Report error Found 99 Enz. Inhib. hit(s) with Target = 'Proto-oncogene tyrosine-protein kinase LCK'
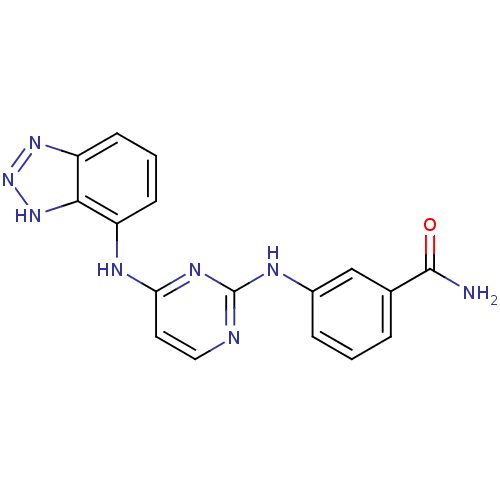
Affinity DataIC50: 8.5nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
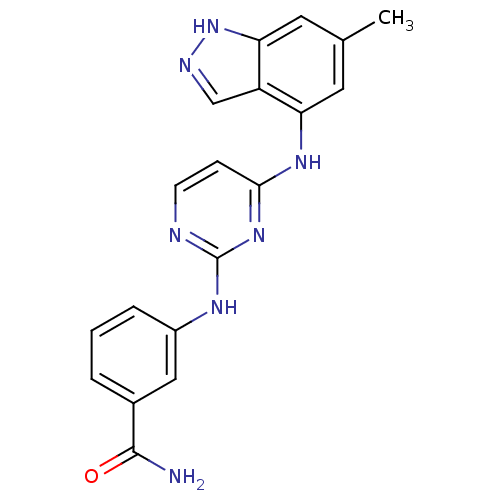
Affinity DataIC50: 12nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
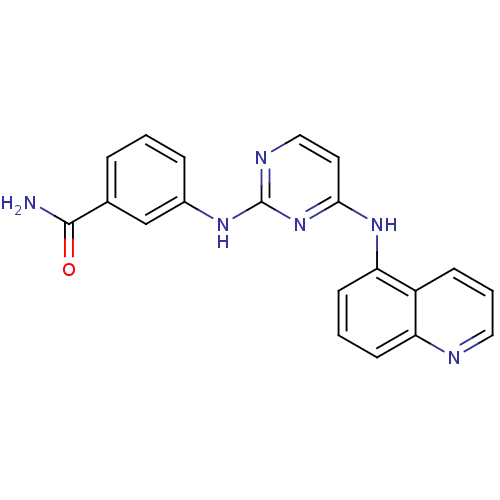
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
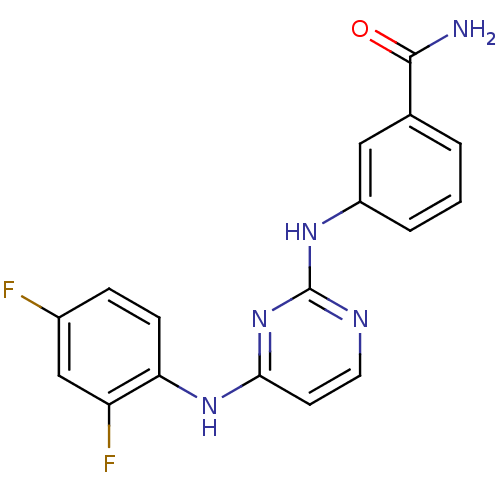
Affinity DataIC50: 19nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataKi: 20nM ΔG°: -43.5kJ/mole IC50: 59nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Inhibition of Lck in the presence of 50uM ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of Lck in the presence of 50uM ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 55nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:IC50 is the inhibitor concentration, which inhibits 50% of kinase activity that catalyzes the transfer of the terminal phosphate from [gamma-33P] lab...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of mouse p56 Lck tyrosine kinaseMore data for this Ligand-Target Pair
Affinity DataIC50: 73nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataEC50: 78nMAssay Description:Inhibition of TCR stimulated IL2 production in mice administered orallyMore data for this Ligand-Target Pair
Affinity DataIC50: 86nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 89nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 90nMAssay Description:IC50 is the inhibitor concentration, which inhibits 50% of kinase activity that catalyzes the transfer of the terminal phosphate from [gamma-33P] lab...More data for this Ligand-Target Pair
Affinity DataIC50: 97nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:IC50 is the inhibitor concentration, which inhibits 50% of kinase activity that catalyzes the transfer of the terminal phosphate from [gamma-33P] lab...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of Lck in the presence of 50uM ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of mouse p56 Lck tyrosine kinaseMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:IC50 is the inhibitor concentration, which inhibits 50% of kinase activity that catalyzes the transfer of the terminal phosphate from [gamma-33P] lab...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 270nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:Inhibition of mouse p56 Lck tyrosine kinaseMore data for this Ligand-Target Pair
Affinity DataIC50: 320nMpH: 7.0 T: 2°CAssay Description:IC50 is the inhibitor concentration, which inhibits 50% of kinase activity that catalyzes the transfer of the terminal phosphate from [gamma-33P] lab...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 380nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 460nMAssay Description:Inhibition of Lck in the presence of 50uM ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 490nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 510nMpH: 7.0 T: 2°CAssay Description:IC50 is the inhibitor concentration, which inhibits 50% of kinase activity that catalyzes the transfer of the terminal phosphate from [gamma-33P] lab...More data for this Ligand-Target Pair
Affinity DataIC50: 520nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 560nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 570nMAssay Description:Inhibition of Lck in the presence of 50uM ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 660nMAssay Description:Inhibition of Lck in the presence of 50uM ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 720nMpH: 7.0 T: 2°CAssay Description:IC50 is the inhibitor concentration, which inhibits 50% of kinase activity that catalyzes the transfer of the terminal phosphate from [gamma-33P] lab...More data for this Ligand-Target Pair
Affinity DataIC50: 720nMAssay Description:Inhibition of mouse p56 Lck tyrosine kinaseMore data for this Ligand-Target Pair
Affinity DataIC50: 750nMAssay Description:Inhibition of Lck in the presence of 50uM ATPMore data for this Ligand-Target Pair
Affinity DataIC50: 880nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMpH: 7.4 T: 2°CAssay Description:Lck activity was assessed using a TR-FRET assay in a 384-well plate format. The degree of phosphorylation of Biotinylated substrate was measured usin...More data for this Ligand-Target Pair