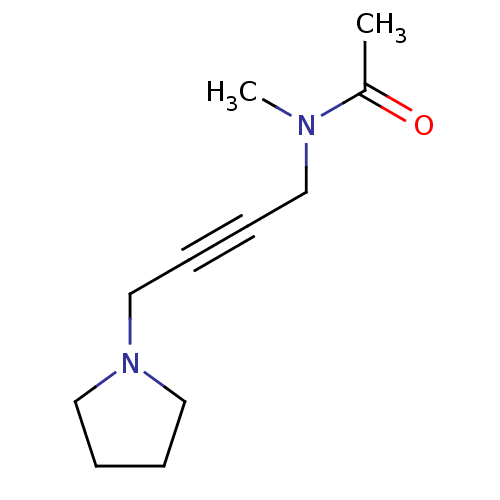
BDBM50009300 CHEMBL11081::N-Methyl-N-(4-pyrrolidin-1-yl-but-2-ynyl)-acetamide::UH5;N-Methyl-N-(4-pyrrolidin-1-yl-but-2-ynyl)-acetamide
SMILES CN(CC#CCN1CCCC1)C(C)=O
InChI Key InChIKey=HWYHDWGGACRVEH-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 9 hits for monomerid = 50009300
Found 9 hits for monomerid = 50009300
Affinity DataKi: 0.920nMAssay Description:Binding affinity to muscarinic acetylcholine receptor was determined in presence of [3H]OXO-M radioligand (in vitro)More data for this Ligand-Target Pair
Affinity DataKi: 260nMAssay Description:Binding affinity to muscarinic acetylcholine receptor was determined in presence of [3H]QNB radioligand (in vitro)More data for this Ligand-Target Pair
Affinity DataKi: 4.50E+3nMAssay Description:Muscarinic acetylcholine receptor binding affinity was determined using [3H]NMS for NG108-15 neuroblastoma cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4.96E+3nMAssay Description:Binding affinity was determined using [3H]-NMS for Muscarinic acetylcholine receptor in SK-N-SH neuroblastoma cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Inhibition of 0.1 nM [3H]cis-methyldioxolane binding to rat neocortex muscarinic receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 2.45E+3nMAssay Description:Inhibition of 0.03 nM [3H]quinuclidinyl benzylate binding to rat neocortex muscarinic receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 2.45E+3nMAssay Description:Displacement of [3H]-QNB in genetically transformed mouse cell line (m1C2) transfected with Muscarinic acetylcholine receptor M1More data for this Ligand-Target Pair
Affinity DataIC50: 2.01E+3nMAssay Description:Displacement of [3H]-QNB from Muscarinic acetylcholine receptor M2 from rat heart homogenatesMore data for this Ligand-Target Pair
Affinity DataKd: 2.20E+3nMAssay Description:Equilibrium dissociation constant muscarinic receptor complex, measured in guinea pig ileumMore data for this Ligand-Target Pair