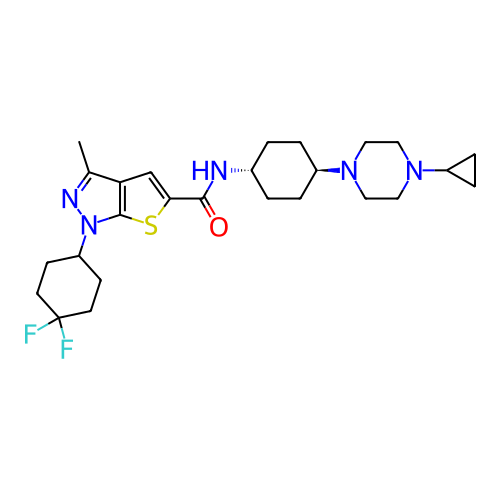Affinity DataIC50: 4.80nMpH: 8.0Assay Description:The assay method used was a scintillation proximity assay (SPA) (obtained from GE Healthcare, Product Code TRKQ7100), with [3H]-cGMP as the substrate...More data for this Ligand-Target Pair
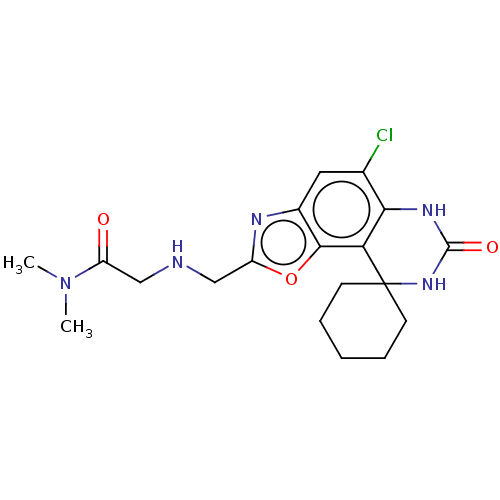
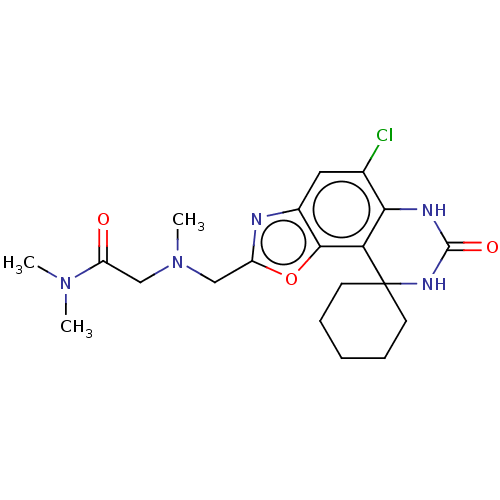
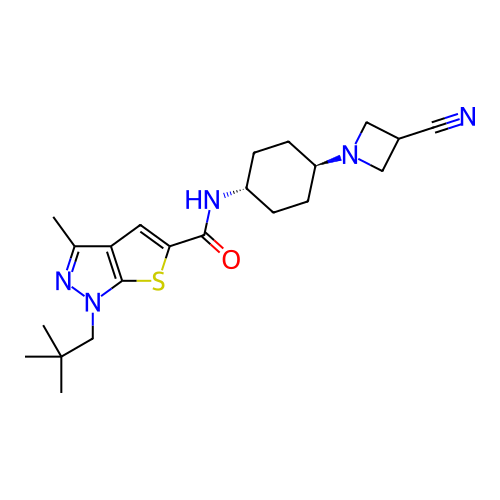
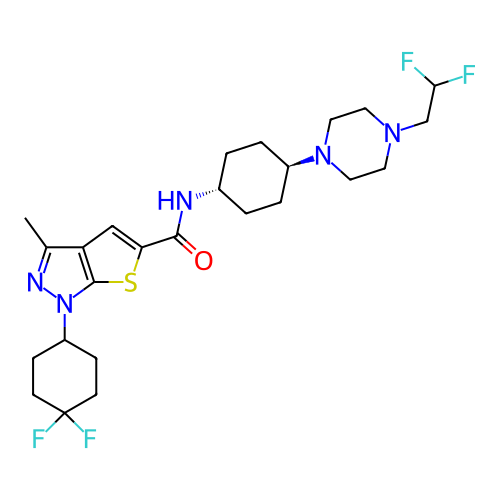
Affinity DataIC50: 8nMAssay Description:Displacement of [3H]-cAMP from human recombinant PDE7BMore data for this Ligand-Target Pair
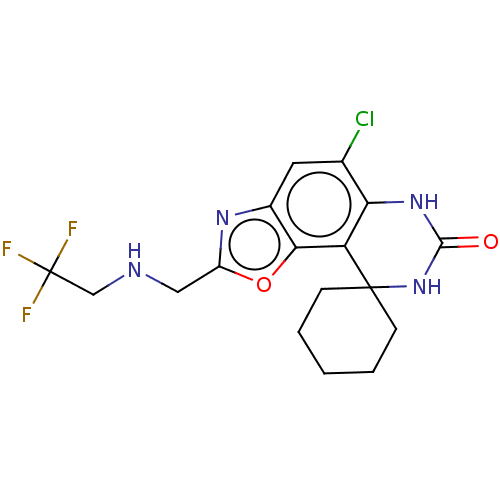
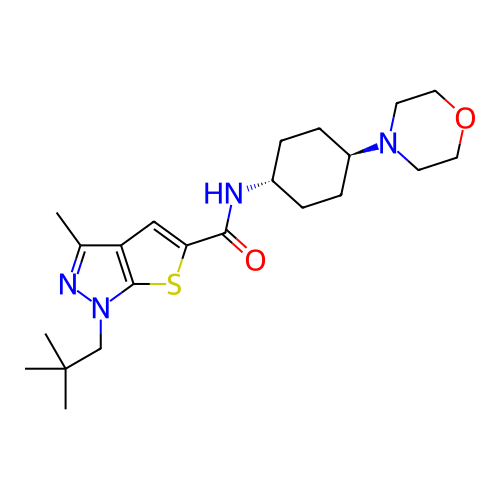
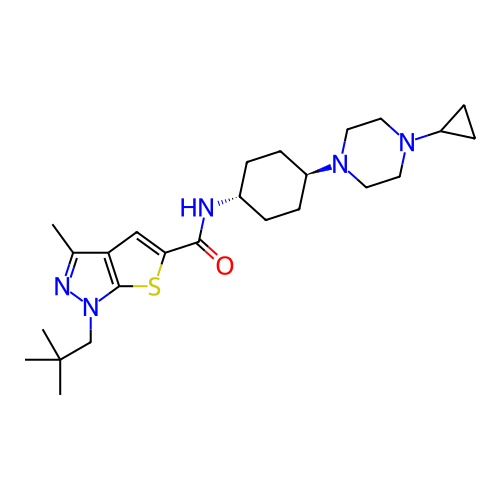
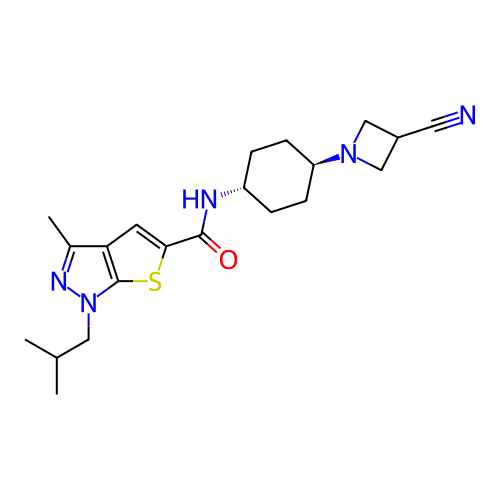
Affinity DataIC50: 9.27nMpH: 8.0Assay Description:The assay method used was a scintillation proximity assay (SPA) (obtained from GE Healthcare, Product Code TRKQ7100), with [3H]-cGMP as the substrate...More data for this Ligand-Target Pair
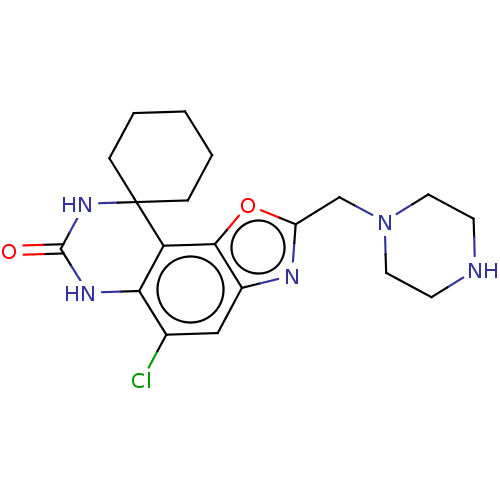
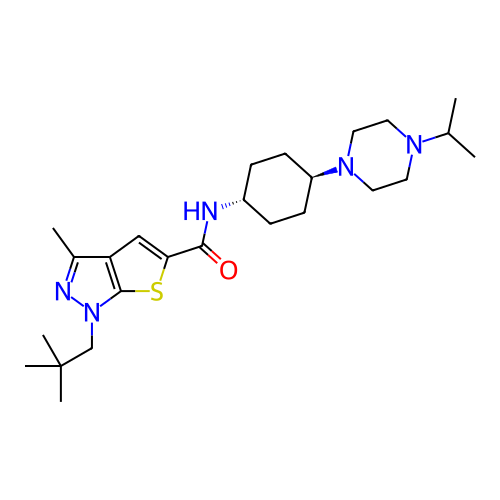
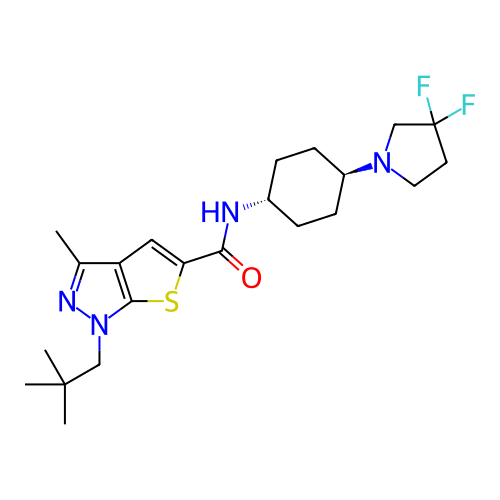
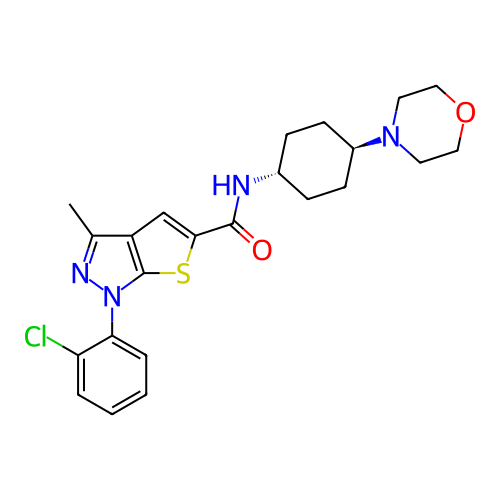
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:PDE7 inhibition was determined by an IMAP TR-FRET assay using PDE7B. The IMAP TR-FRET PDE assay was optimized for concentration of enzyme, FAM-cAMP s...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of cloned human recombinant PDE7B assessed as [3H]cAMP hydrolysis by radiometric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of PDE7B (unknown origin) by 3H-cAMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Displacement of [3H]-cAMP from human recombinant PDE7BMore data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Inhibition of cloned human recombinant PDE7B assessed as [3H]cAMP hydrolysis by radiometric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Compounds were tested for their activity by measuring the inhibition of PDE7A or PDE7B hydrolysis of [3H]cAMP to [3H]AMP. Generally, eight dilutions ...More data for this Ligand-Target Pair