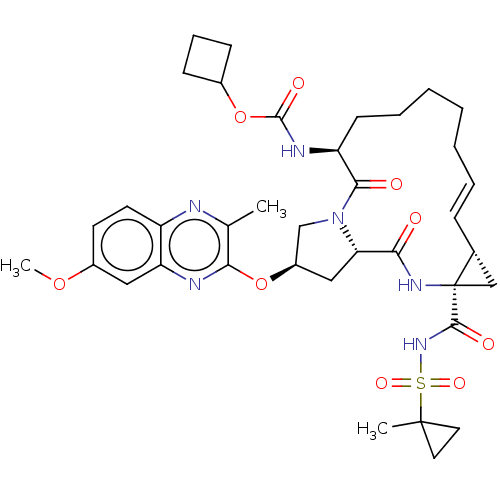
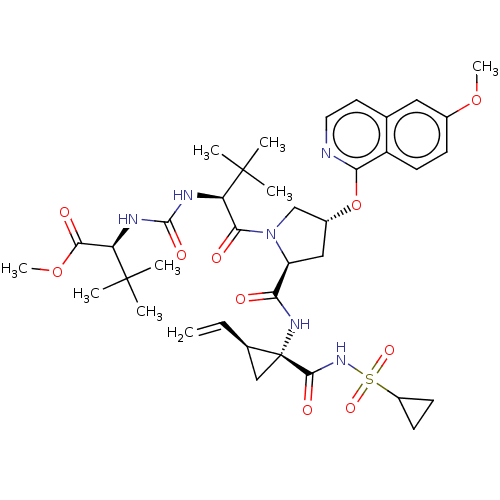
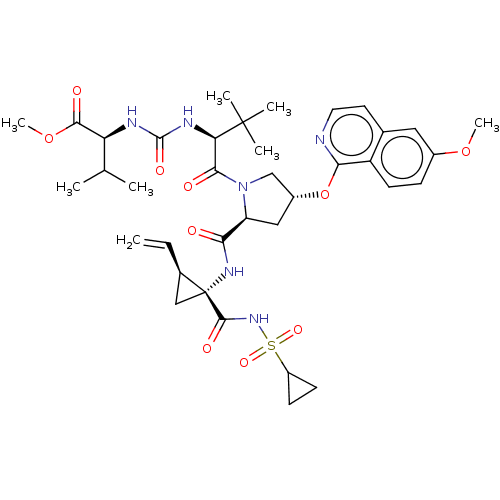
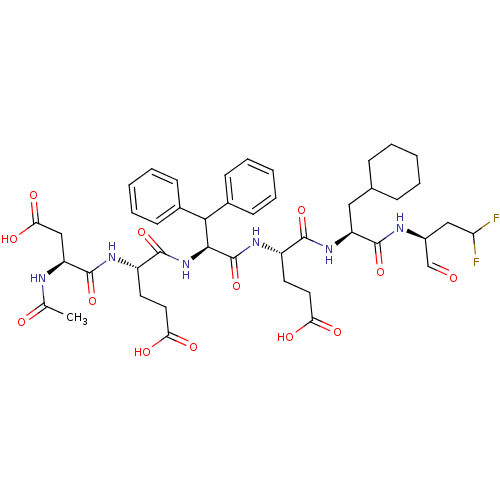
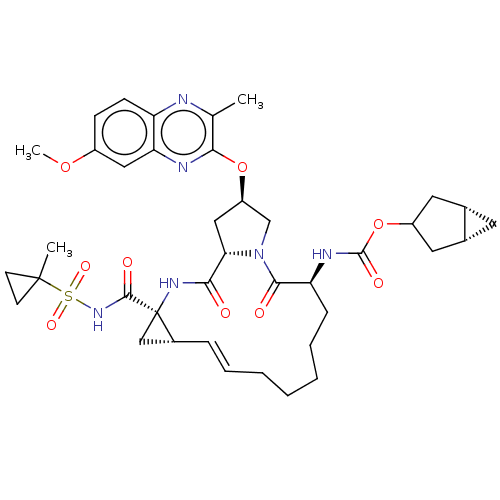
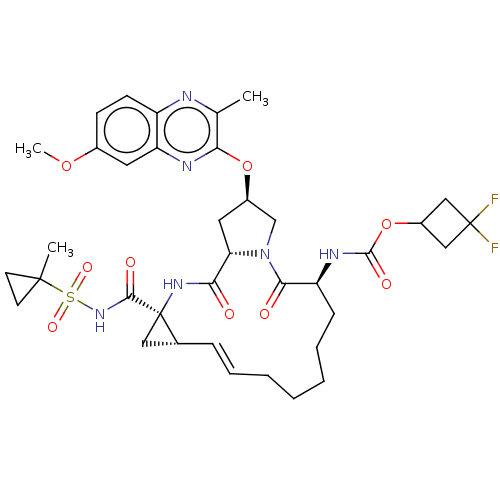
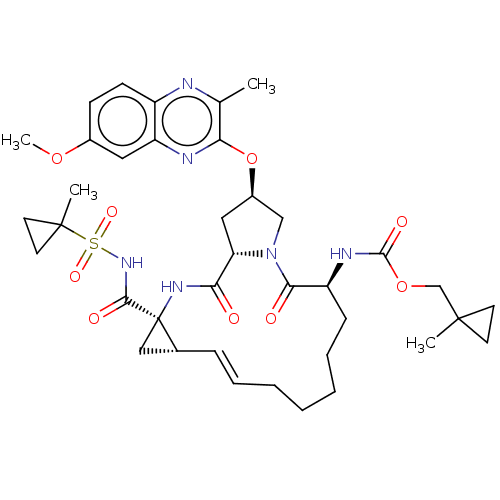
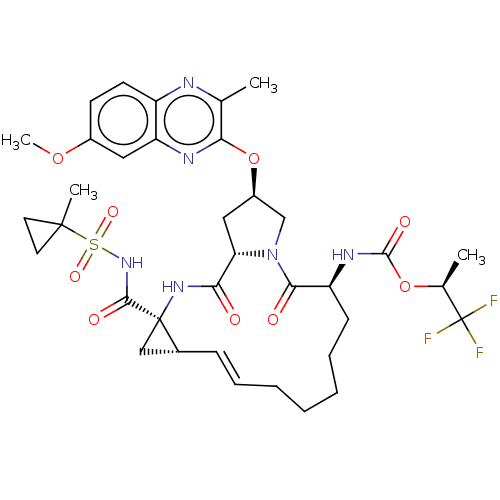
Report error Found 552 Enz. Inhib. hit(s) with Target = 'Genome polyprotein/Non-structural protein 4A'
Affinity DataKi: 0.0100nMAssay Description:Inhibitory activity against hepatitis C virus (HCV) NS3/NS4A serine proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:Inhibition of hepatitis C virus (HCV) NS3/NS4A serine proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataIC50: 0.190nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:Inhibition of HCV NS3/4a protease using Ac-DE-Dap(QXL520)-EE-Abu-shi-[COO]AS-C(5-FAMsp)-NH2 as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Inhibition of wild type HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) using Ac-DE-Dap(QXL 520)-EE-Abu-psi-[COO]AS-C(5-FAMsp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.210nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 0.220nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 0.240nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 0.270nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataIC50: 0.340nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 0.440nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 0.460nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Inhibition of hepatitis C virus (HCV) NS3/NS4A serine proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Inhibition of HCV GT-3a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Inhibitory activity against hepatitis C virus (HCV) NS3/NS4A serine proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease R155K mutant expressed in Escherichia coli BL21(DE3) using Ac-DE-Dap(QXL 520)-EE-Abu-psi-[COO]AS-C(5-FA...More data for this Ligand-Target Pair
Affinity DataIC50: 0.820nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of recombinant full length HCV genotype 1a NS3/4A protease (1027 to 1711 residues) expressed in Escherichia coli strain BL21 (DE3) using R...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of recombinant full length HCV genotype 1a NS3/4A protease (1027 to 1711 residues) expressed in Escherichia coli strain BL21 (DE3) using R...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibition of hepatitis C virus (HCV) NS3/NS4A serine proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition of N-terminal poly-His tagged recombinant HCV genotype 1a NS3/4A protease expressed in Escherichia coli BL21(DE3) pLysS using HCV-FRET pep...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair
Affinity DataKi: 1.90nMAssay Description:Inhibition of HCV GT-1a NS3/4a protease using Ac-DE-D(Edans)-EE-Abu-c-[COO]-AS-K(Dabcy1)-NH2 preincubated for 1 hr followed by substrate additionMore data for this Ligand-Target Pair

 3D Structure (crystal)
3D Structure (crystal)