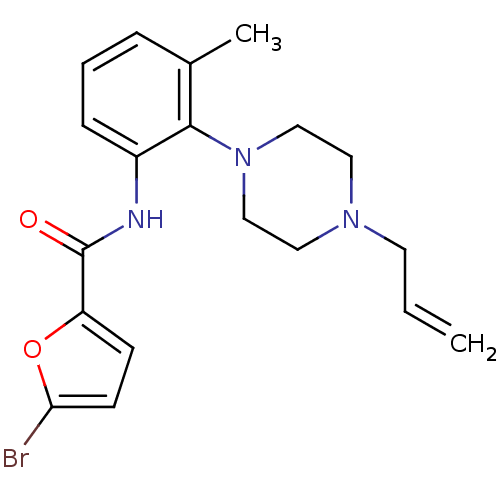
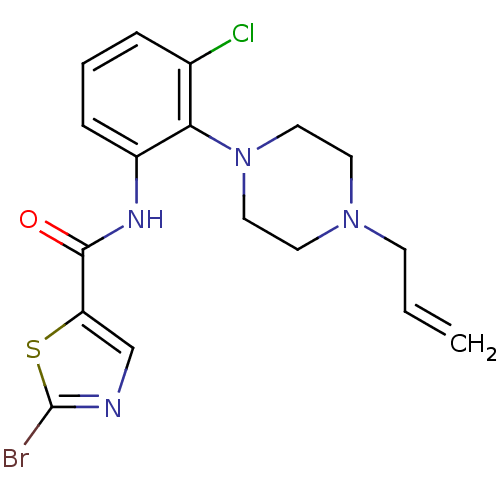
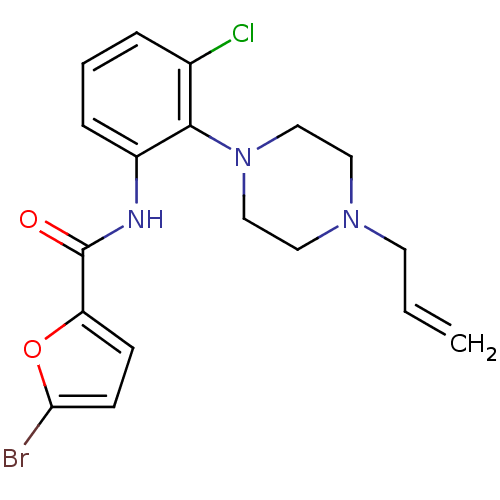
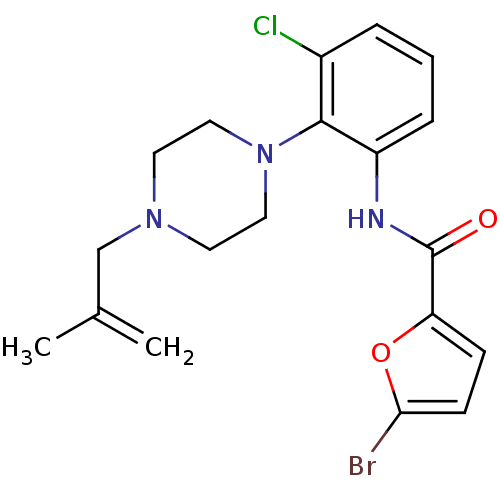
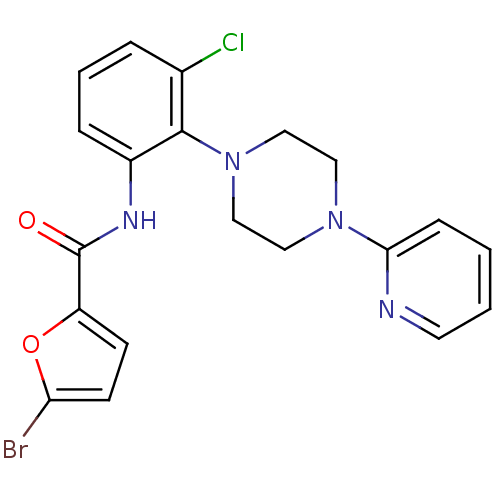
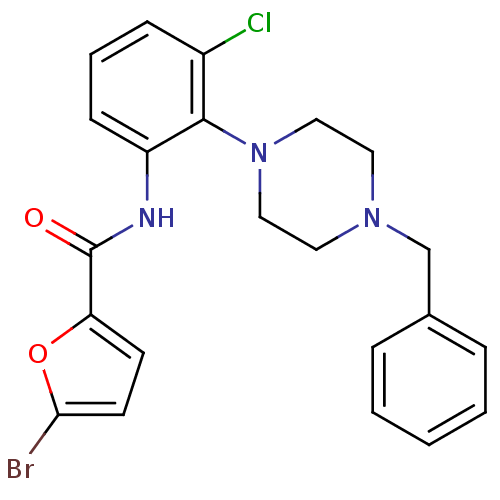
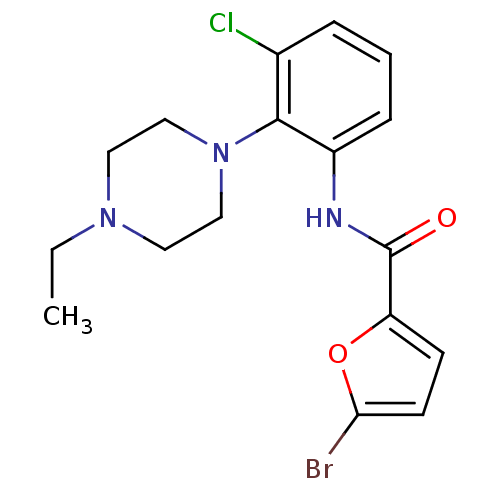
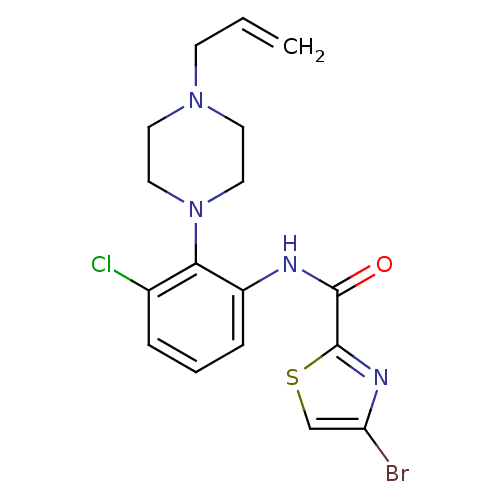
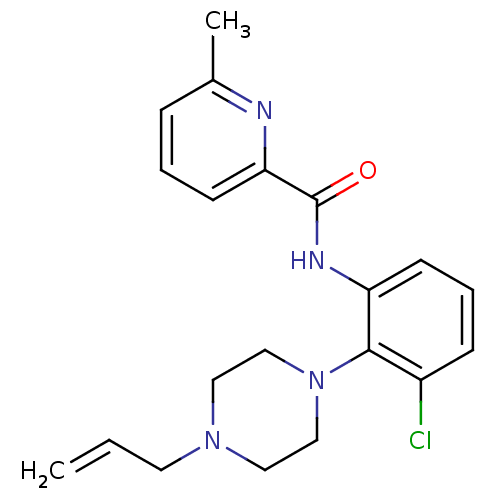
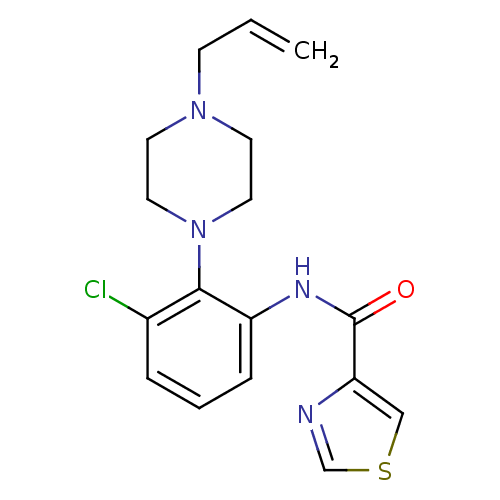
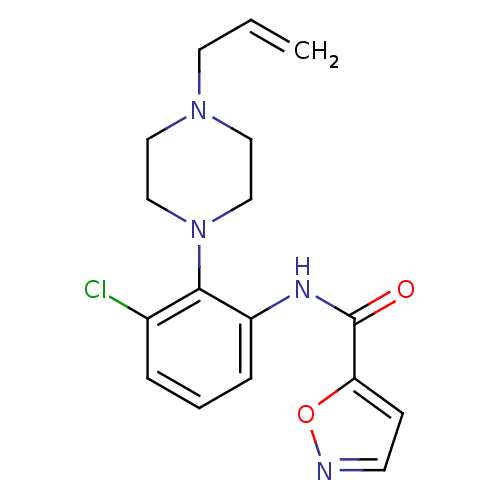
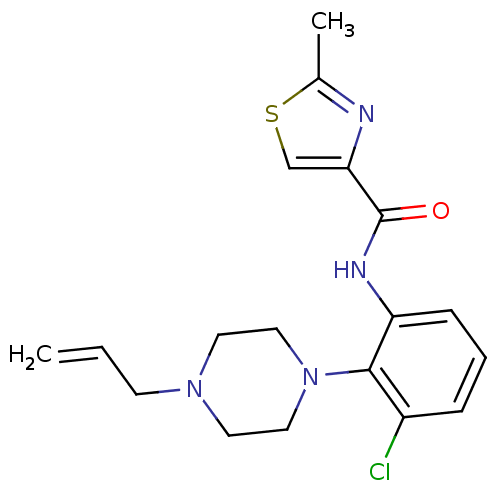
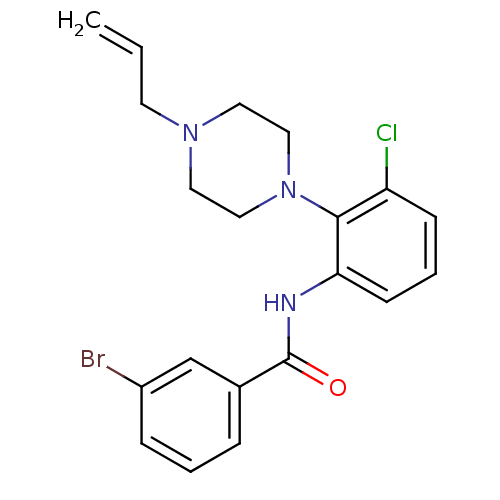
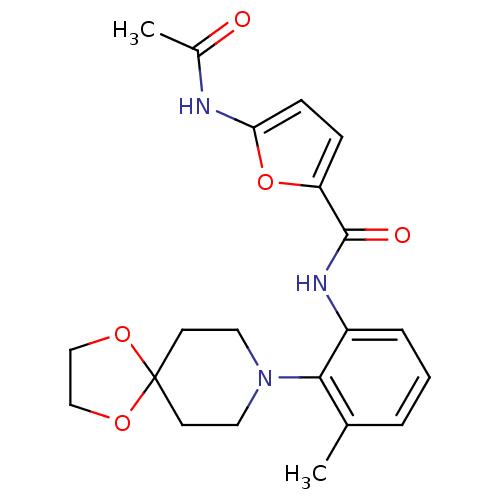
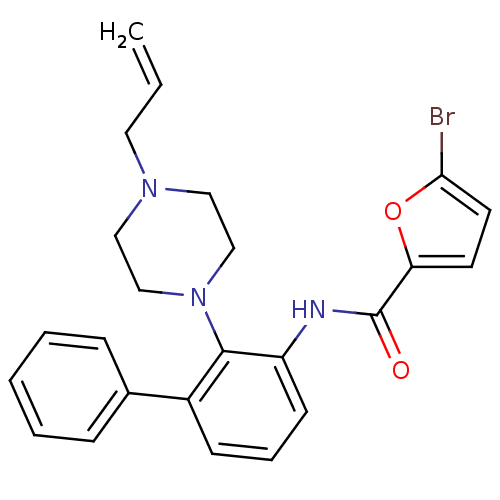
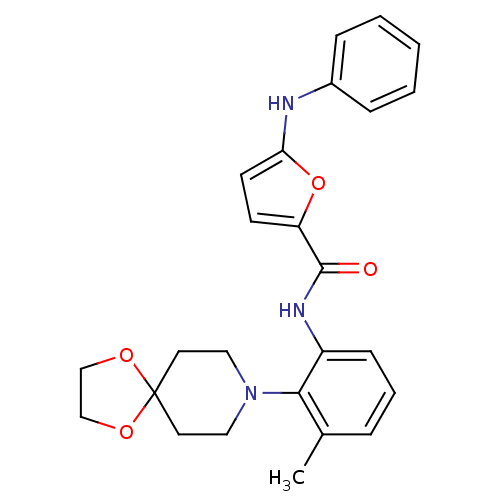
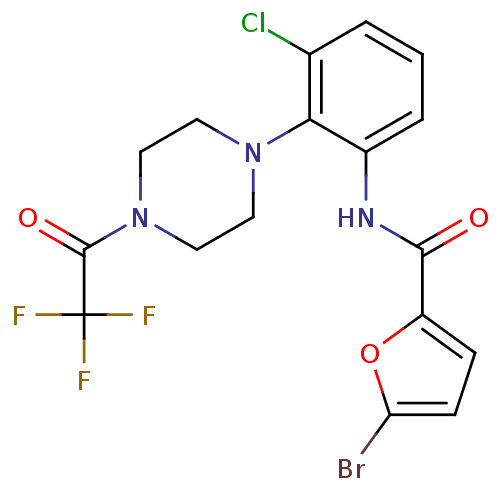
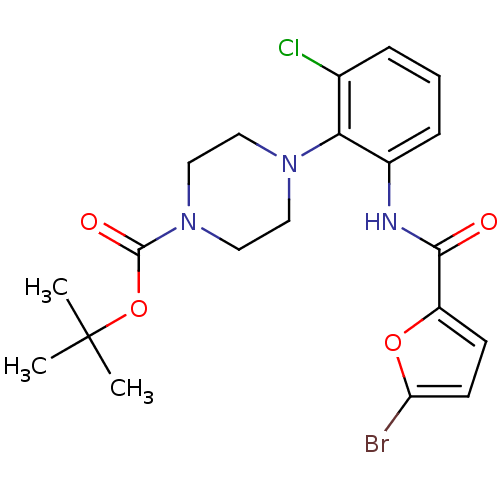
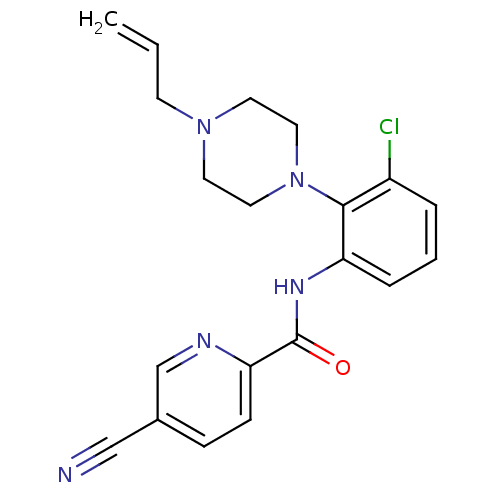
Report error Found 46 Enz. Inhib. hit(s) with Target = 'Isoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402]'
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 60nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 80nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 110nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 160nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 200nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 210nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 250nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 290nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 310nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 330nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 350nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 410nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 530nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 540nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 620nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 810nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 900nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 960nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.00E+3nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.00E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.10E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.10E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.20E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.26E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.40E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.80E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+3nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.20E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.30E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 4.80E+3nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 5.00E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 5.80E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 6.40E+3nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 8.90E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 9.90E+3nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 1.10E+4nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair
TargetIsoform Alpha-1 of Mitogen-activated protein kinase 10 (Alpha-1) 9-402](Human)
The Scripps Research Institute
The Scripps Research Institute
Affinity DataIC50: 2.00E+4nMpH: 7.0 T: 2°CAssay Description:Biochemical IC50s for JNK were determined using HTRF. In each assay the phosphor-Thr71-ATF-2 product was detected using a Europium-cryptate labeled a...More data for this Ligand-Target Pair


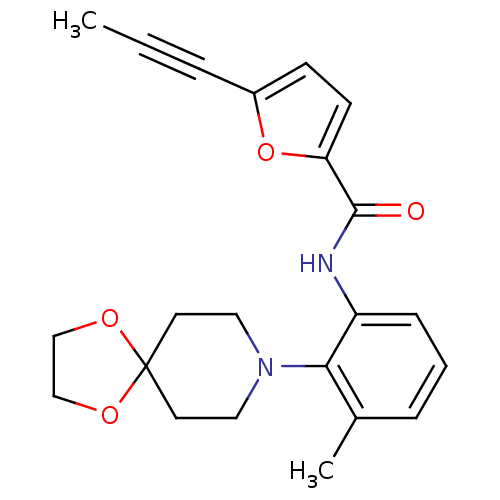
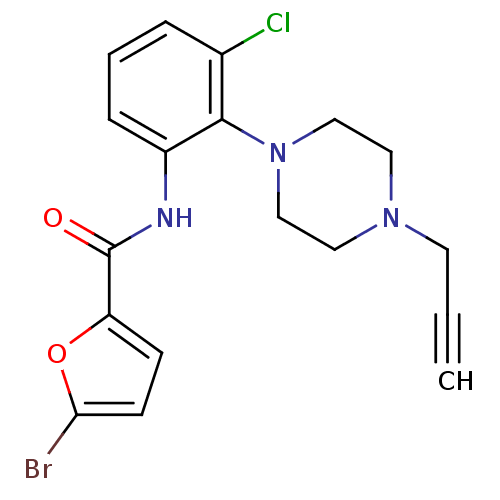
 3D Structure (crystal)
3D Structure (crystal)