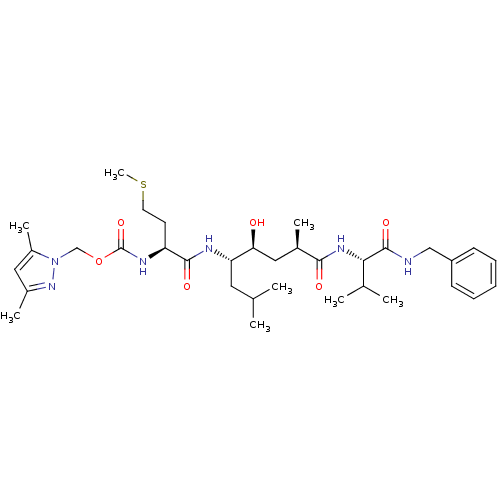
BDBM16251 (3,5-dimethyl-1H-pyrazol-1-yl)methyl N-[(1S)-1-{[(1R,3S,4S)-1-{[(1S)-1-(benzylcarbamoyl)-2-methylpropyl]carbamoyl}-3-hydroxy-1,6-dimethylheptan-4-yl]carbamoyl}-3-(methylsulfanyl)propyl]carbamate::N-benzyl-N2-{(2R,4S,5S)-5-[(N-{[(3,5-dimethyl-1H-pyrazol-1-yl)methoxy]carbonyl}-L-methionyl)amino]-4-hydroxy-2,7-dimethyloctanoyl}-L-valinamide::pyrazole-bearing inhibitor 3
SMILES CSCC[C@H](NC(=O)OCn1nc(C)cc1C)C(=O)N[C@@H](CC(C)C)[C@@H](O)C[C@@H](C)C(=O)N[C@@H](C(C)C)C(=O)NCc1ccccc1
InChI Key InChIKey=ULDYEMFEBUTYDX-UYVIBWFJSA-N
Data 3 KI
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 3 hits for monomerid = 16251
Found 3 hits for monomerid = 16251
Affinity DataKi: 14nM ΔG°: -11.1kcal/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 25nM ΔG°: -10.8kcal/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 811nM ΔG°: -8.64kcal/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair