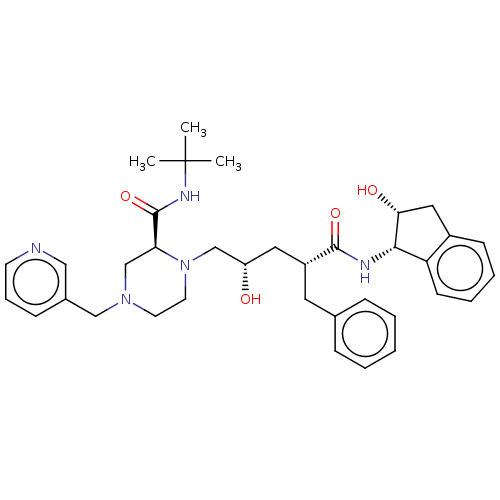
BDBM517 (2S)-1-[(2S,4R)-4-benzyl-2-hydroxy-4-{[(1S,2R)-2-hydroxy-2,3-dihydro-1H-inden-1-yl]carbamoyl}butyl]-N-tert-butyl-4-(pyridin-3-ylmethyl)piperazine-2-carboxamide::CHEMBL115::Crixivan::INDINAVIR SULFATE::Indinavir::Indinavir, 19::L-735, 524::MK639
SMILES CC(C)(C)NC(=O)[C@@H]1CN(Cc2cccnc2)CCN1C[C@@H](O)C[C@@H](Cc1ccccc1)C(=O)N[C@@H]1[C@H](O)Cc2ccccc12
InChI Key InChIKey=CBVCZFGXHXORBI-PXQQMZJSSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 4 hits for monomerid = 517
Found 4 hits for monomerid = 517
TargetDimer of Gag-Pol polyprotein [489-587,Q496K](Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
Affinity DataKi: 0.543nM ΔG°: -12.6kcal/molepH: 5.0 T: 2°CAssay Description:Protease activity was determined by following the increase in fluorescence with hydrolysis of the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Ty...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587,Q496K,I573V](Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
Affinity DataKi: 4.40nM ΔG°: -11.4kcal/molepH: 5.0 T: 2°CAssay Description:Protease activity was determined by following the increase in fluorescence with hydrolysis of the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Ty...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587,Q496K,L499I,M525I,S526D,M535I,R546K,L552P,A560V,G562S,L579M,I582L](Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
Affinity DataKi: 481nM ΔG°: -8.61kcal/molepH: 5.0 T: 2°CAssay Description:Protease activity was determined by following the increase in fluorescence with hydrolysis of the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Ty...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587,Q496K,L499I,M525I,S526D,M535I,R546K,L552P,A560V,G562S,I573V,L579M,I582L](Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
Affinity DataKi: 1.10E+3nM ΔG°: -8.12kcal/molepH: 5.0 T: 2°CAssay Description:Protease activity was determined by following the increase in fluorescence with hydrolysis of the fluorogenic substrate Arg-Glu(EDANS)-Ser-Gln-Asn-Ty...More data for this Ligand-Target Pair
Activity Spreadsheet -- ITC Data from BindingDB
 Found 18 hits for monomerid = 517
Found 18 hits for monomerid = 517
CellHIV-1 Protease Mutant (V82F/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -9.19kcal/mole −TΔS°: -15.1kcal/mole ΔH°: 5.89kcal/mole
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -11.8kcal/mole −TΔS°: -15.7kcal/mole ΔH°: 3.89kcal/mole
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (M46I/I54V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -12.2kcal/mole −TΔS°: -15.7kcal/mole ΔH°: 3.49kcal/mole logk: 8.90E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.7kcal/mole −TΔS°: -14.8kcal/mole ΔH°: 2.10kcal/mole logk: 2.08E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.1kcal/mole −TΔS°: -14.5kcal/mole ΔH°: 2.40kcal/mole logk: 7.09E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -11.6kcal/mole −TΔS°: -14.2kcal/mole ΔH°: 2.60kcal/mole logk: 3.13E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease B Subtype Mutant (V82F/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -10.2kcal/mole −TΔS°: -13.7kcal/mole ΔH°: 3.49kcal/mole logk: 3.13E+7
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.7kcal/mole −TΔS°: -14.8kcal/mole ΔH°: 2.10kcal/mole logk: 2.08E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -9.28kcal/mole −TΔS°: -17.7kcal/mole ΔH°: 8.39kcal/mole logk: 6.50E+6
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -10.4kcal/mole −TΔS°: -16.8kcal/mole ΔH°: 6.39kcal/mole logk: 4.50E+7
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -12.4kcal/mole −TΔS°: -14.2kcal/mole ΔH°: 1.80kcal/mole logk: 1.30E+9
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (V82A/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -10.8kcal/mole −TΔS°: -14.5kcal/mole ΔH°: 3.69kcal/mole logk: 8.50E+7
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease Mutant (L10I/L90M)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -11.8kcal/mole −TΔS°: -14.8kcal/mole ΔH°: 3.00kcal/mole logk: 4.30E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease C Subtype Mutant (V82F/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -9.48kcal/mole −TΔS°: -12.6kcal/mole ΔH°: 3.09kcal/mole logk: 1.00E+7
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -8.54kcal/mole −TΔS°: -14.7kcal/mole ΔH°: 6.15kcal/mole logk: 1.86E+6
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -7.79kcal/mole −TΔS°: -13.4kcal/mole ΔH°: 5.59kcal/mole logk: 5.26E+5
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
ITC DataΔG°: -11.9kcal/mole −TΔS°: -15.5kcal/mole ΔH°: 3.59kcal/mole logk: 5.56E+8
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
CellHIV-1 Protease A Subtype Mutant (V82F/I84V)(Human immunodeficiency virus type 1)
The Johns Hopkins University
The Johns Hopkins University
ITC DataΔG°: -9.19kcal/mole −TΔS°: -12.1kcal/mole ΔH°: 2.90kcal/mole logk: 5.20E+6
pH: 5.0 T: 25.00°C
pH: 5.0 T: 25.00°C
 3D Structure (crystal)
3D Structure (crystal)