Report error Found 3239 Enz. Inhib. hit(s) with Target = 'cAMP-specific 3',5'-cyclic phosphodiesterase 4D'
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of human PDE4D using [3H]cAMP by packard topcount scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of human PDE4D using [3H]cAMP by packard topcount scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0210nMAssay Description:Inhibition of human PDE4D by scintillation counting methodMore data for this Ligand-Target Pair
TargetcAMP-specific 3',5'-cyclic phosphodiesterase 4D [372-376,381-715,S375G,I381V,S383G,G384S,V385H,K386M)(Human)
Plexxikon
Plexxikon
Affinity DataIC50: 0.0210nMAssay Description:Phosphodiesterase activities were assayed in the presence of inhibitor compounds. Measurement takes advantage of the selective binding of 5-AMP or 5-...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0210nMAssay Description:Inhibition of human His-tagged PDE4D catalytic domain expressed in Escherichia coli BL21-CodonPlus(DE3) cells using [3H]cAMP or [3H]cGMP as substrate...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0210nMAssay Description:Inhibition of human His-tagged PDE4D catalytic domain expressed in Escherichia coli BL21-CodonPlus(DE3) cells using [3H]cAMP or [3H]cGMP as substrate...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0210nMAssay Description:Inhibition of human PDE4DMore data for this Ligand-Target Pair
Affinity DataKi: 0.0617nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Inhibition of human GST-PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Inhibition of human PDE4D3 assessed as inhibition of [3H]cAMP hydrolysis to [3H]AMP after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Inhibition of human GST-PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Inhibition of human GST-PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Inhibition of human GST-PDE4DMore data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataKi: 0.123nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:Inhibition of human recombinant PDE4D using [3H]cAMP as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured af...More data for this Ligand-Target Pair
Affinity DataKi: 0.155nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataKi: 0.155nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataKi: 0.162nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataKi: 0.170nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.190nMAssay Description:Inhibitory activity against PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:Inhibitory activity against PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:Inhibition of PDE4D (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:Inhibitory activity against PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:Inhibition of human PDE4DMore data for this Ligand-Target Pair
Affinity DataKi: 0.224nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataKi: 0.234nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.270nMAssay Description:Inhibition of human PDE4D3 assessed as inhibition of [3H]cAMP hydrolysis to [3H]AMP after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Inhibition of human recombinant PDE4D catalytic domain cloned from human HL60 cells assessed as inhibition of cAMP hydrolysisMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Inhibition of human GST-PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Inhibition of human recombinant PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Inhibition of human GST-PDE4DMore data for this Ligand-Target Pair
Affinity DataKi: 0.302nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataKi: 0.316nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataKi: 0.316nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.350nMAssay Description:Inhibition of PDE4DMore data for this Ligand-Target Pair
Affinity DataKi: 0.355nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Inhibition of PDE4D (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.389nMAssay Description:Inhibition of human PDE4D2 expressed in Escherichia coli BL21 (DE3) using cAMP as substrate by PDELight HTS cAMP phosphodiesterase Kit basedMore data for this Ligand-Target Pair
Affinity DataIC50: 0.390nMAssay Description:Inhibition of his-tagged recombinant human PDE4D catalytic domain (T86 to S413 residues) expressed in Escherichia coli BL21(DE3) using [3H]-cAMP as s...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibitory activity against PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of PDE4D2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMpH: 7.5 T: 2°CAssay Description:PDE4 catalytic activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using PDE-SPA kit (Amersham International). [3H]-AMP was c...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of human PDE4D3 assessed as inhibition of [3H]cAMP hydrolysis to [3H]AMP after 15 mins by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of human recombinant PDE4D catalytic domain cloned from human HL60 cells assessed as inhibition of cAMP hydrolysisMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of human PDE4DMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of PDE4D (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of PDE4D2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of PDE4D (unknown origin) using cAMP as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 0.460nMAssay Description:Inhibition of PDE4DMore data for this Ligand-Target Pair

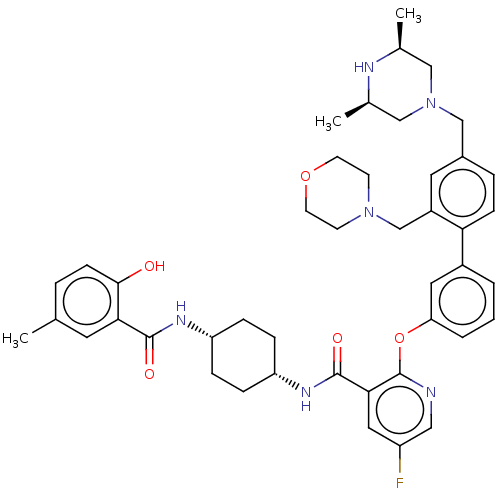
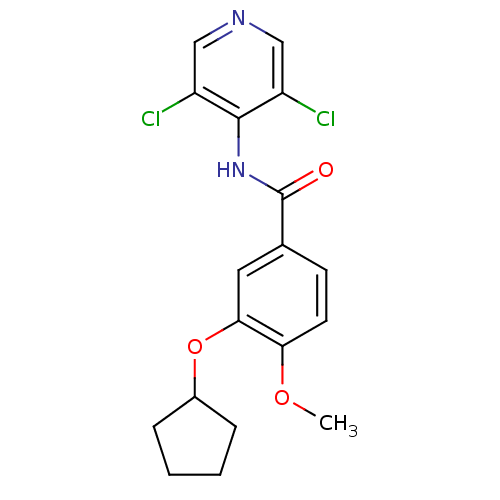
 3D Structure (crystal)
3D Structure (crystal)