Report error Found 198 Enz. Inhib. hit(s) with Target = 'Melanin-concentrating hormone receptor 2'
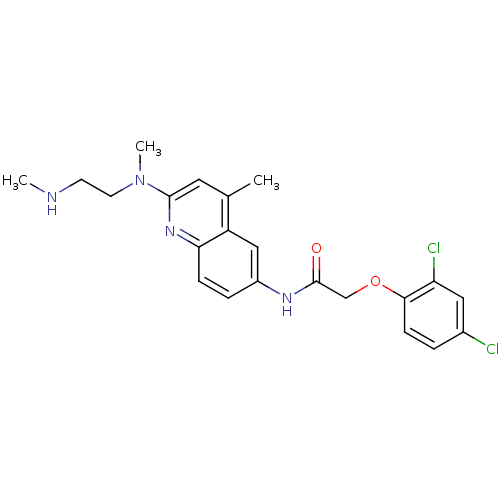
Affinity DataIC50: 0.100nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
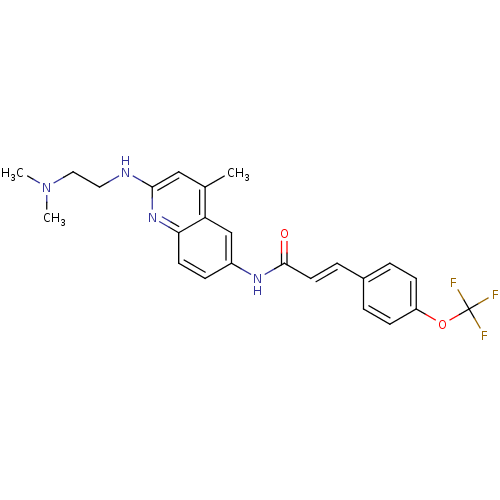
Affinity DataIC50: 0.100nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
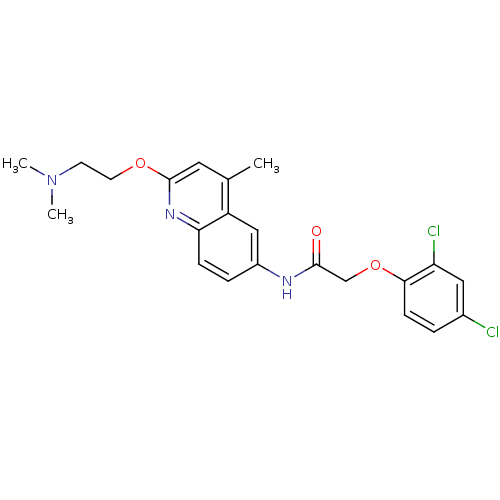
Affinity DataIC50: 1nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
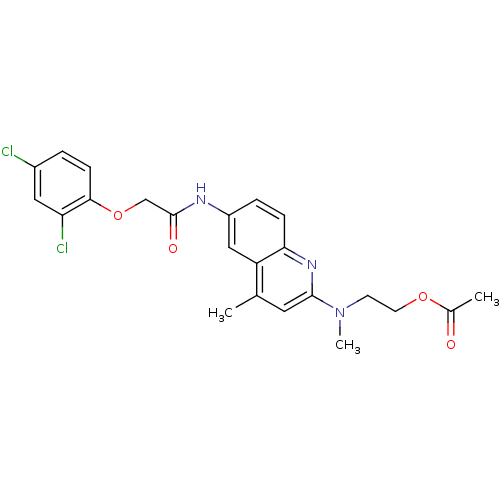
Affinity DataIC50: 1nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
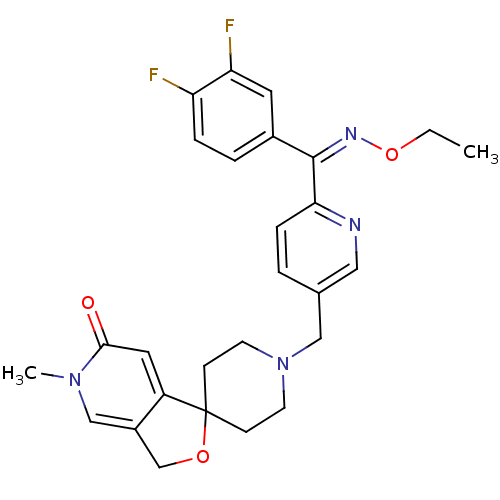
Affinity DataKi: 1.60nMAssay Description:Displacement of [35S] 4-(1-(3,4-difluorophenyl)-2-(ethyl(3-(6-fluoro-3H-spiro[isobenzofuran-1,4'-piperidine]-1'-yl)propyl)amino)-2-oxoethyl)-3-oxopip...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.30nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.40nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.80nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.80nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.10nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.40nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.70nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human MCH2RMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Displacement of 15 pM [125I]-MCH from human MCH2R expressed in CHO-K1 in whole-cell binding assayMore data for this Ligand-Target Pair
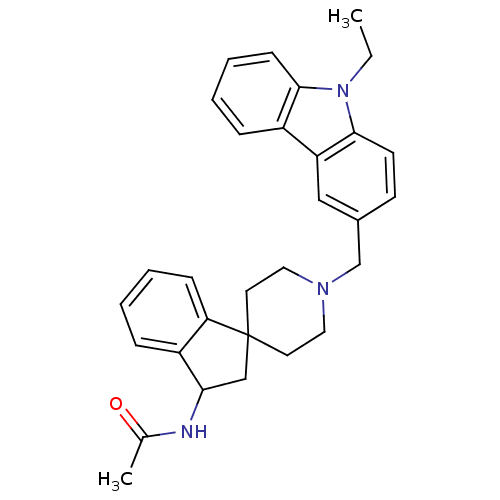
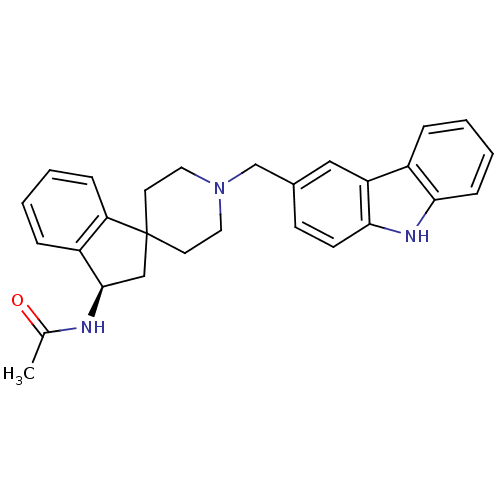
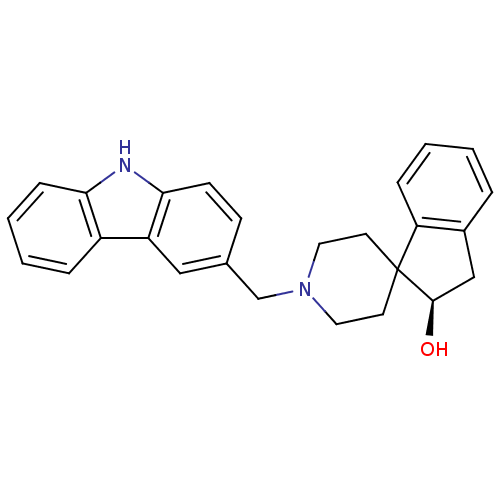
Affinity DataKi: 13nMAssay Description:Displacement of [3H]N-(1'-((9-ethyl-9H-carbazol-3-yl)methyl)-2,3-dihydrospiro[indene-1,4'-piperidine]-3-yl)acetamide from human MCHR2 expressed in HE...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Antagonist activity at human MCHR2 receptor expressed in CHO cells assessed as inhibition of MCH-stimulated Ca2+ flux preincubated for 10 mins prior ...More data for this Ligand-Target Pair