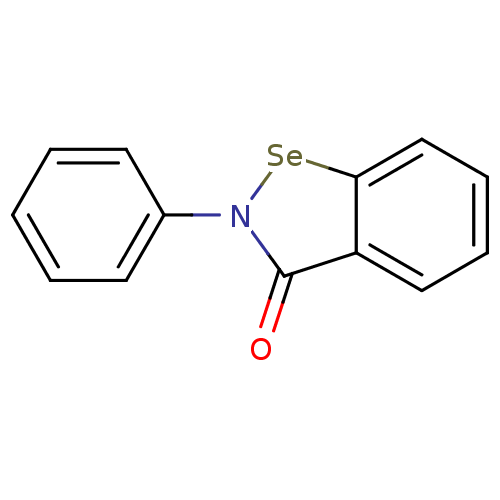
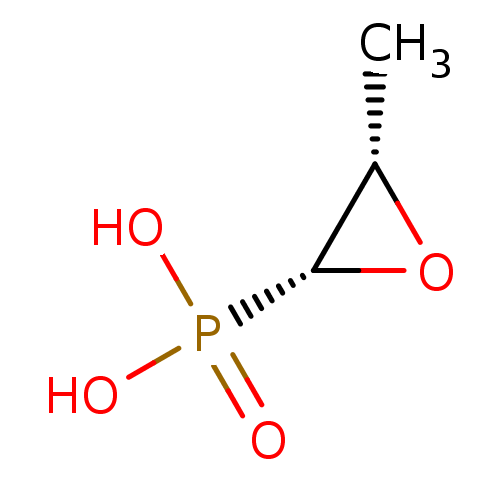
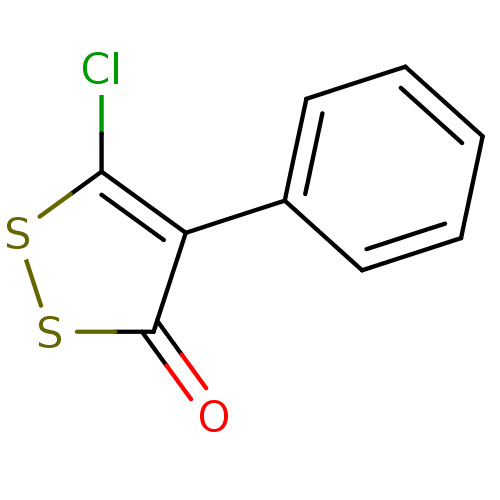
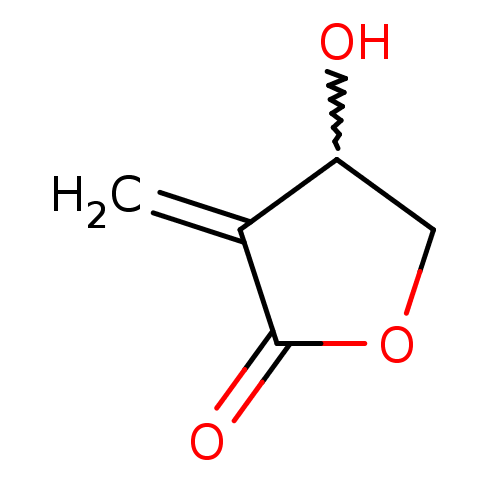
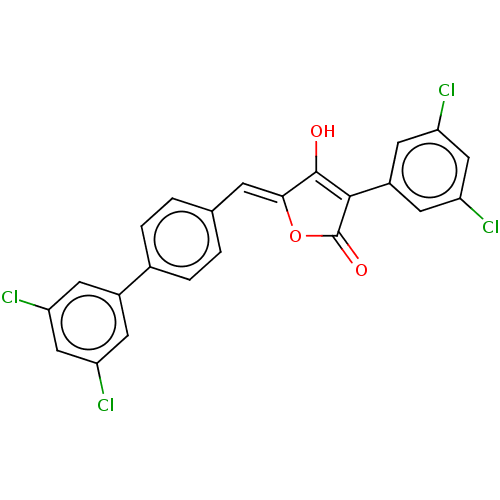
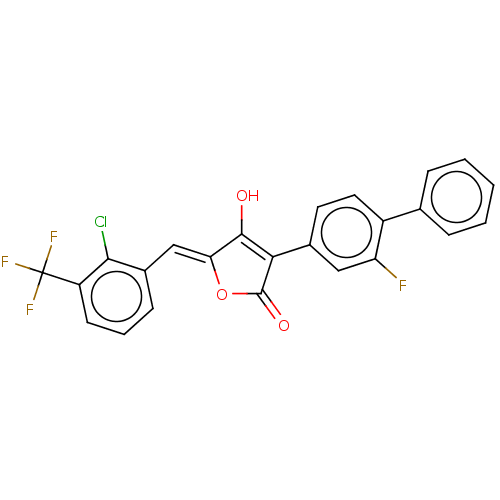
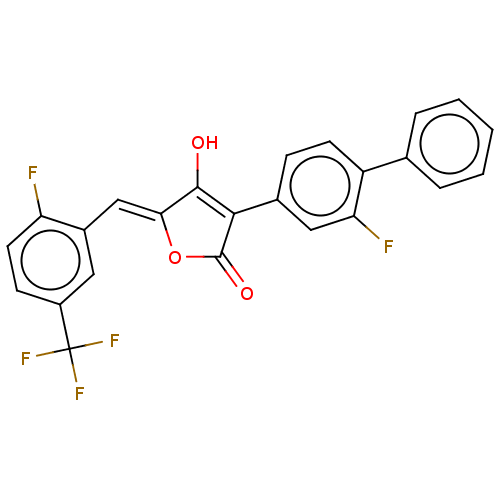
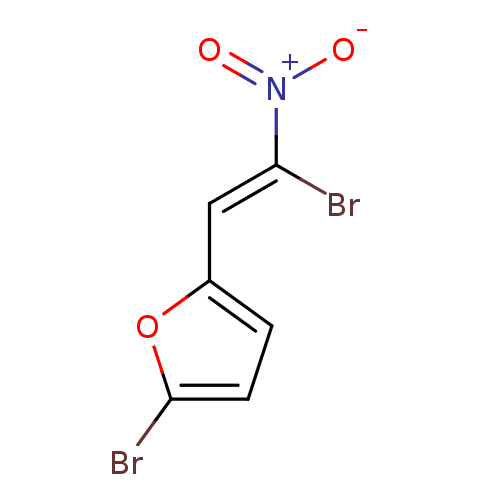
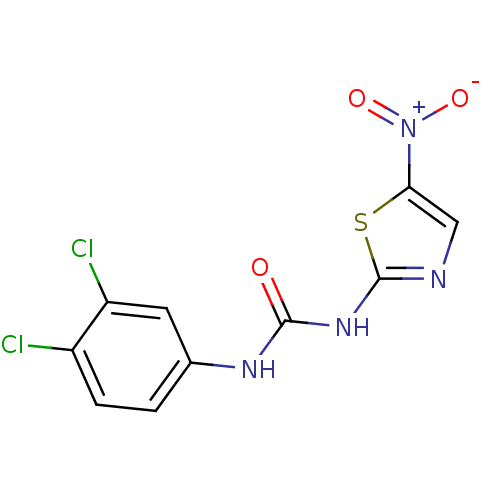
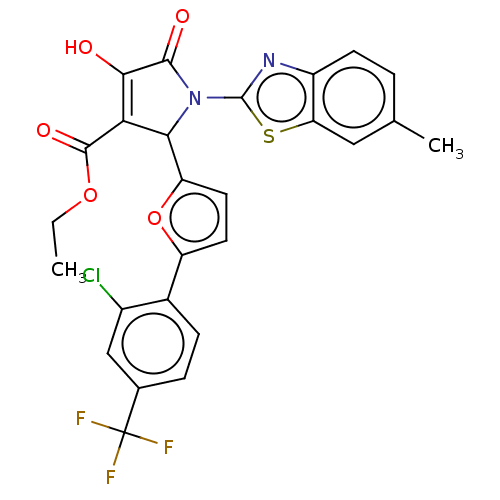
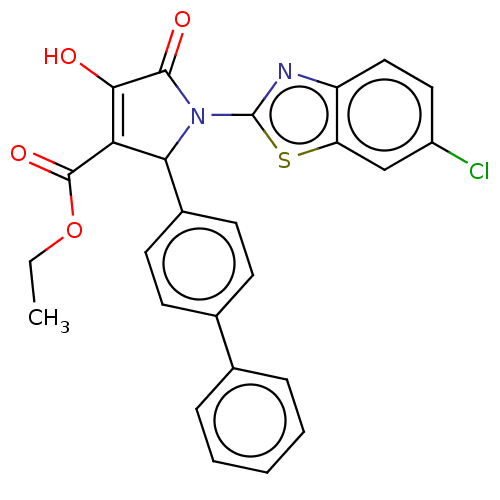
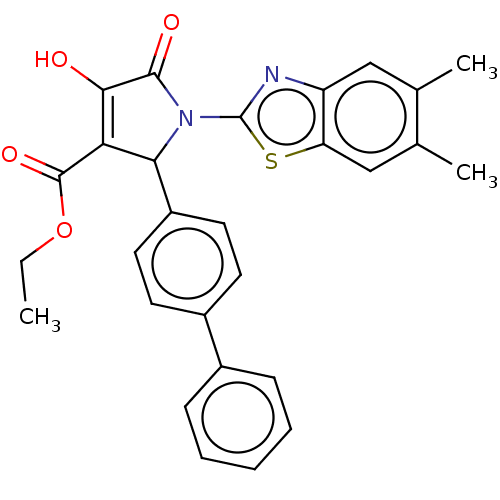
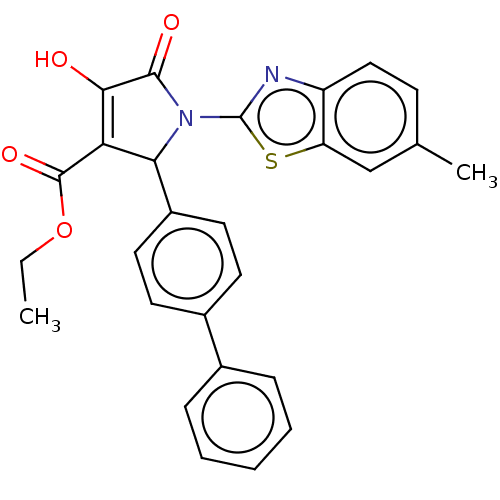
Report error Found 177 Enz. Inhib. hit(s) with Target = 'UDP-N-acetylglucosamine 1-carboxyvinyltransferase'
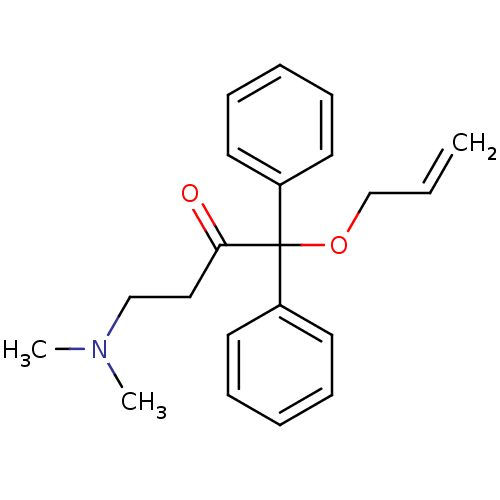
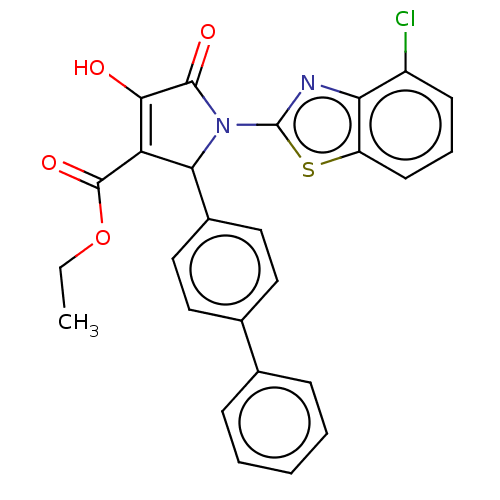
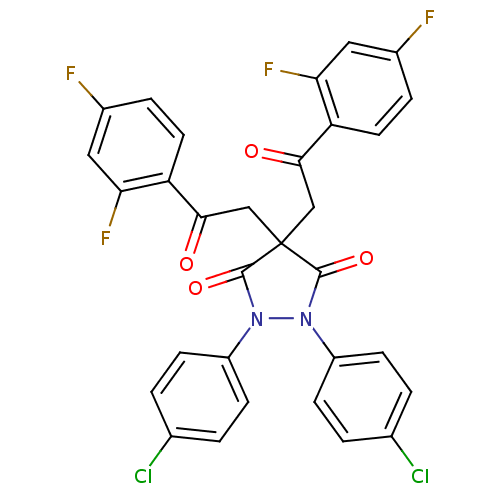
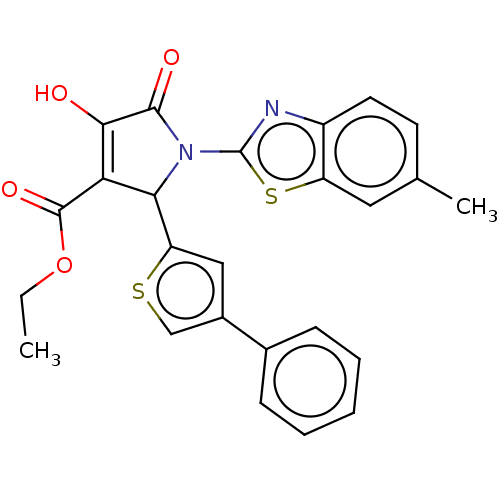
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Haemophilus influenzae (G-proteobacteria))
University of Ljubljana
University of Ljubljana
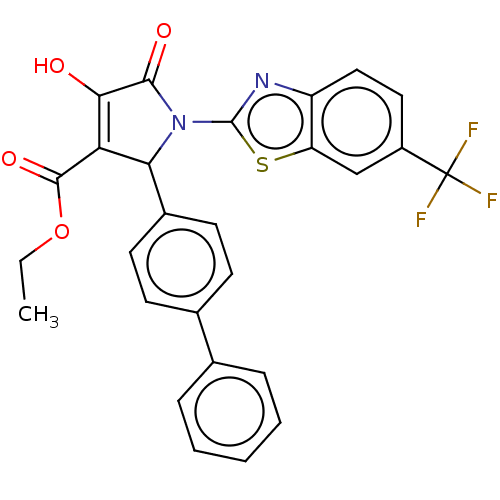
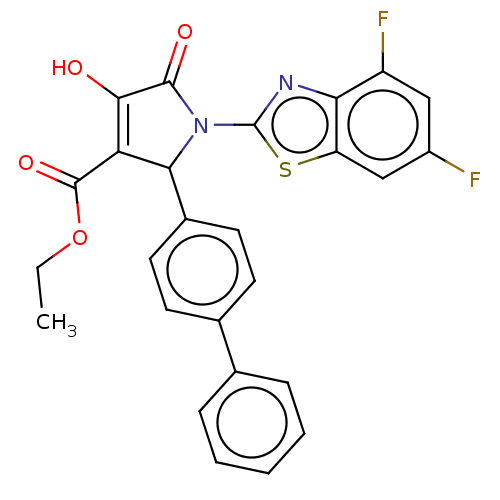
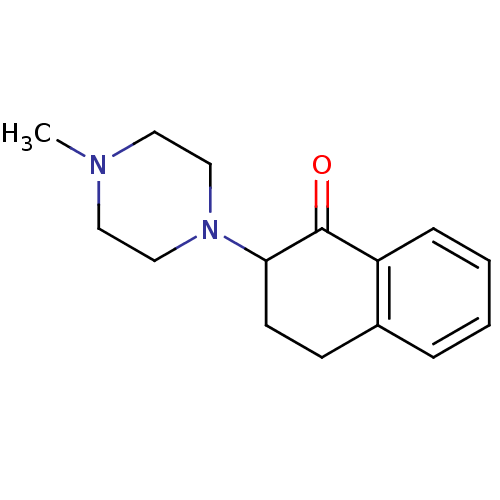
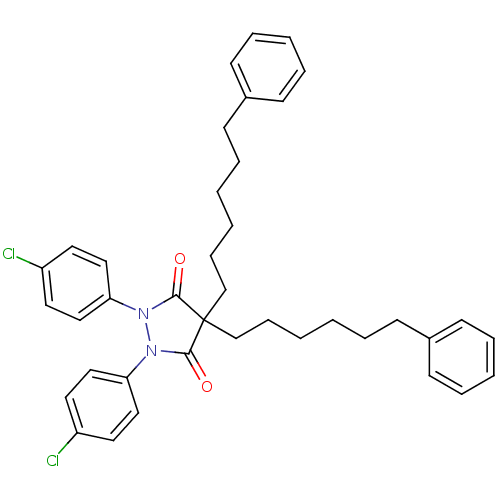
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
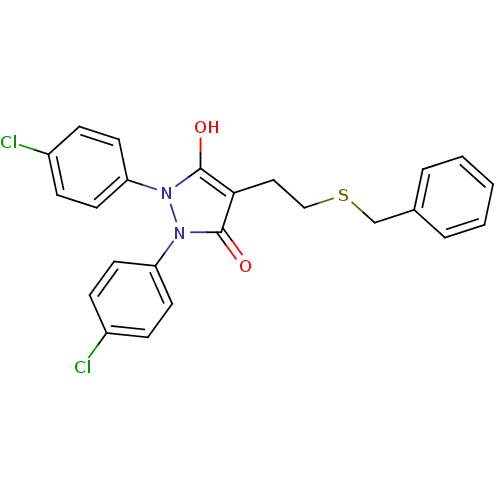
Affinity DataIC50: 100nMAssay Description:Inhibition of Escherichia coli MurA expressed in Escherichia coli BL21(lambdaDE3) using UNAG and PEP as substrate incubated for 10 mins prior to PEP ...More data for this Ligand-Target Pair
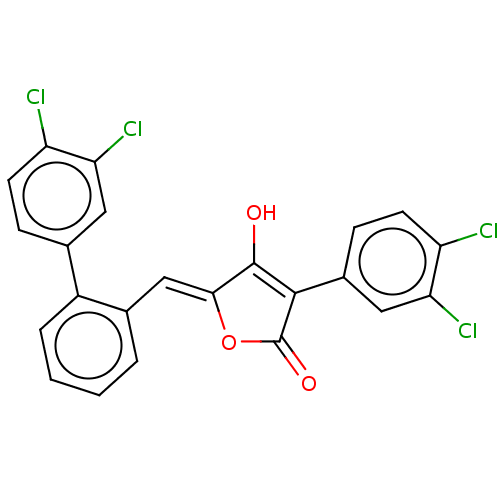
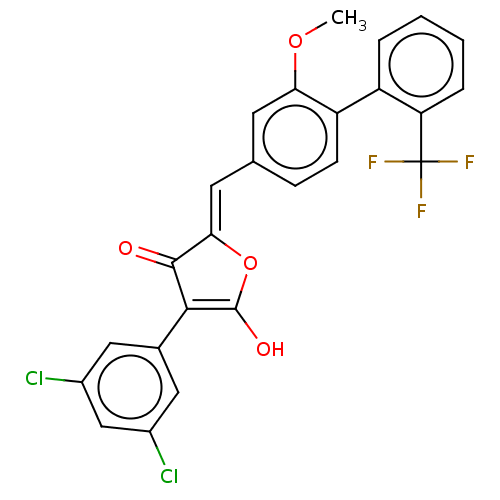
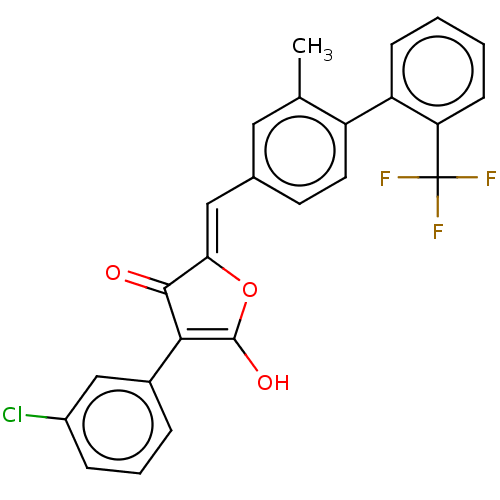
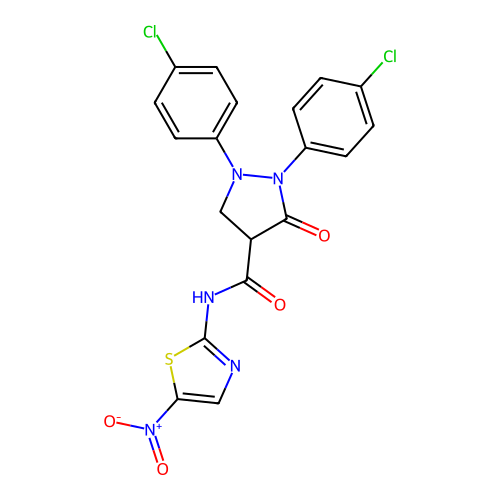
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
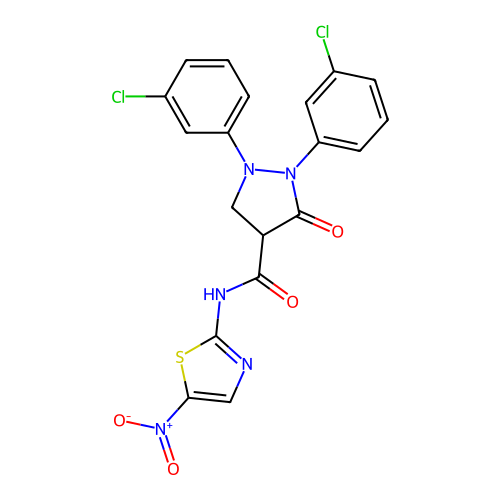
Affinity DataIC50: 118nMAssay Description:Inhibition of Escherichia coli K12 Mur A in presence of UNAGMore data for this Ligand-Target Pair
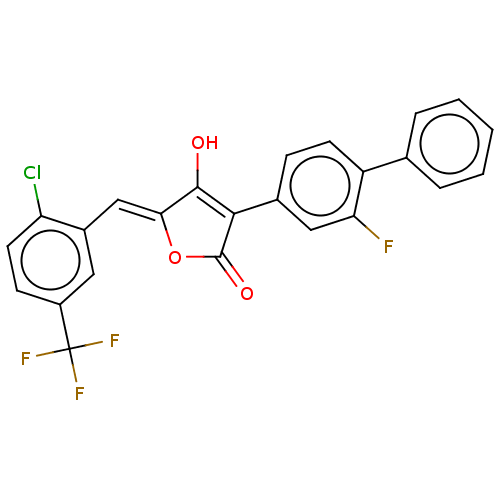
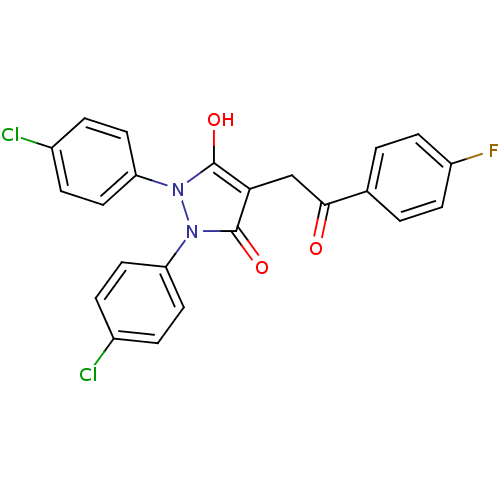
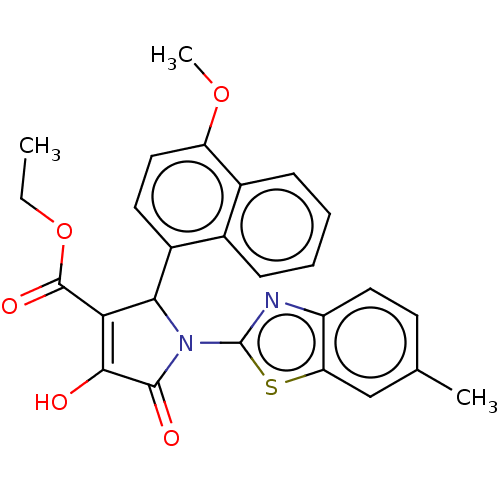
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
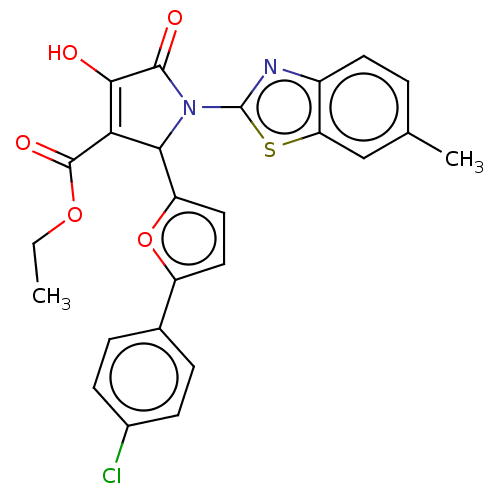
Affinity DataIC50: 210nMAssay Description:Inhibition of recombinant Escherichia coli MurA expressed in Escherichia coli NiCo21 using UNAG and PEP as substrate preincubated for 30 mins in pres...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase 2(Staphylococcus aureus subsp. aureus Mu50)
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 2.09E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 2.17E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 2.25E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of Escherichia coli MurA expressed in Escherichia coli BL21(lambdaDE3) using UNAG and PEP as substrate incubated for 10 mins prior to PEP ...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Haemophilus influenzae (G-proteobacteria))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase 2(Staphylococcus aureus subsp. aureus Mu50)
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase 2(Staphylococcus aureus subsp. aureus Mu50)
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase 1(Staphylococcus aureus subsp. aureus N315)
Wyeth Research
Curated by ChEMBL
Wyeth Research
Curated by ChEMBL
Affinity DataIC50: 4.07E+3nMAssay Description:Inhibitory activity against MurA in Staphylococcus aureusMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 4.18E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 4.57E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 4.60E+3nMAssay Description:Inhibition of Escherichia coli MurA expressed in Escherichia coli BL21(lambdaDE3) using UNAG and PEP as substrate incubated for 10 mins prior to PEP ...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 4.80E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of his tagged Escherichia coli K12 MurA C115D mutant using UDP-N-acetylglucosamine as substrate in presence of DTT and PEP by microplate r...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 5.10E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 5.80E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 6.10E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 6.27E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 6.51E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 6.70E+3nMAssay Description:Inhibition of Escherichia coli K12 His-tagged MurA expressed in Escherichia coli BL21 cells using UNAG as substrate pre incubated for 15 mins followe...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 7.50E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 7.88E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 8.40E+3nMAssay Description:Inhibitory concentration against MurA enzyme in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 8.50E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 8.75E+3nMAssay Description:Inhibitory activity against MurA in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 8.80E+3nMAssay Description:Inhibition of Escherichia coli K-12 MBP-fused C-terminal His-tagged MurA (6508 to 7768 residues) using UNAG as substrate preincubated for 10 mins fol...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 9.50E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 9.60E+3nMAssay Description:Inhibitory concentration against MurA enzyme in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 9.80E+3nMAssay Description:Inhibitory concentration against MurA enzyme in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 9.80E+3nMAssay Description:Inhibition of Escherichia coli K12 His-tagged MurA expressed in Escherichia coli BL21 cells using UNAG as substrate pre incubated for 15 mins followe...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 9.90E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 9.90E+3nMAssay Description:Inhibition of wild type his tagged Escherichia coli K12 MurA using UDP-N-acetylglucosamine as substrate preincubated with substrate for 30 mins prior...More data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 9.90E+3nMAssay Description:Inhibitory concentration against MurA enzyme in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylglucosamine 1-carboxyvinyltransferase(Escherichia coli (strain K12))
Heidelberg University
Curated by ChEMBL
Heidelberg University
Curated by ChEMBL
Affinity DataIC50: 1.02E+4nMAssay Description:Inhibition of Escherichia coli K12 His-tagged MurA expressed in Escherichia coli BL21 cells using UNAG as substrate pre incubated for 15 mins followe...More data for this Ligand-Target Pair