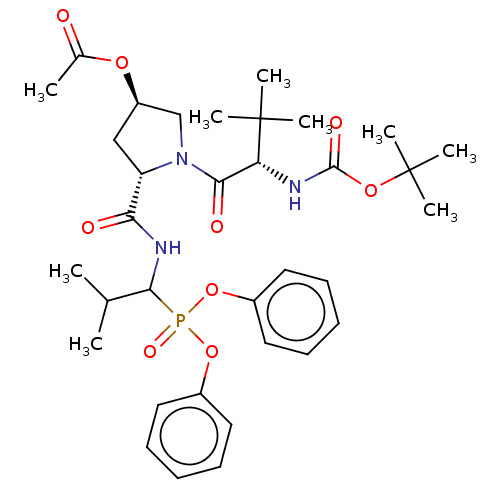
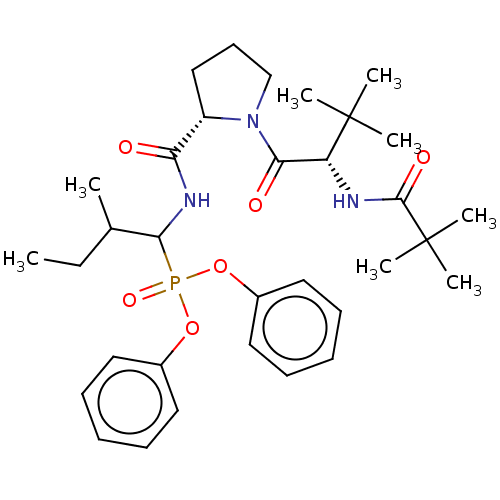
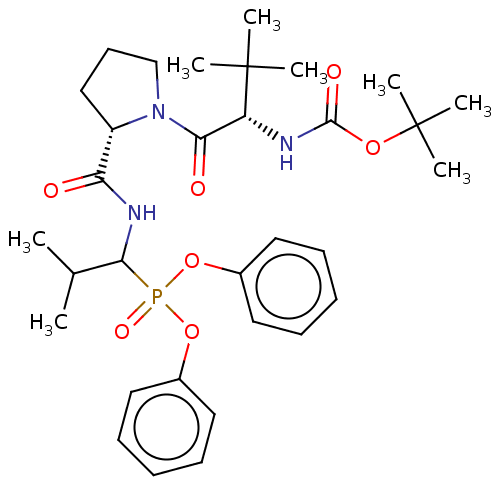
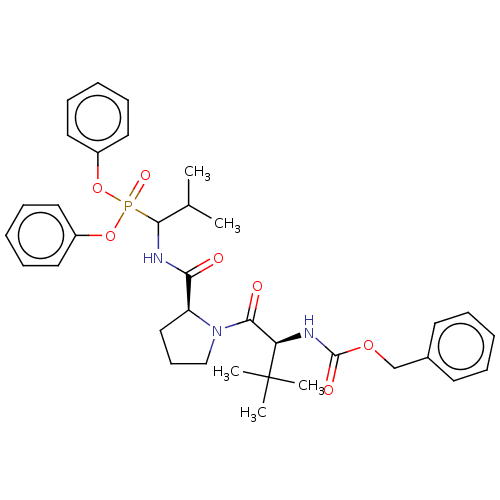
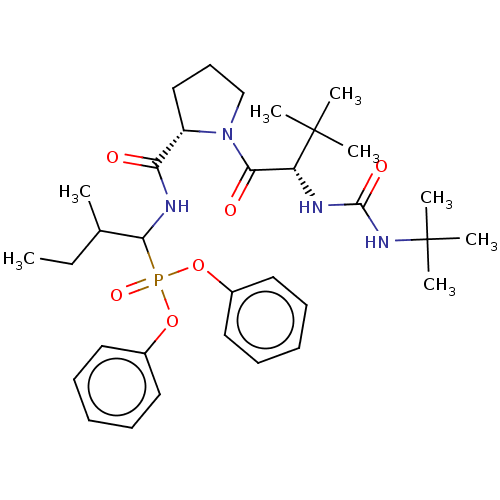
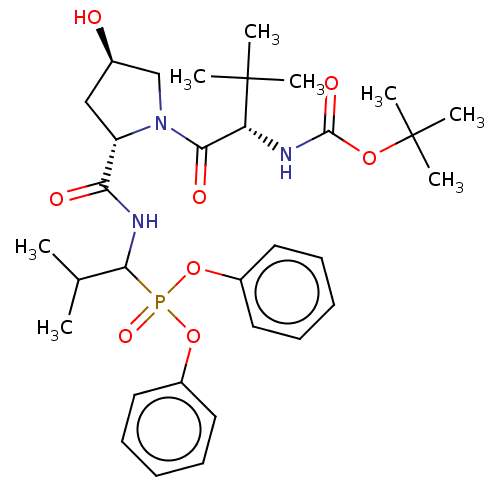
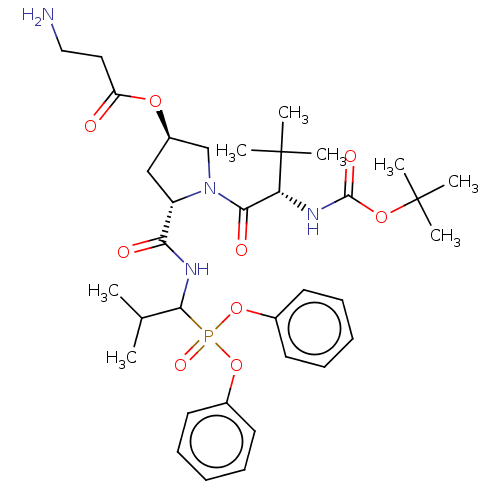
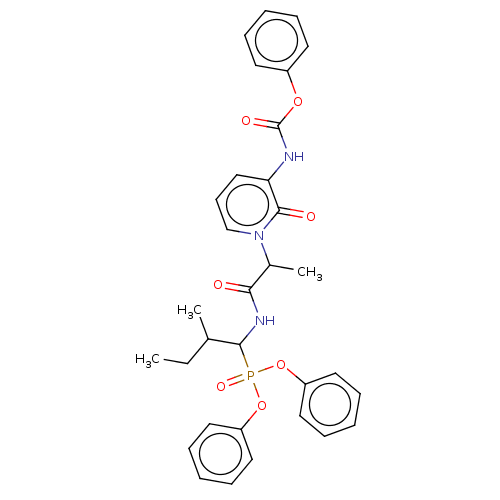
Report error Found 29 Enz. Inhib. hit(s) with Target = 'Probable periplasmic serine endoprotease DegP-like'
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
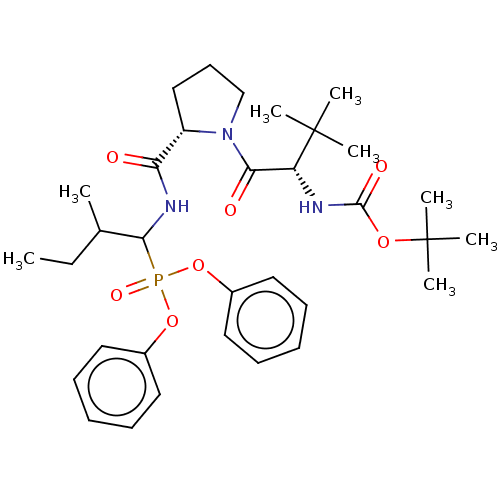
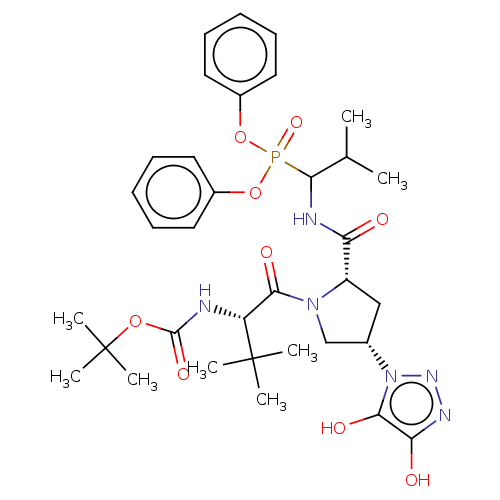
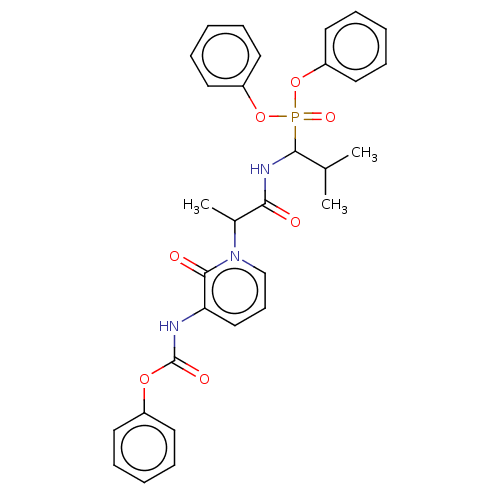
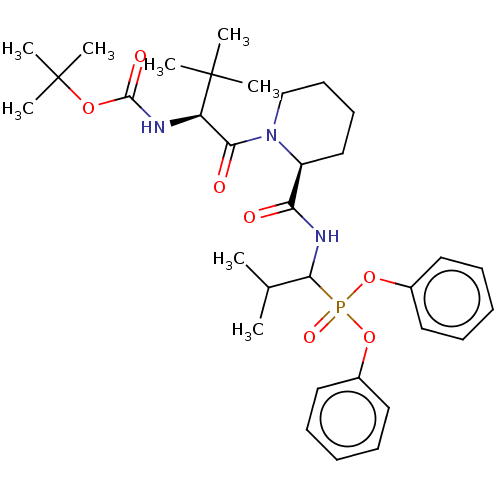
Affinity DataIC50: 3.86E+3nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
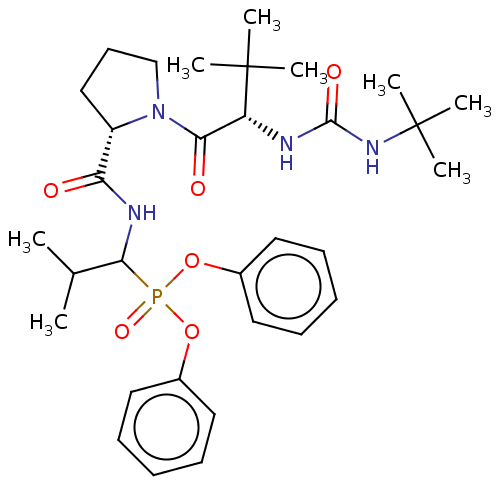
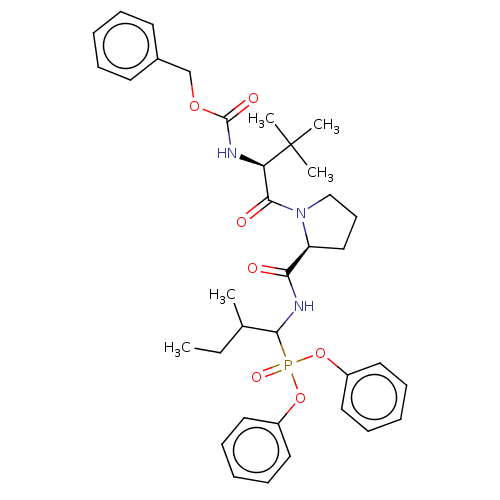
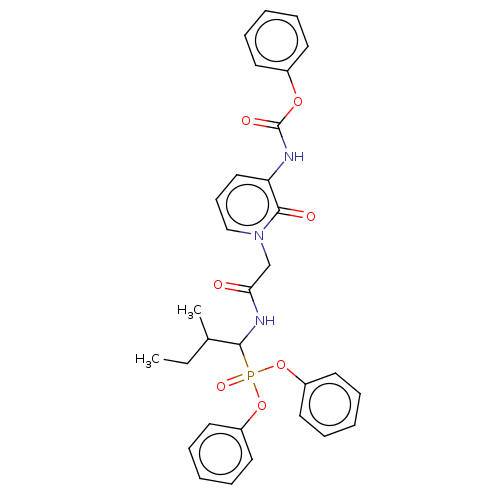
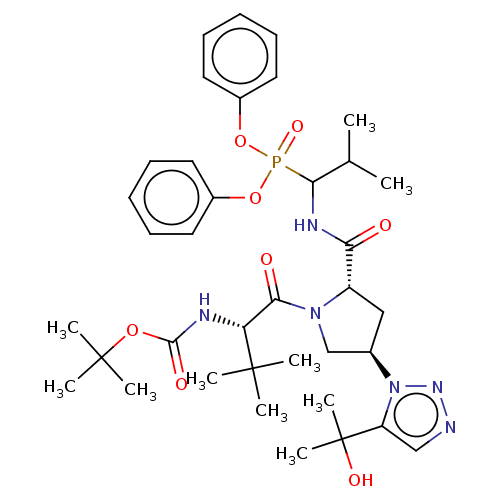
Affinity DataIC50: 4.57E+3nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
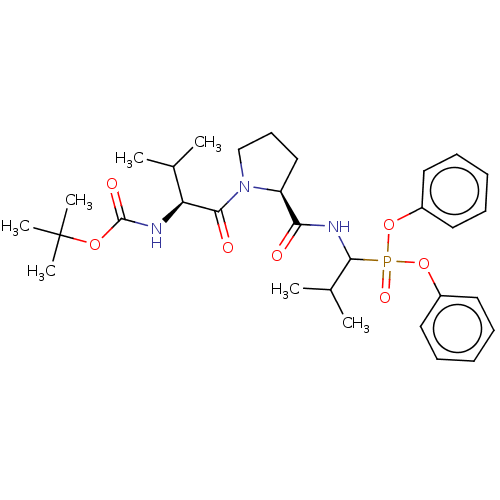
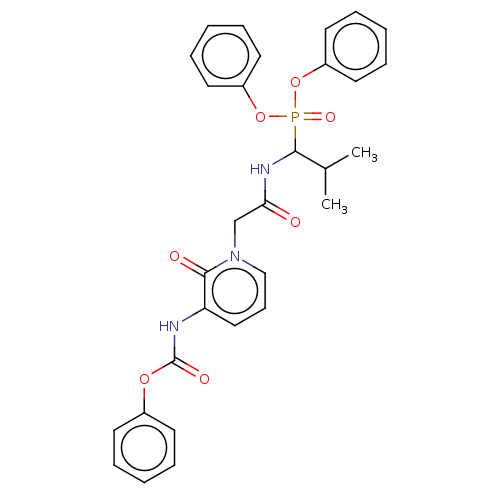
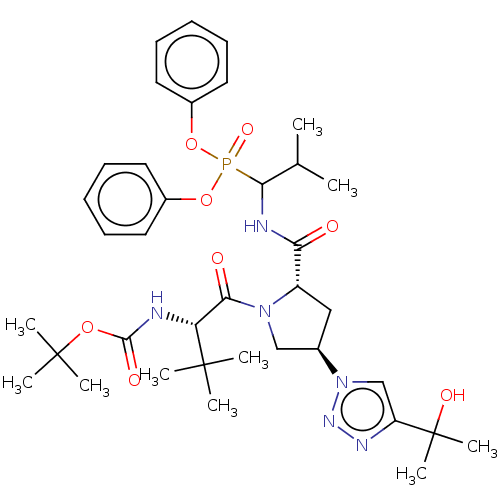
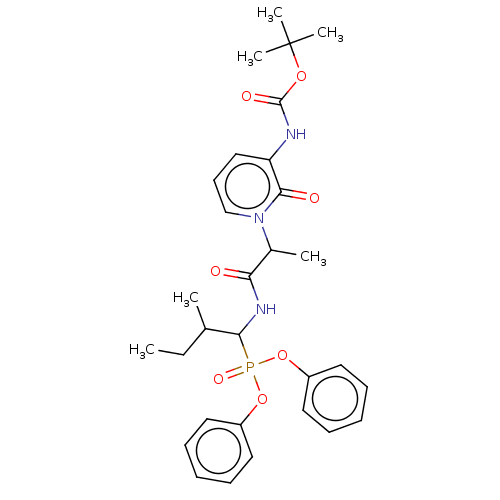
Affinity DataIC50: 9.30E+3nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
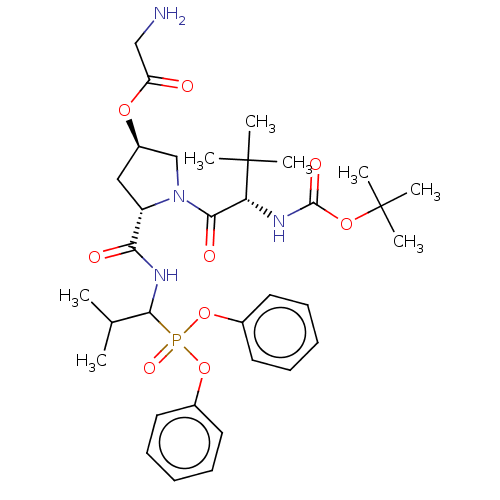
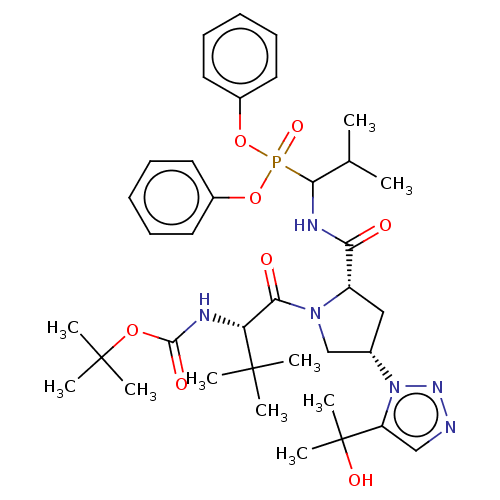
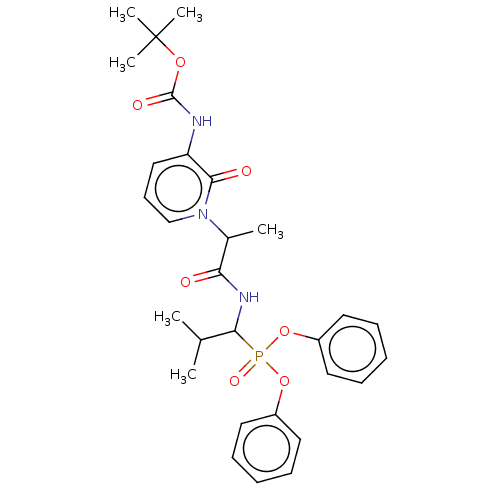
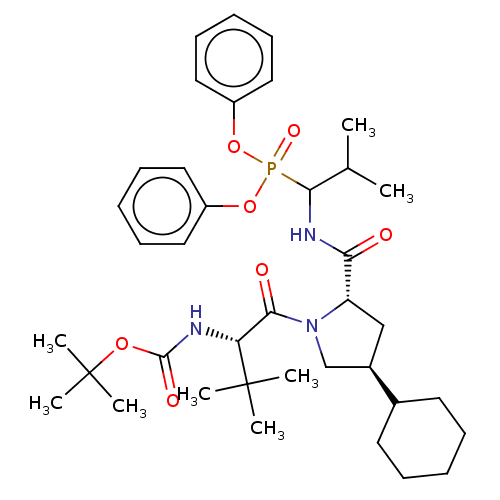
Affinity DataIC50: 9.30E+3nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.03E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.07E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.39E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.41E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.42E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.53E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.59E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.98E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 2.19E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 2.30E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 2.49E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 2.90E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 3.03E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 4.18E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 4.83E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 4.87E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 5.95E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 7.16E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 7.70E+4nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 7.97E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 8.18E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 9.57E+4nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.29E+5nMAssay Description:Inhibition of recombinant ctHtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate pretreated for 10 min followed by substrate addition and me...More data for this Ligand-Target Pair
TargetProbable periplasmic serine endoprotease DegP-like(Chlamydia trachomatis (strain D/UW-3/Cx))
University of Otago
Curated by ChEMBL
University of Otago
Curated by ChEMBL
Affinity DataIC50: 1.63E+5nMAssay Description:Inhibition of recombinant Chlamydia trachomatis HtrA serine protease using MeOCoum-ENLHLPLPI1F-D as substrate measured after 30 minsMore data for this Ligand-Target Pair