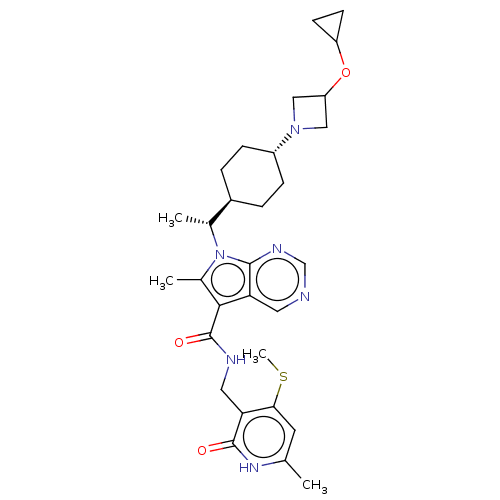
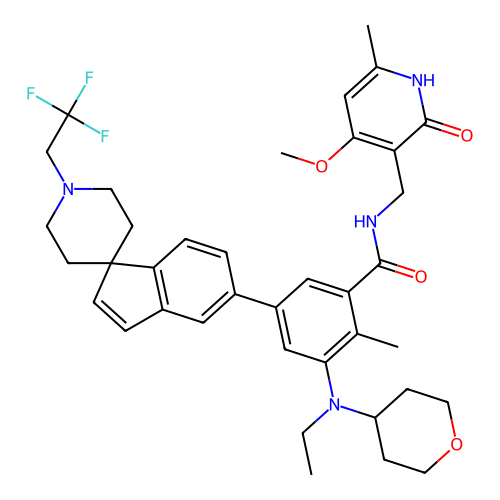
Report error Found 6479 Enz. Inhib. hit(s) with Target = 'Histone-lysine N-methyltransferase EZH2'
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.00970nMAssay Description:Inhibition of wild-type EZH2 (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.0100nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.0100nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0100nMAssay Description:Inhibition of EZH2 (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKd: 0.0159nMAssay Description:Inhibition of recombinant PRC2 complex (unknown origin) assessed as dissociation rate constant using H3K27me0 peptide substrate incubated for 3 hrs b...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.0200nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKd: 0.0203nMAssay Description:Inhibition of recombinant PRC2 complex (unknown origin) assessed as dissociation rate constant using H3K27me0 peptide substrate incubated for 3 hrs b...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0280nMAssay Description:Inhibition of EZH2 (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0320nMAssay Description:Inhibition of EZH2 (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0370nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0380nMAssay Description:Inhibition of EZH2 (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0380nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0390nMAssay Description:Inhibition of EZH2 (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0390nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.0400nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0430nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKd: 0.0430nMAssay Description:Inhibition of recombinant PRC2 complex (unknown origin) assessed as dissociation rate constant using H3K27me0 peptide substrate incubated for 3 hrs b...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0540nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0540nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0570nMAssay Description:Inhibition of EZH2 (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0570nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0590nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0630nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0670nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKd: 0.0693nMAssay Description:Inhibition of recombinant PRC2 complex (unknown origin) assessed as dissociation rate constant using H3K27me0 peptide substrate incubated for 3 hrs b...More data for this Ligand-Target Pair
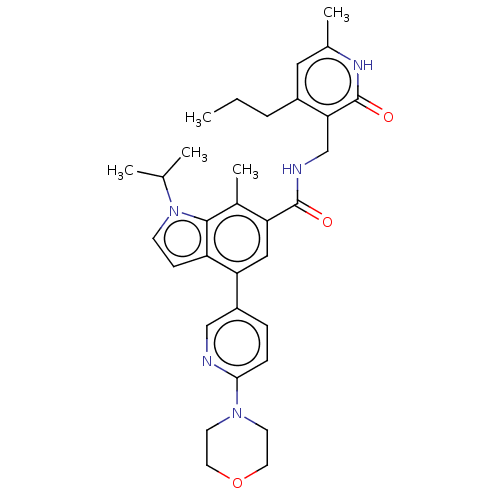
TargetHistone-lysine N-methyltransferase EZH2 [Y641N](Human)
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
Ligand Info
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0720nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2 [Y641N](Human)
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
TargetHistone-lysine N-methyltransferase EZH2 [Y641N](Human)
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
Ligand Info
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0840nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
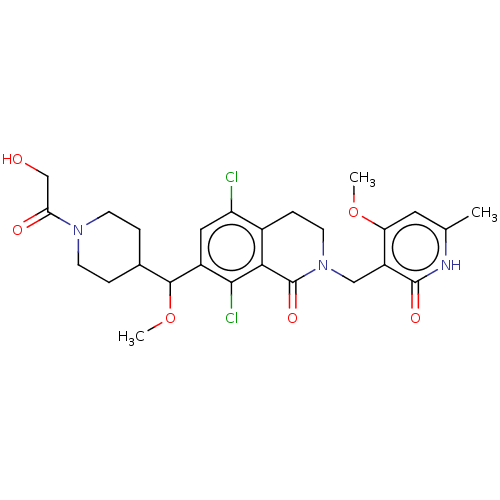
TargetHistone-lysine N-methyltransferase EZH2 [Y641N](Human)
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
Ligand Info
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0910nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKd: 0.0916nMAssay Description:Inhibition of recombinant PRC2 complex (unknown origin) assessed as dissociation rate constant using H3K27me0 peptide substrate incubated for 3 hrs b...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0920nMAssay Description:EZH2 biochemical assay (IC50): Compound potencies were assessed through incorporation of 3H-SAM into a biotinylated H3 peptide. Specifically, 30 pM P...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.0920nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKd: 0.0935nMAssay Description:Inhibition of recombinant PRC2 complex (unknown origin) assessed as dissociation rate constant using H3K27me0 peptide substrate incubated for 3 hrs b...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
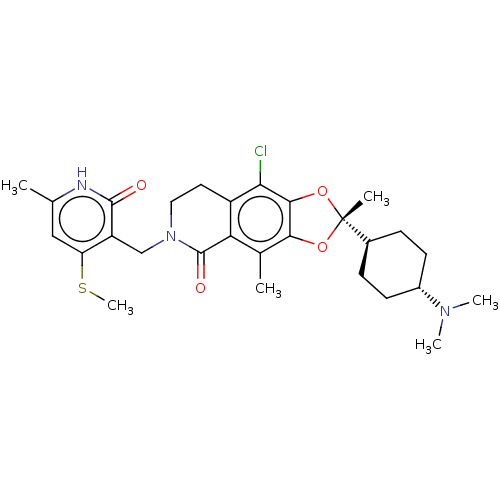
TargetHistone-lysine N-methyltransferase EZH2 [Y641N](Human)
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
Ligand Info
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: <0.100nMAssay Description:Binding affinity to EZH2 Y641N mutant (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:A. Compound preparation1. Prepare 10 mM stock solutions in 100% DMSO from solid material2. Serial dilute 10 mM compound stocks either 2 or 3-fold in ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2/Polycomb protein EED/Polycomb protein SUZ12(Human)
Novartis
US Patent
Novartis
US Patent
Affinity DataIC50: 0.100nMAssay Description:Representative compounds of the present invention were serially and separately diluted 3-fold in DMSO to obtain a total of eight or twelve concentrat...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: <0.100nMAssay Description:Inhibition of EZH2 Y641N mutant (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of N-terminal FLAG-tagged EZH2 in PRC2 complex (unknown origin) expressed in baculovirus infected Sf9 cells using H2N-RKQLATKAAR(Kme0)SAPA...More data for this Ligand-Target Pair
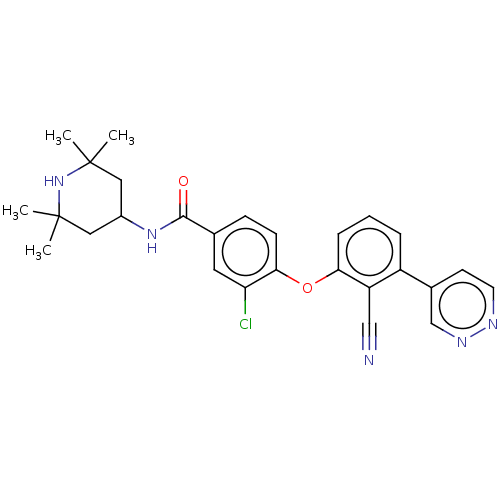
TargetHistone-lysine N-methyltransferase EZH2 [Y641N](Human)
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
SHANGHAI SYNERGY PHARMACEUTICAL SCIENCES CO., LTD.; ZHEJIANG HUAHAI PHARMACEUTICAL CO., LTD.
US Patent
Ligand Info
TargetHistone-lysine N-methyltransferase EZH2(Human)
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affiliated Hospital of Guangdong Medical University
Curated by ChEMBL
Affinity DataKi: <0.100nMAssay Description:Binding affinity to EZH2 (unknown origin)More data for this Ligand-Target Pair
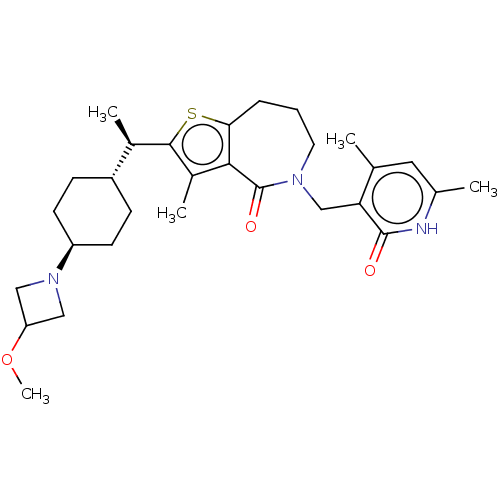
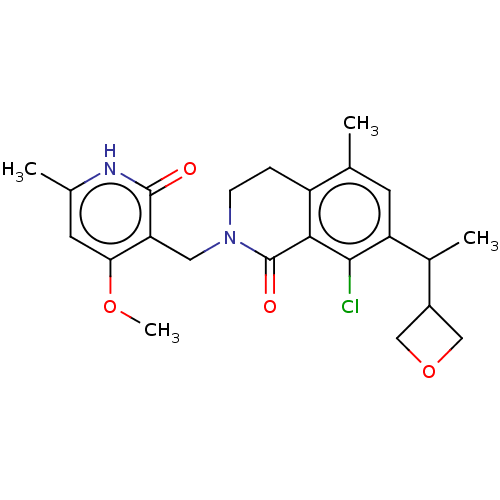



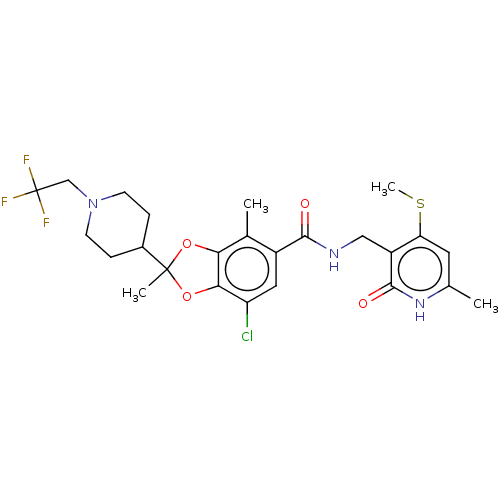
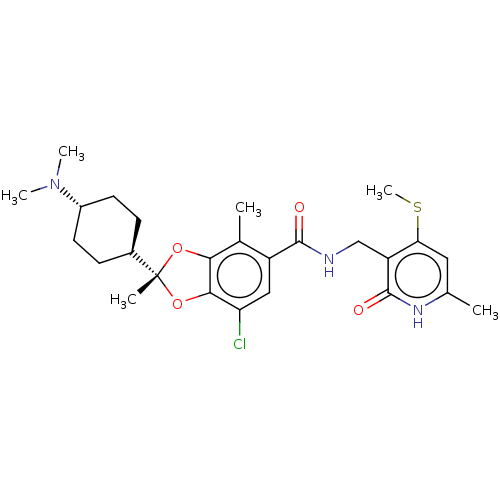

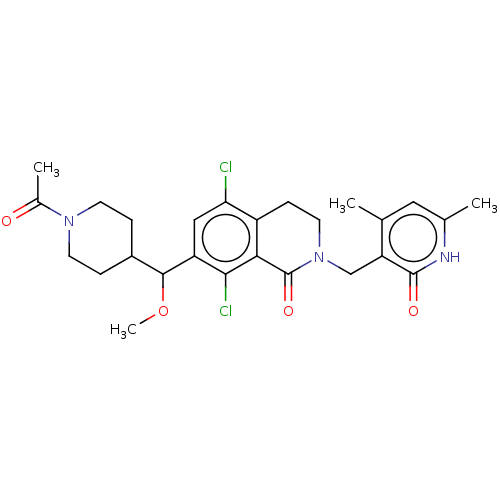
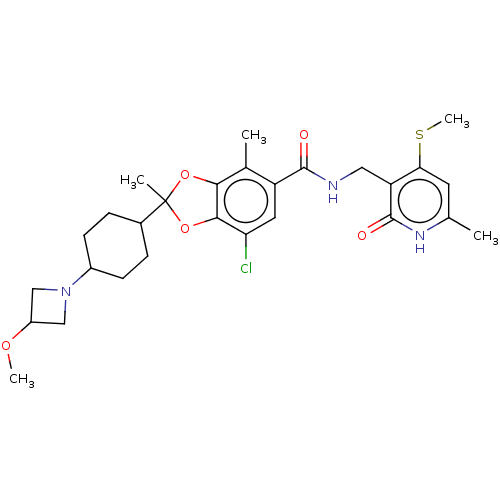
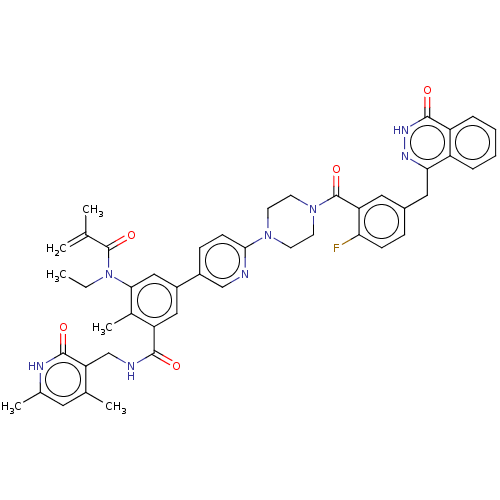
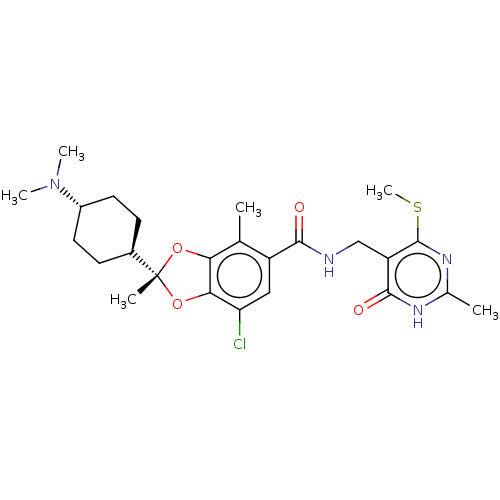
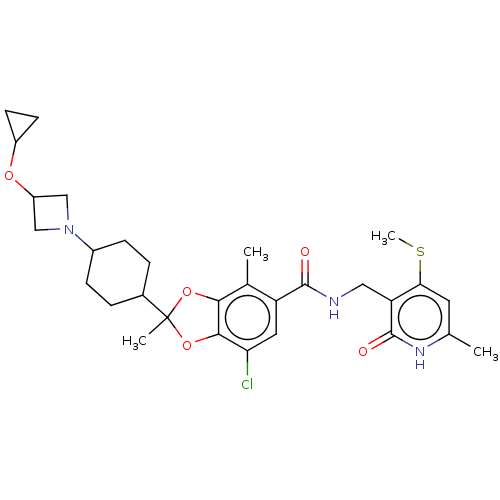
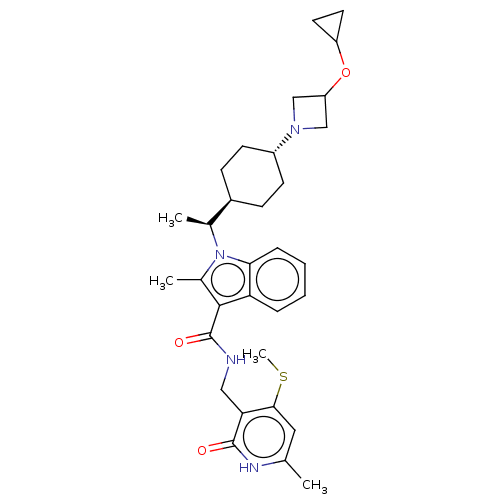



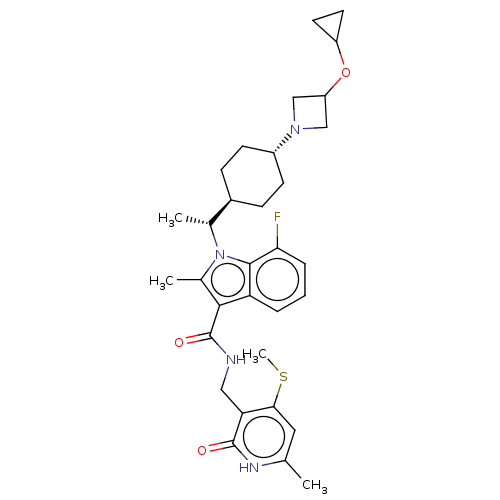

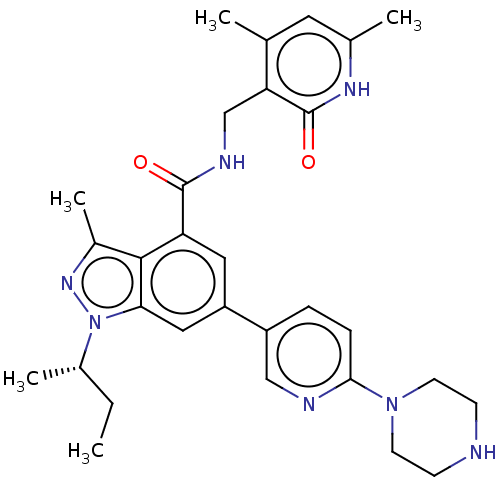
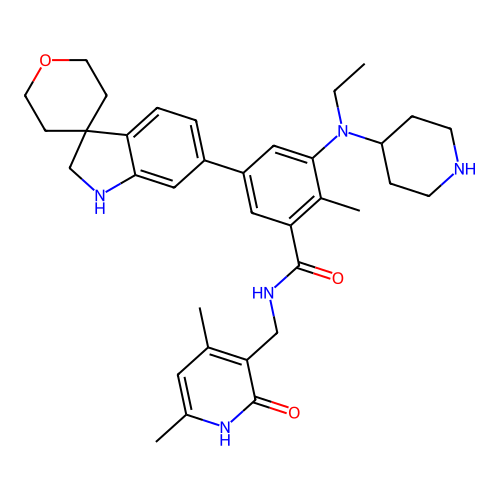

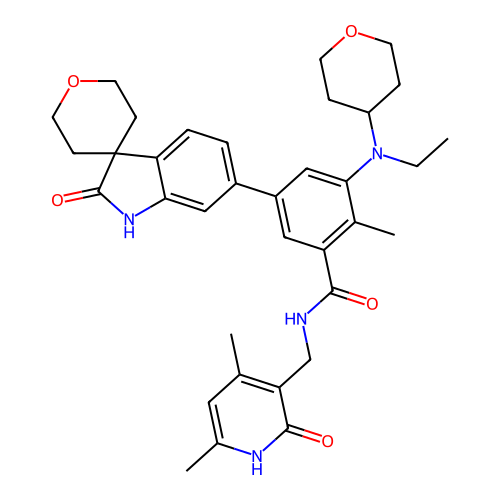
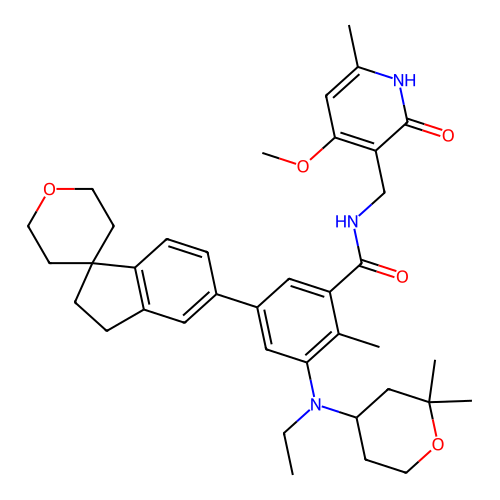
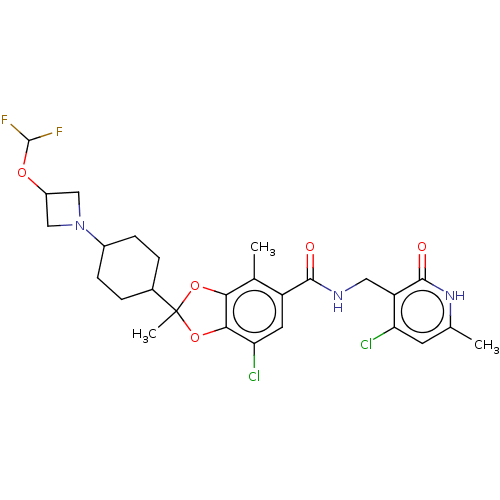
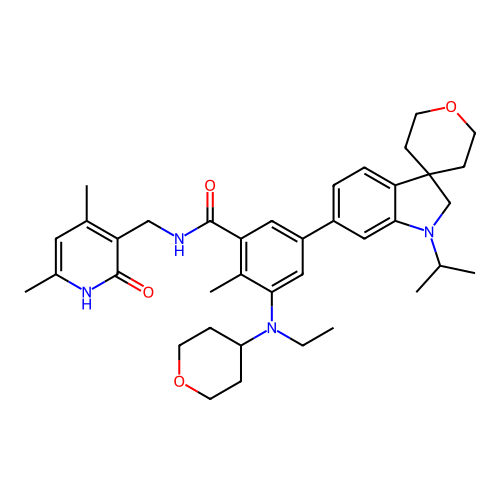
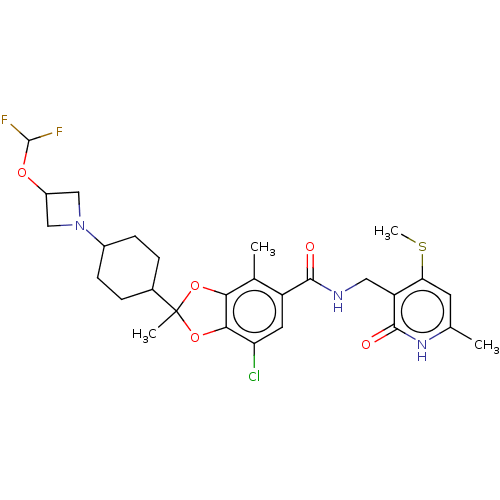

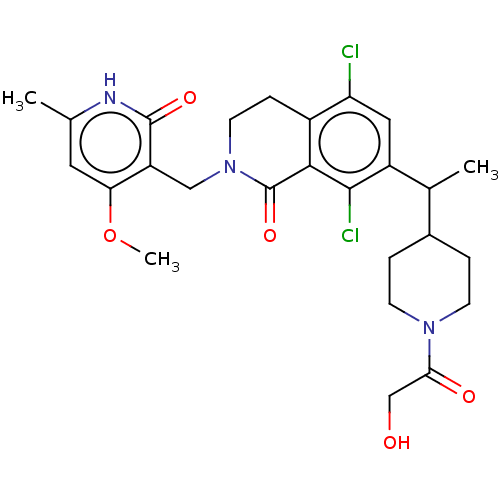
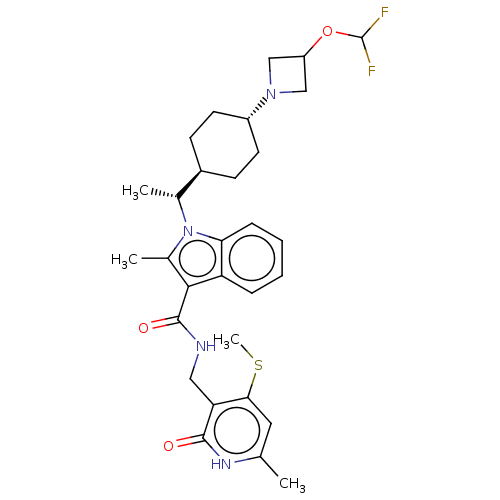
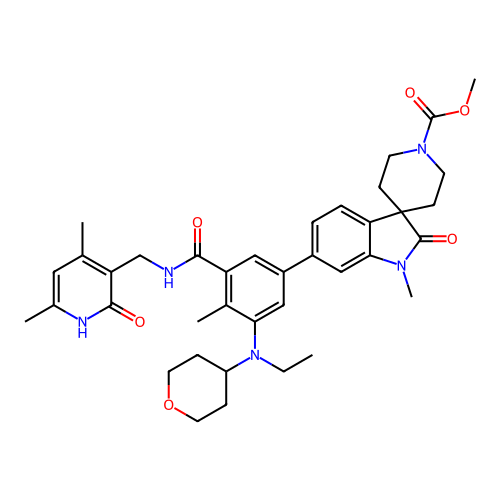
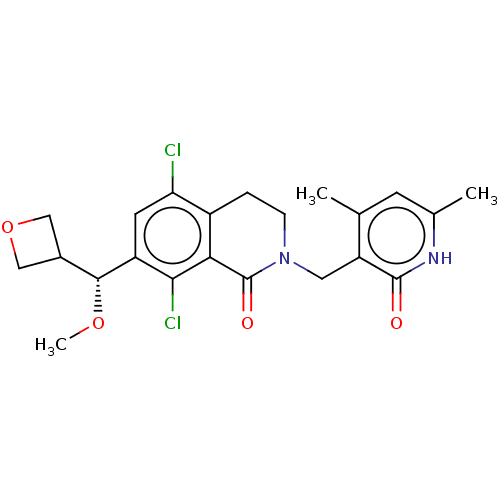

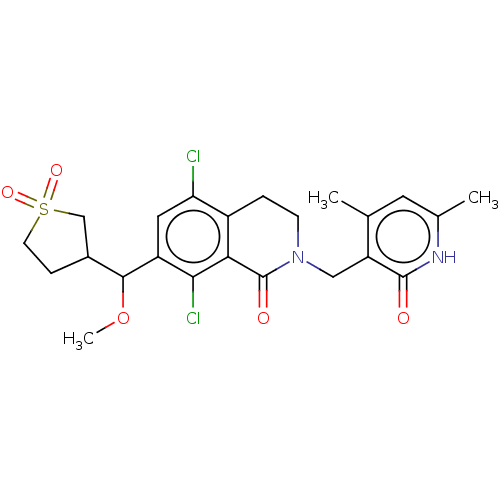
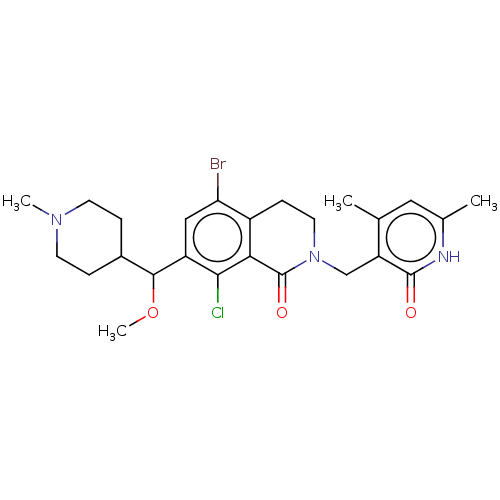

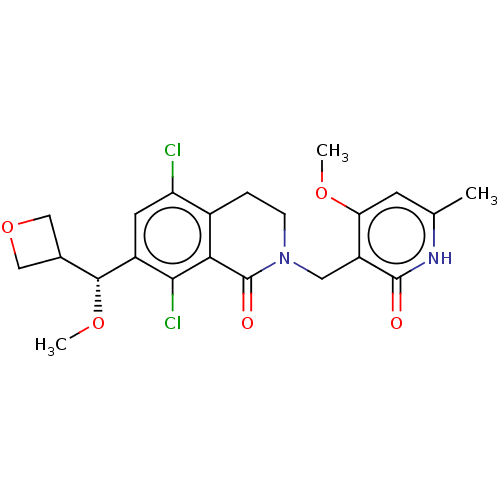
 3D Structure (crystal)
3D Structure (crystal)