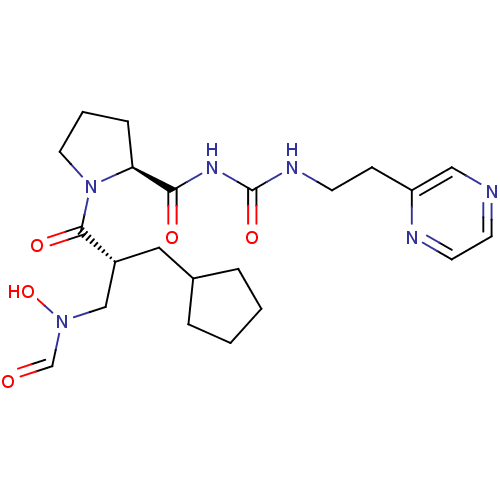
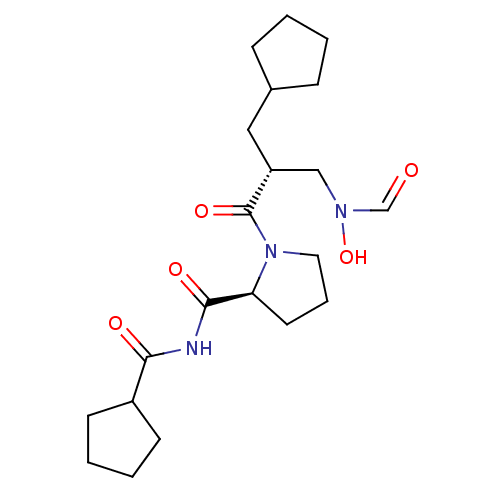
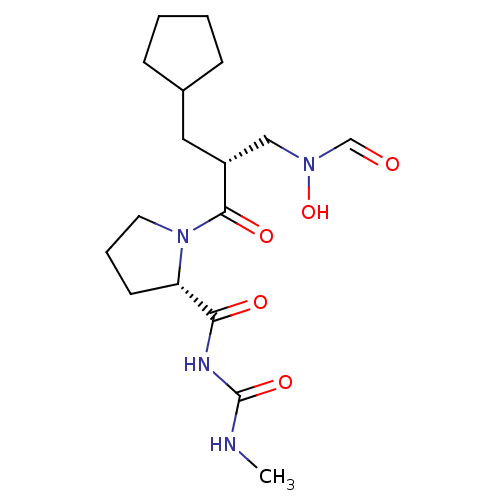
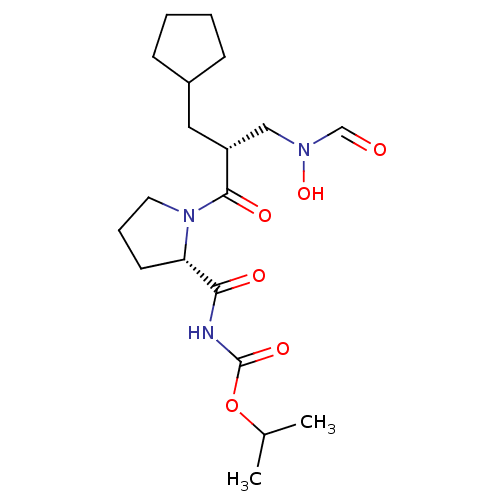
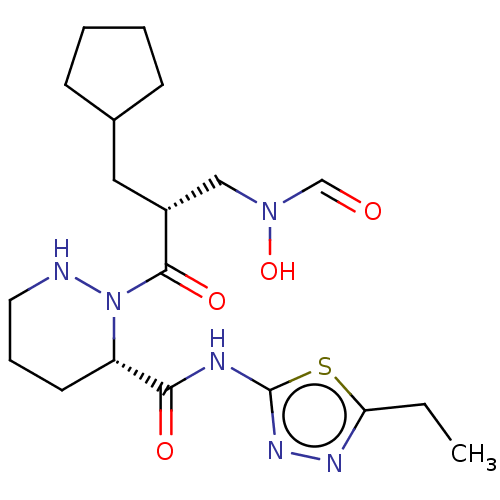
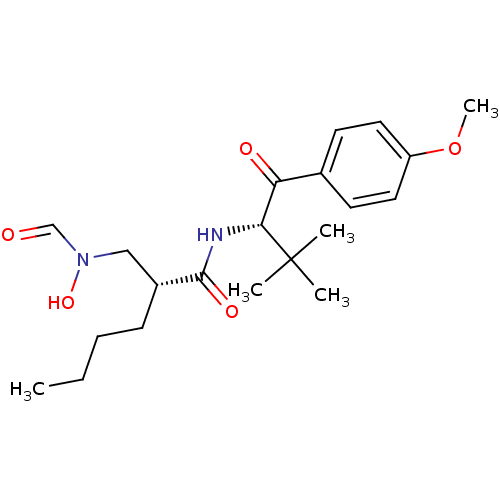
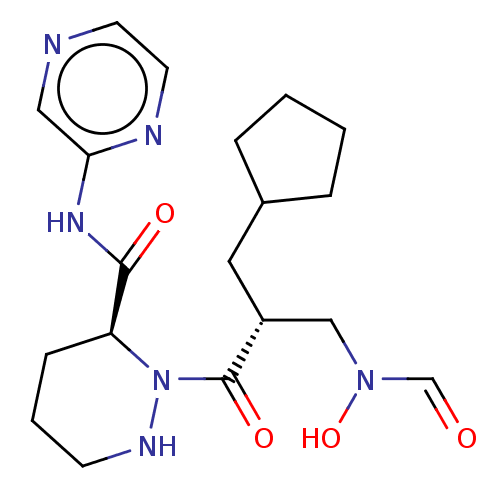
Report error Found 612 Enz. Inhib. hit(s) with Target = 'Peptide deformylase'
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.120nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.150nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.190nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.190nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataKi: 0.220nMAssay Description:Inhibitory effect against Escherichia coli peptide deformylase (PDF) by AAP assayMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.220nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Inhibitory effect against Escherichia coli peptide deformylase (PDF) by AAP assayMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.260nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.280nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity towards Peptide deformylaseMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.310nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataKi: 0.330nMAssay Description:Inhibitory effect against Escherichia coli peptide deformylase (PDF) by AAP assayMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.350nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.350nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.370nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.380nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.410nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.430nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.440nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.460nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.480nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.5nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Inhibitory activity against Escherichia coli peptide deformylase (PDF) Nickel containing enzymeMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.510nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.520nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataIC50: 0.520nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.540nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inhibitory activity against Escherichia coli peptide deformylase (PDF) Nickel containing enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 0.630nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.660nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataKi: 0.670nMAssay Description:Binding affinity against Co(II)-substituted Escherichia coli peptide deformylase was determinedMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.700nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.720nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.770nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.800nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataIC50: 0.860nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
TargetPeptide deformylase(Streptococcus pneumoniae serotype 4 (strain ATCC B...)
Glaxosmithkline
Curated by ChEMBL
Glaxosmithkline
Curated by ChEMBL
Affinity DataIC50: 0.910nMAssay Description:Inhibition of Streptococcus pneumoniae PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled ass...More data for this Ligand-Target Pair
Affinity DataIC50: 0.930nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Compound was evaluated for inhibition of peptide deformylase, PDF.Ni of Escherichia coliMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of Staphylococcus aureus PDF by fluorescence detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibitory activity against Escherichia coli peptide deformylase (PDF) Nickel containing enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibitory activity against Escherichia coli peptide deformylase (PDF) Nickel containing enzymeMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
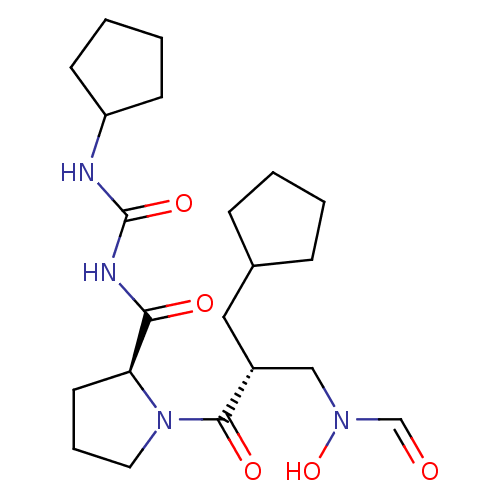
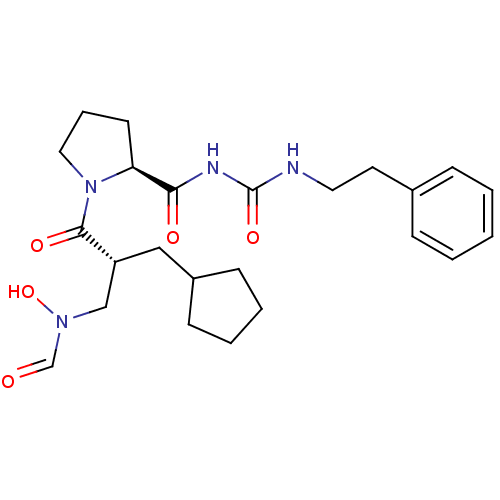
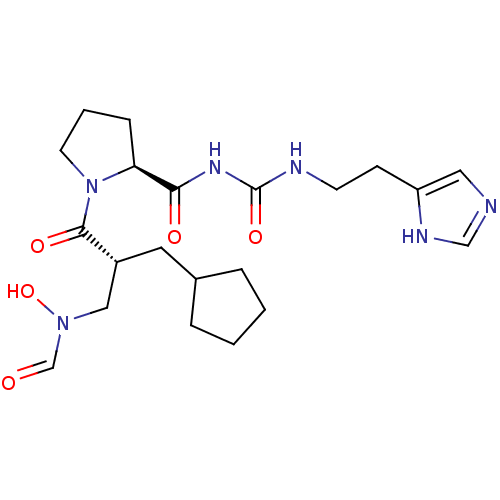

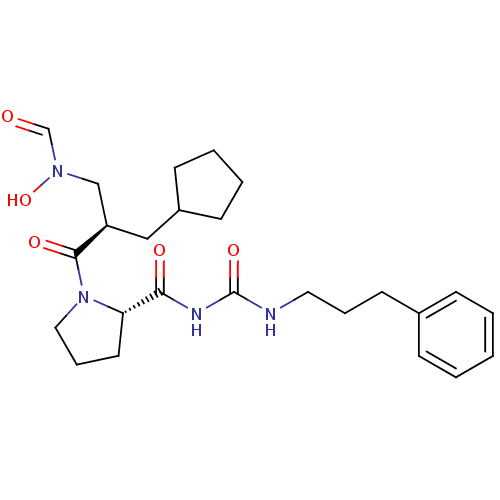
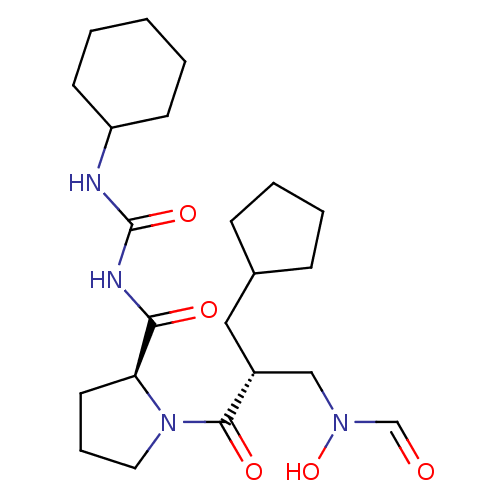
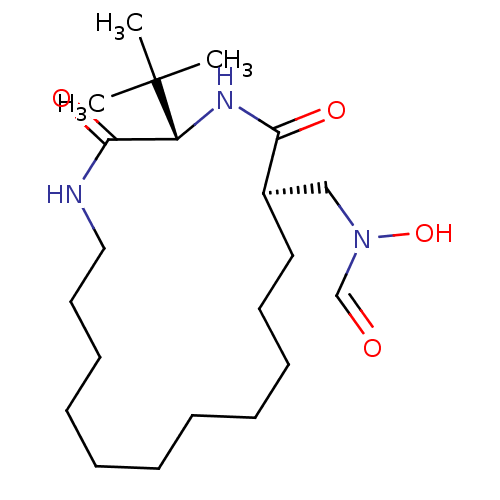



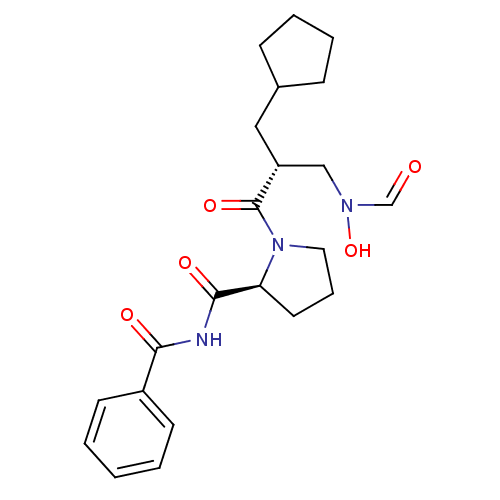
 3D Structure (crystal)
3D Structure (crystal)