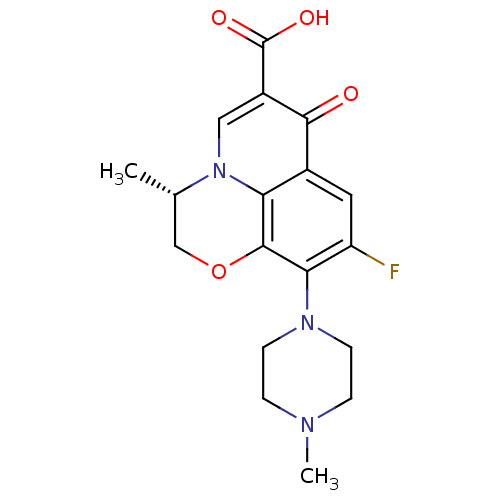
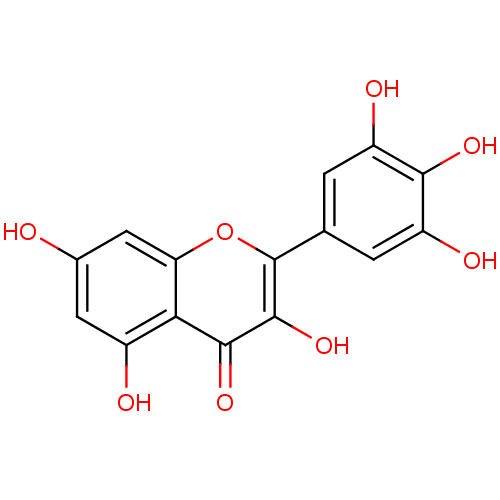
Report error Found 20 Enz. Inhib. hit(s) with Target = 'DNA primase'
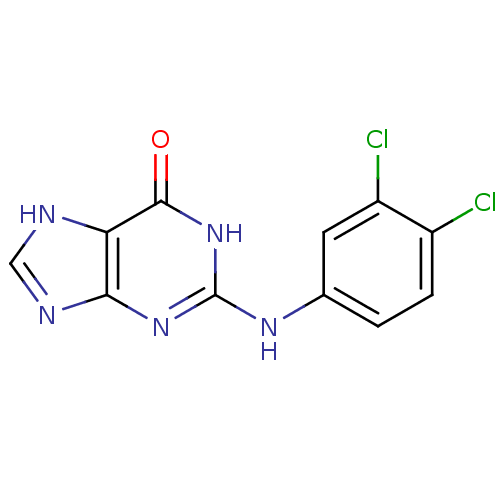
Affinity DataIC50: 770nMAssay Description:Inhibition against DNA-Dependent GTPase activity of HSV-1 helicase-primase in HSV-1-infected CV-1 cellsMore data for this Ligand-Target Pair
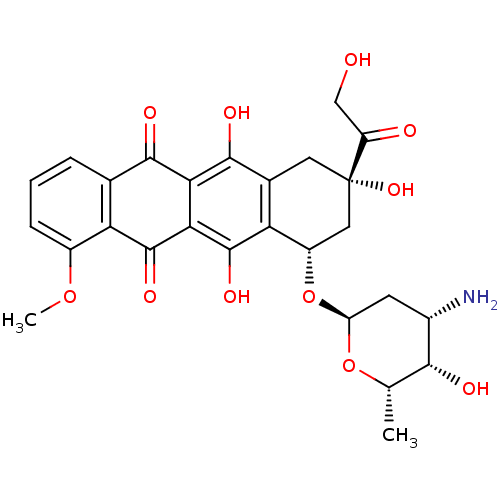
Affinity DataIC50: 850nMAssay Description:Inhibition against DNA-Dependent GTPase activity of HSV-1 helicase-primase in HSV-1-infected CV-1 cellsMore data for this Ligand-Target Pair
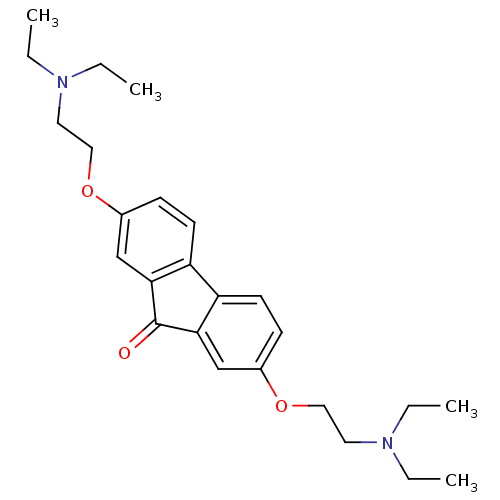
Affinity DataIC50: 850nMAssay Description:Inhibition against DNA-Dependent GTPase activity of HSV-1 helicase-primase in HSV-1-infected CV-1 cellsMore data for this Ligand-Target Pair
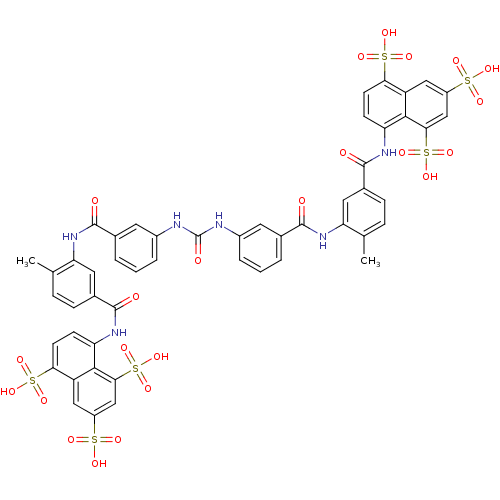
Affinity DataIC50: 880nMAssay Description:Inhibition against DNA-Dependent GTPase activity of HSV-1 helicase-primase in HSV-1-infected CV-1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 910nMAssay Description:Inhibition against DNA-Dependent GTPase activity of HSV-1 helicase-primase in HSV-1-infected CV-1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 920nMAssay Description:Inhibition against DNA-Dependent GTPase activity of HSV-1 helicase-primase in HSV-1-infected CV-1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition against DNA-Dependent GTPase activity of HSV-1 helicase-primase in HSV-1-infected CV-1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition against DNA-Dependent GTPase activity of HSV-1 helicase-primase in HSV-1-infected CV-1 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.40E+3nMpH: 8.8Assay Description:The reaction mixture contained DNA (1.25 uM or as specified), NTP (110 uM or as specified), 50 mM NaCl, 150 mM potassium glutamate, buffer [20 mM CAP...More data for this Ligand-Target Pair
Affinity DataIC50: 7.10E+3nMpH: 8.8Assay Description:The reaction mixture contained DNA (1.25 uM or as specified), NTP (110 uM or as specified), 50 mM NaCl, 150 mM potassium glutamate, buffer [20 mM CAP...More data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+3nMpH: 8.8Assay Description:The reaction mixture contained DNA (1.25 uM or as specified), NTP (110 uM or as specified), 50 mM NaCl, 150 mM potassium glutamate, buffer [20 mM CAP...More data for this Ligand-Target Pair
Affinity DataIC50: 8.70E+3nMAssay Description:Inhibition against Primase by coupled primase-DNA polymerase-I assay with the two subunit recombinant HSV-1 helicase primaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+3nMpH: 8.8Assay Description:The reaction mixture contained DNA (1.25 uM or as specified), NTP (110 uM or as specified), 50 mM NaCl, 150 mM potassium glutamate, buffer [20 mM CAP...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition against Primase by coupled primase-DNA polymerase-I assay with the two subunit recombinant HSV-1 helicase primaseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.90E+4nMAssay Description:Inhibition against Primase by coupled primase-DNA polymerase-I assay with the two subunit recombinant HSV-1 helicase primaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.90E+4nMpH: 8.8Assay Description:The reaction mixture contained DNA (1.25 uM or as specified), NTP (110 uM or as specified), 50 mM NaCl, 150 mM potassium glutamate, buffer [20 mM CAP...More data for this Ligand-Target Pair
Affinity DataIC50: 7.50E+4nMpH: 8.8Assay Description:The reaction mixture contained DNA (1.25 uM or as specified), NTP (110 uM or as specified), 50 mM NaCl, 150 mM potassium glutamate, buffer [20 mM CAP...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+4nMAssay Description:Inhibitory concentration against herpes simplex virus helicase primase 8 (HSV)More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 8.8Assay Description:The reaction mixture contained DNA (1.25 uM or as specified), NTP (110 uM or as specified), 50 mM NaCl, 150 mM potassium glutamate, buffer [20 mM CAP...More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+5nMAssay Description:Inhibition of Escherichia coli primaseMore data for this Ligand-Target Pair