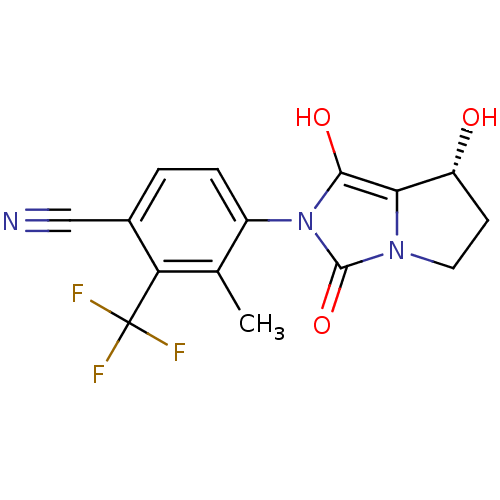
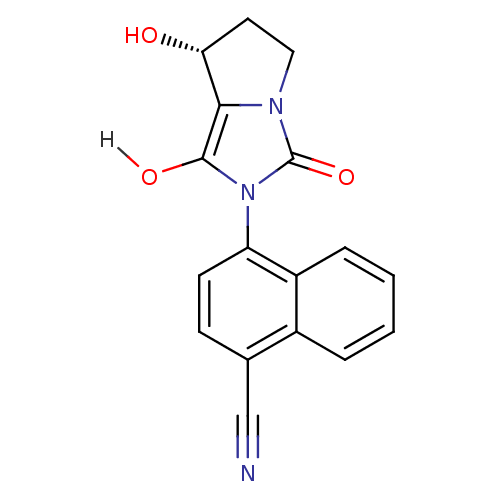
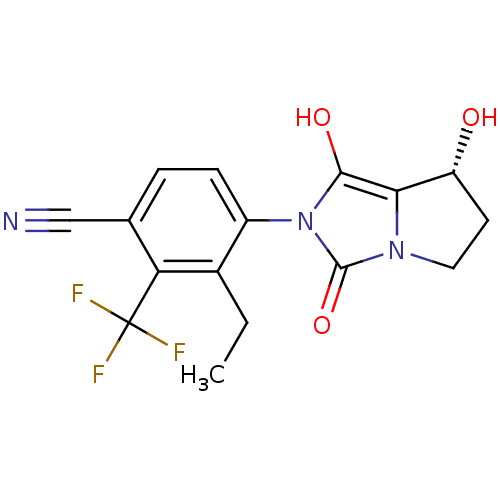
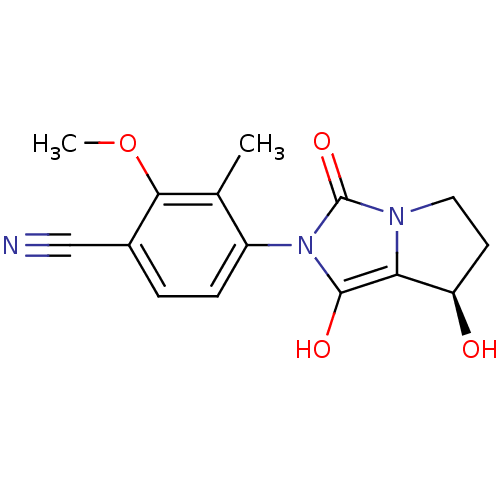
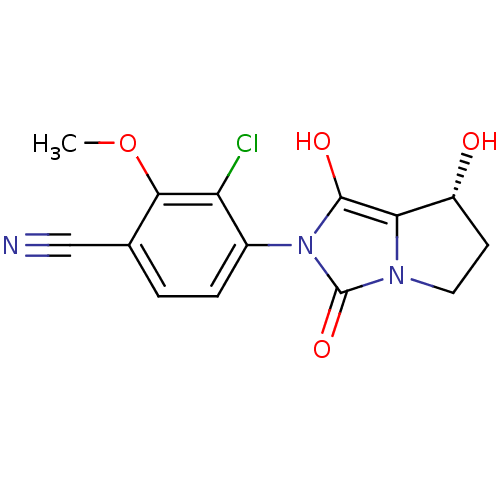
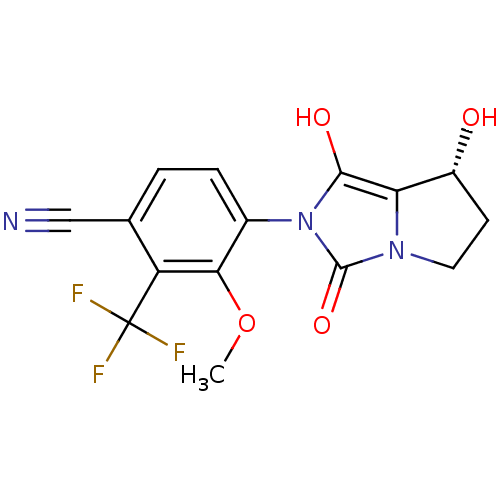
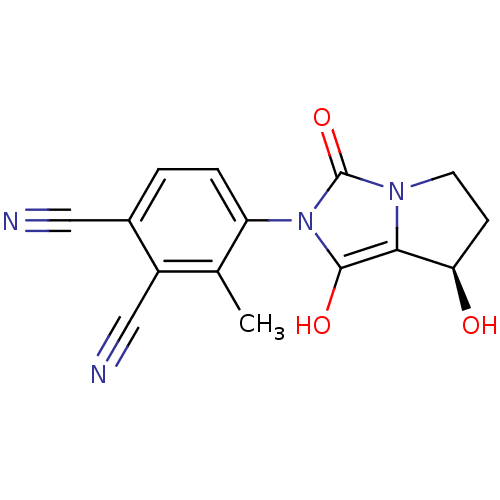
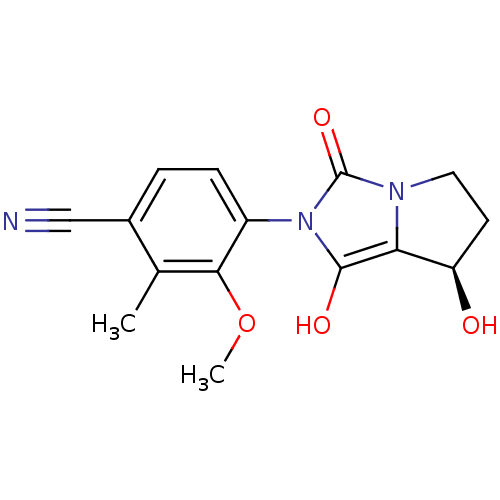
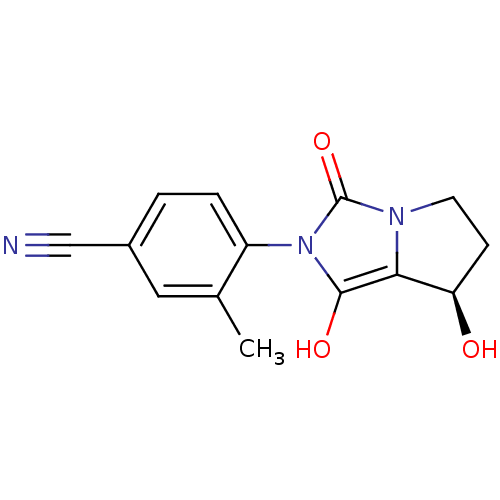
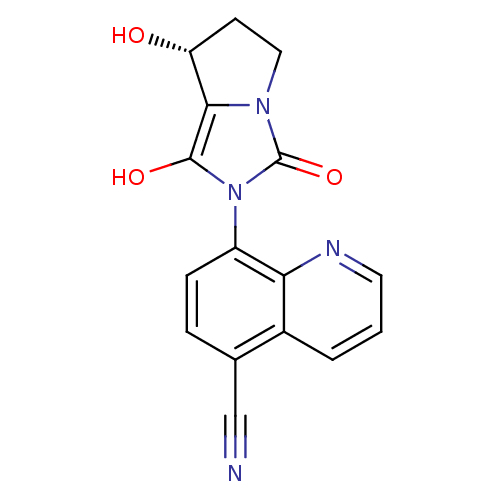
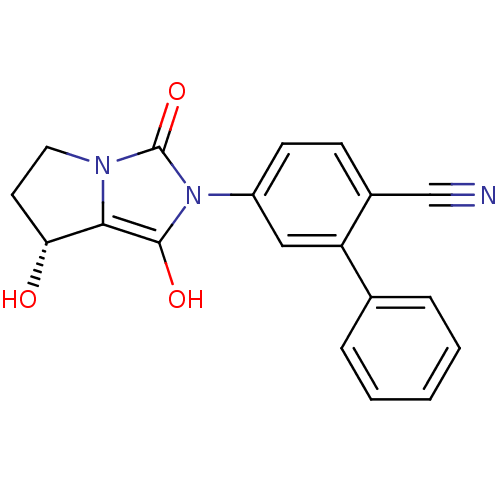
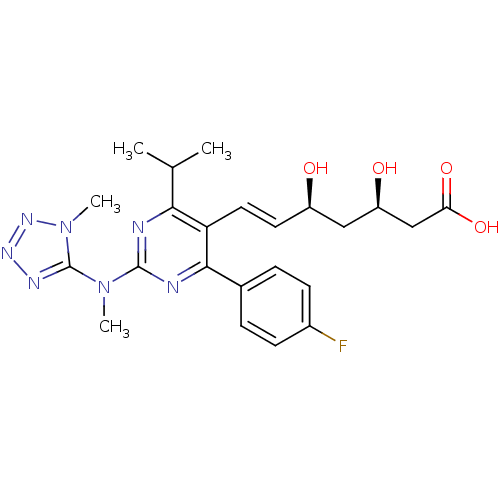
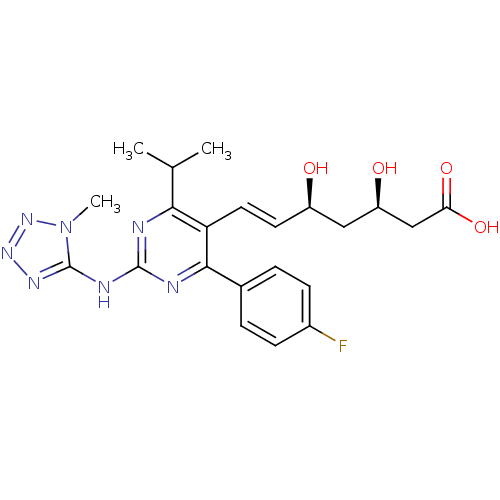
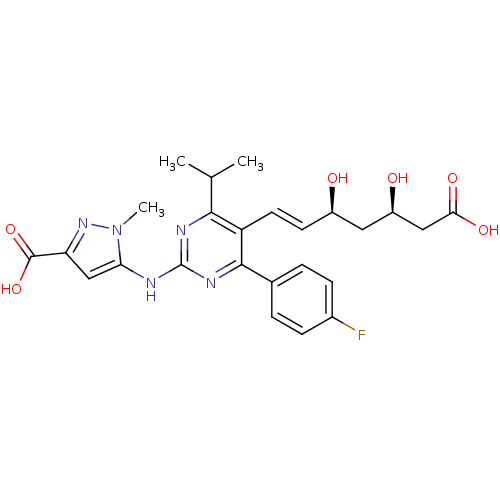
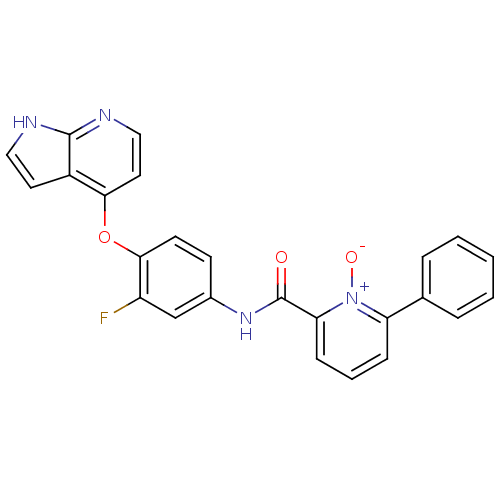
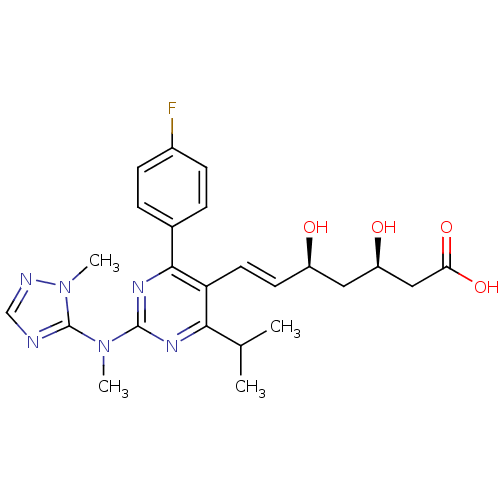
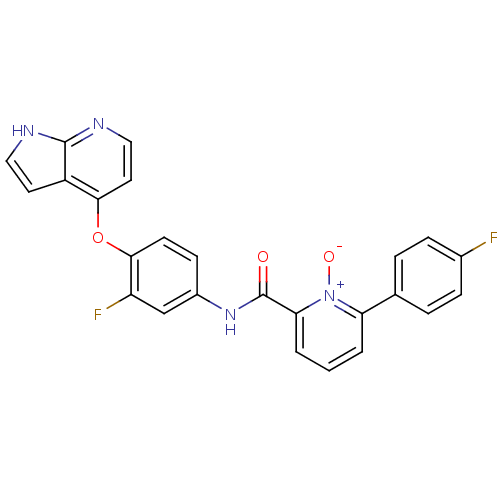
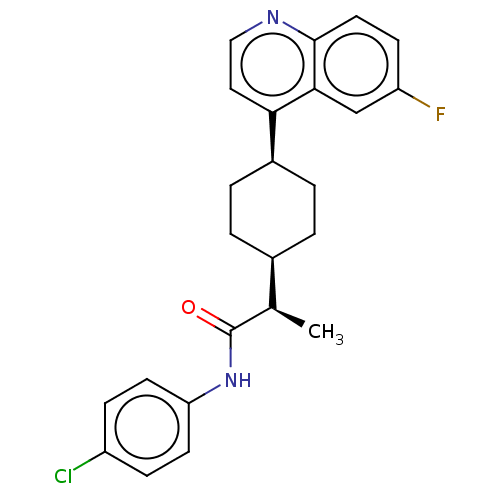
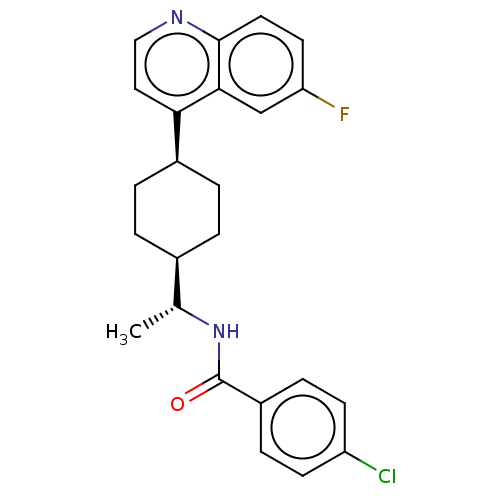
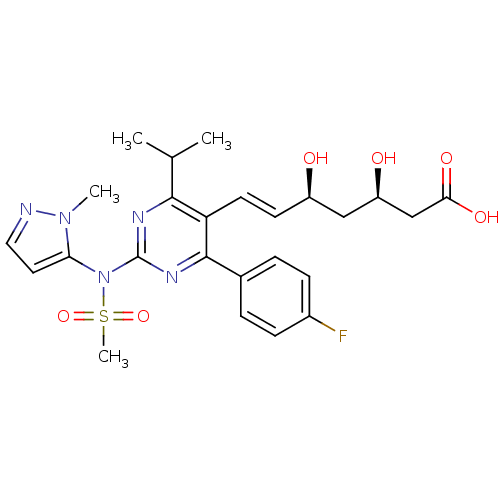
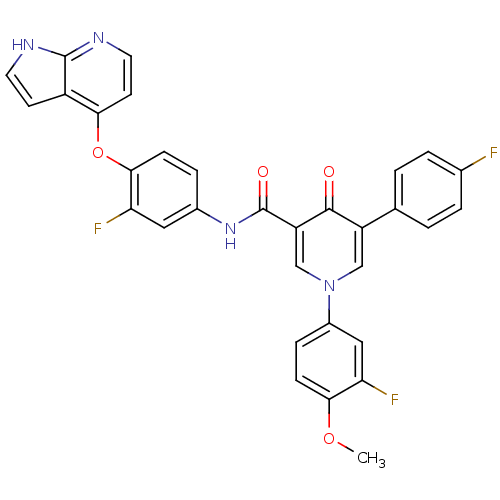
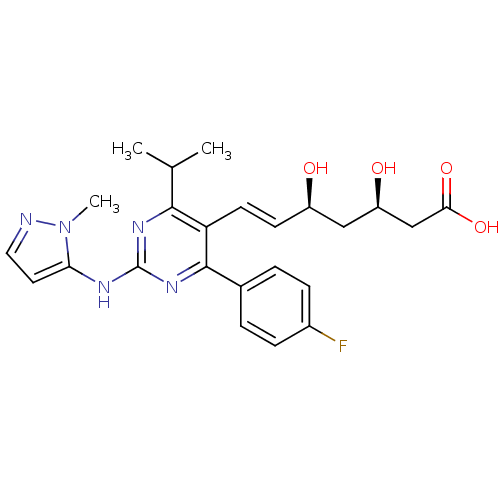
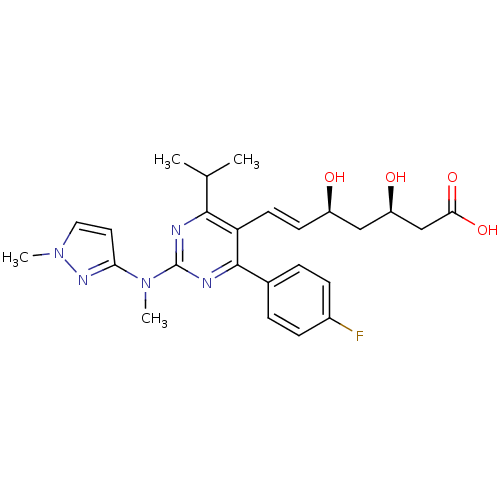
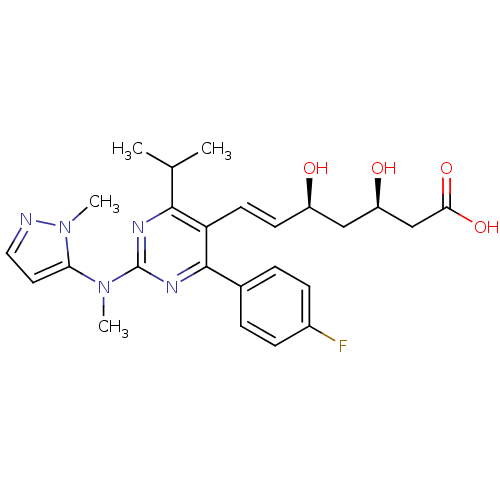
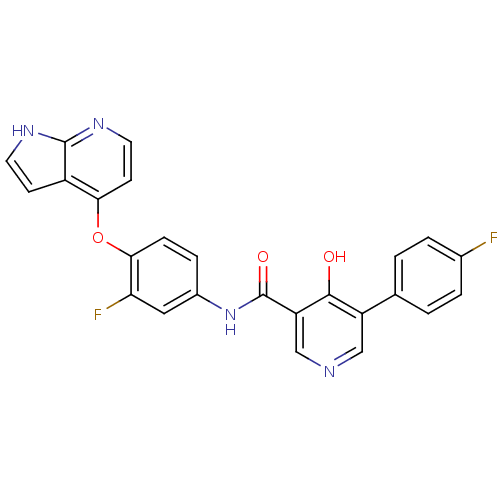
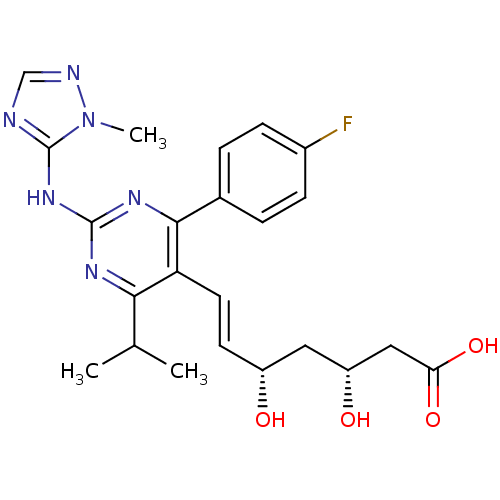
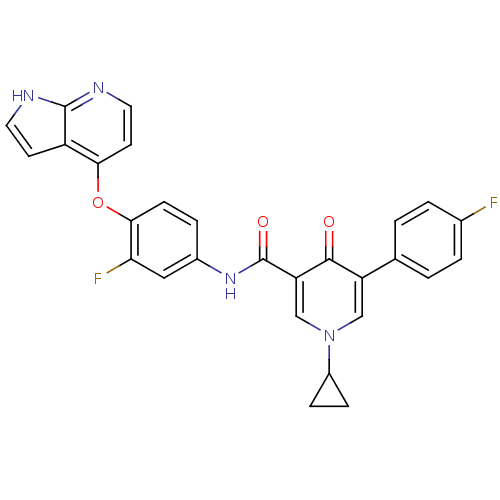
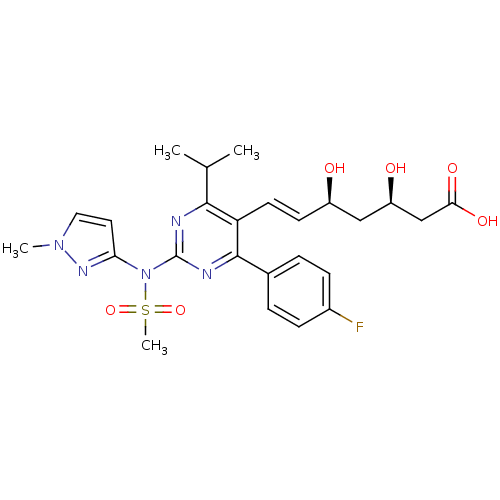
Affinity DataKi: 0.700nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 3.20nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 3.70nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 6.10nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 7.80nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 8.10nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 8.5nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataKi: 48nMAssay Description:Binding affinity to human ARMore data for this Ligand-Target Pair
Affinity DataIC50: 0.5nM EC50: 1.40nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nM EC50: 1.5nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nM EC50: 13.7nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMpH: 7.5 T: 2°CAssay Description:Kinase activity was assayed using baculovirus expressed GST-Met, and poly(Glu/Tyr) as the substrate. Dose response curves were generated to determine...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40nM EC50: 3.95nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMpH: 7.5 T: 2°CAssay Description:Kinase activity was assayed using baculovirus expressed GST-Met, and poly(Glu/Tyr) as the substrate. Dose response curves were generated to determine...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMpH: 7.5 T: 2°CAssay Description:Kinase activity was assayed using baculovirus expressed GST-Met, and poly(Glu/Tyr) as the substrate. Dose response curves were generated to determine...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nM EC50: 3.70nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90nMpH: 7.5 T: 2°CAssay Description:Kinase activity was assayed using baculovirus expressed GST-Met, and poly(Glu/Tyr) as the substrate. Dose response curves were generated to determine...More data for this Ligand-Target Pair
TargetIndoleamine 2,3-dioxygenase 1(Homo sapiens (Human))
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells preincubated for 2 hrs followed by IFNgamma stimulation and measured after 18 hrs byMore data for this Ligand-Target Pair
TargetIndoleamine 2,3-dioxygenase 1(Homo sapiens (Human))
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells preincubated for 2 hrs followed by IFNgamma stimulation and measured after 18 hrs byMore data for this Ligand-Target Pair
TargetIndoleamine 2,3-dioxygenase 1(Mus musculus)
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated mouse M109 cells preincubated for 2 hrs followed by IFNgamma stimulation and measured after 18 hrs byMore data for this Ligand-Target Pair
TargetIndoleamine 2,3-dioxygenase 1(Homo sapiens (Human))
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated human HeLa cells preincubated for 2 hrs followed by IFNgamma stimulation and measured after 18 hrs byMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nM EC50: 5.40nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMpH: 7.5 T: 2°CAssay Description:Kinase activity was assayed using baculovirus expressed GST-Met, and poly(Glu/Tyr) as the substrate. Dose response curves were generated to determine...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30nM EC50: 1.40nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40nM EC50: 1.30nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 2.5nM EC50: 1.20nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 2.5nM EC50: 2.5nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80nMpH: 7.5 T: 2°CAssay Description:Kinase activity was assayed using baculovirus expressed GST-Met, and poly(Glu/Tyr) as the substrate. Dose response curves were generated to determine...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90nM EC50: 7.5nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
TargetIndoleamine 2,3-dioxygenase 1(Mus musculus)
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Bristol Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of IDO1 in IFNgamma-stimulated mouse M109 cells preincubated for 2 hrs followed by IFNgamma stimulation and measured after 18 hrs byMore data for this Ligand-Target Pair
Affinity DataIC50: 3nM EC50: 3.70nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10nMpH: 7.5 T: 2°CAssay Description:Kinase activity was assayed using baculovirus expressed GST-Met, and poly(Glu/Tyr) as the substrate. Dose response curves were generated to determine...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10nM EC50: 0.600nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMpH: 7.5 T: 2°CAssay Description:Kinase activity was assayed using baculovirus expressed GST-Met, and poly(Glu/Tyr) as the substrate. Dose response curves were generated to determine...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70nM EC50: 7.70nMpH: 7.0 T: 2°CAssay Description:Enzyme Assay for HMG-CoA reductase was based on the conversion of isotopically labeled HMG-CoA to mevalonic acid using rat liver microsomes as enzyme...More data for this Ligand-Target Pair

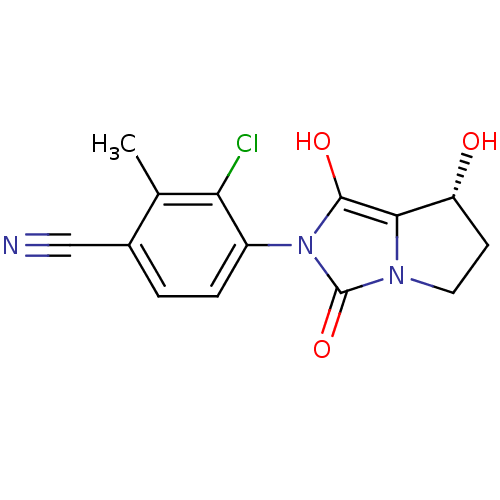

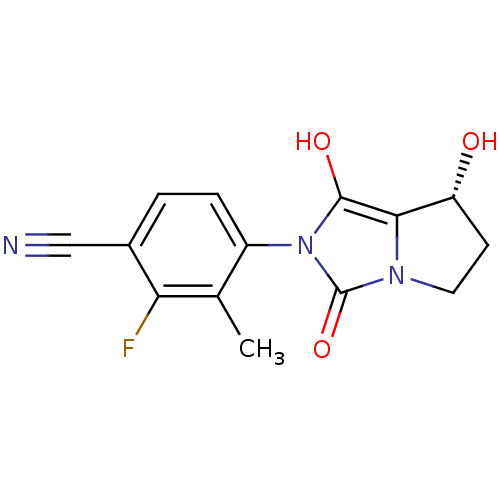


 3D Structure (crystal)
3D Structure (crystal)