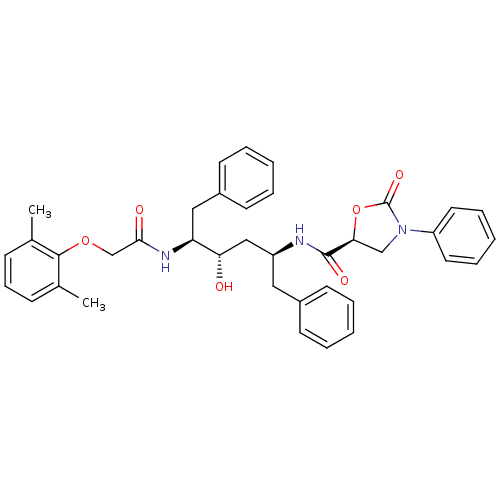
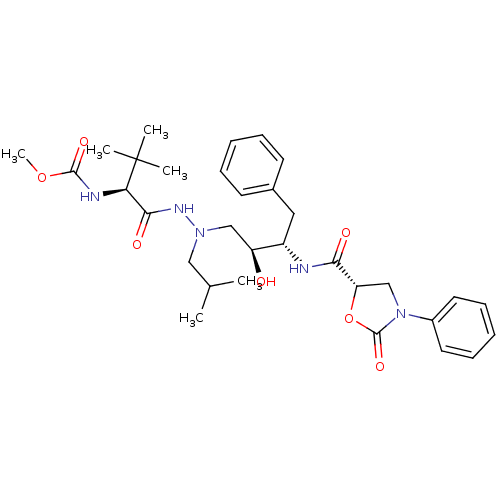
Report error Found 19 Enz. Inhib. hit(s) with Target = 'Dimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D]'
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
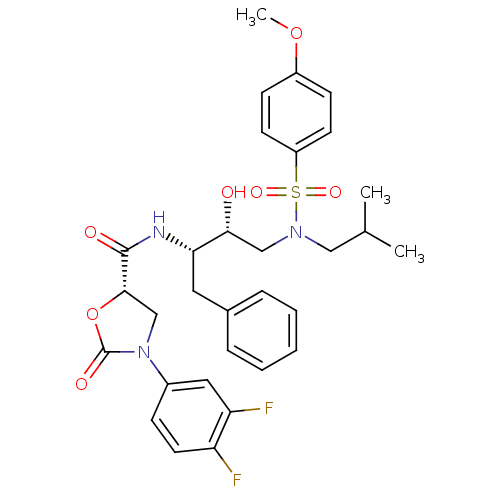
Affinity DataKi: 0.00900nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
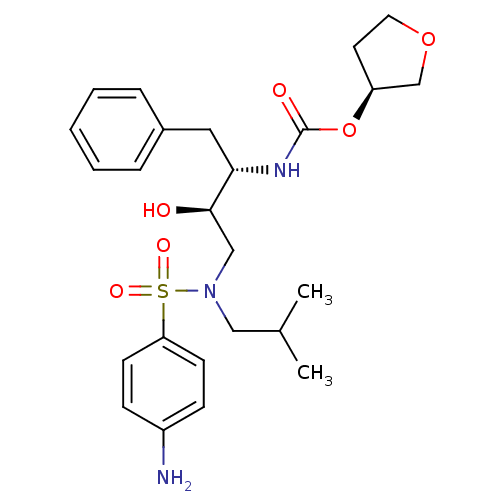
Affinity DataKi: 0.0390nM ΔG°: -60.4kJ/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
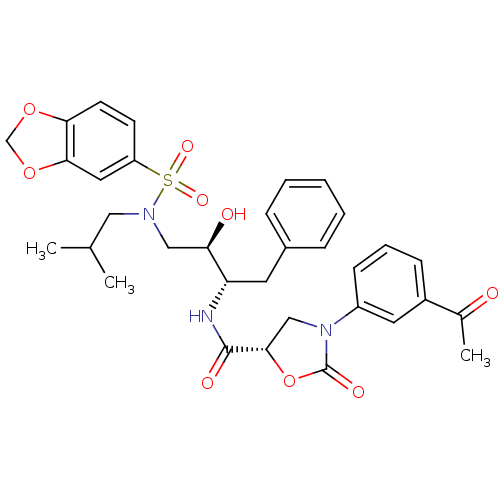
Affinity DataKi: 0.0400nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
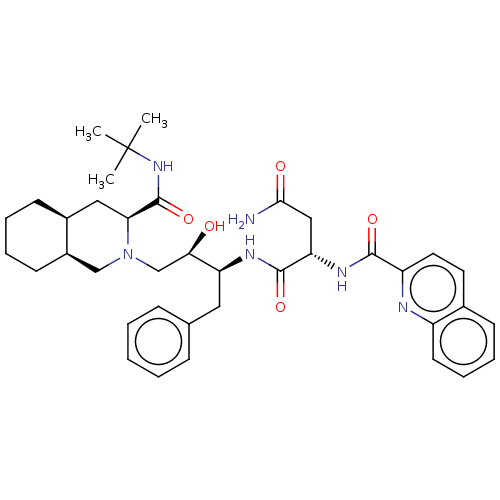
Affinity DataKi: 0.0800nM ΔG°: -58.6kJ/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.152nM ΔG°: -57.0kJ/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.179nM ΔG°: -56.6kJ/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.210nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.235nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.360nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.460nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.920nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 1.03nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 1.67nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 3.10nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 3.90nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 13.9nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 14nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 18.3nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 39.2nMAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair