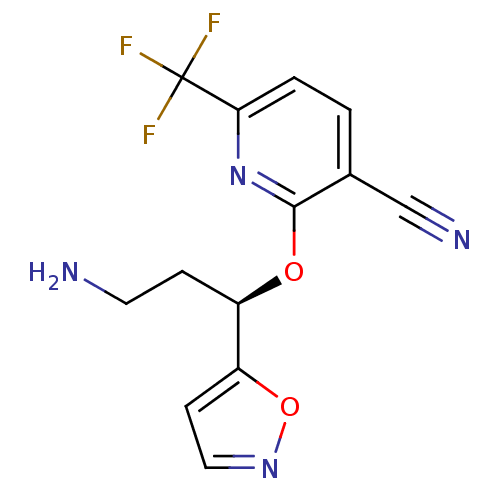
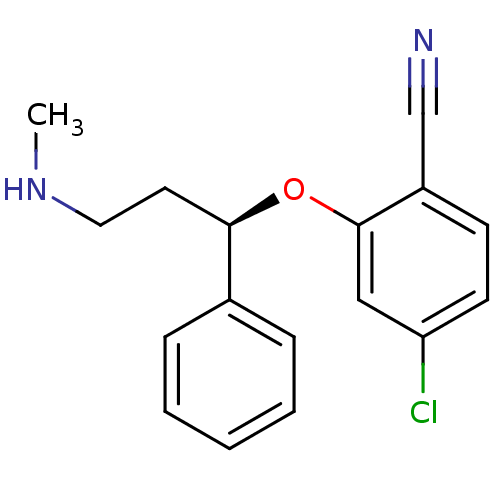
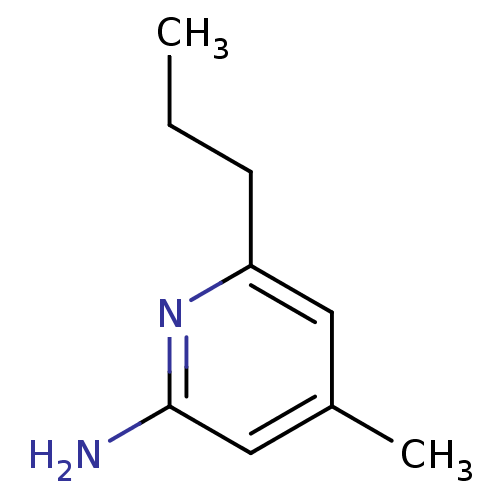
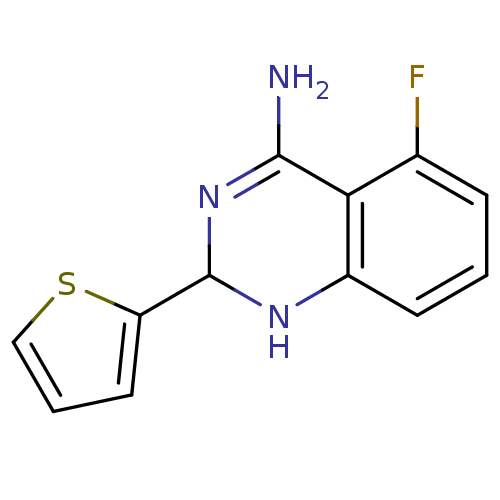
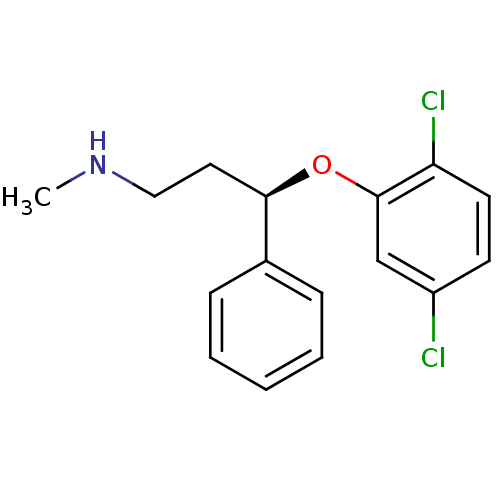
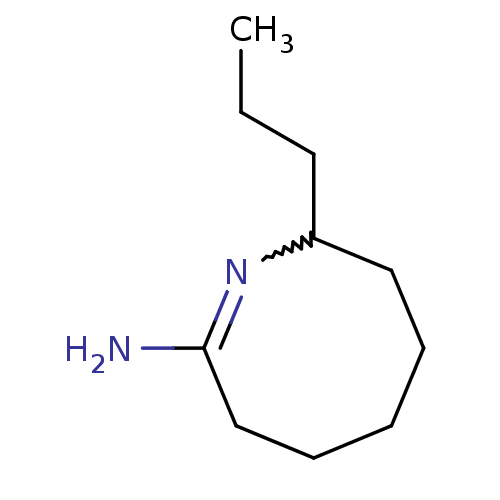
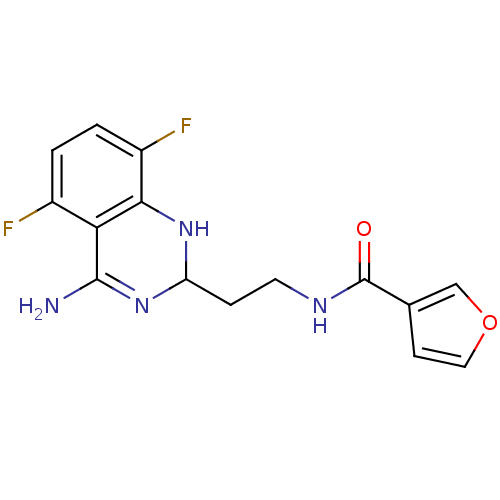
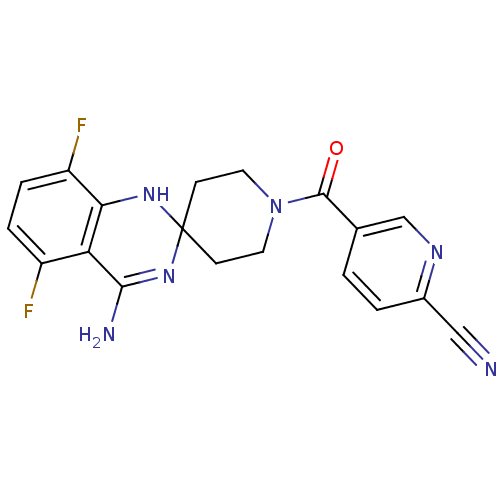
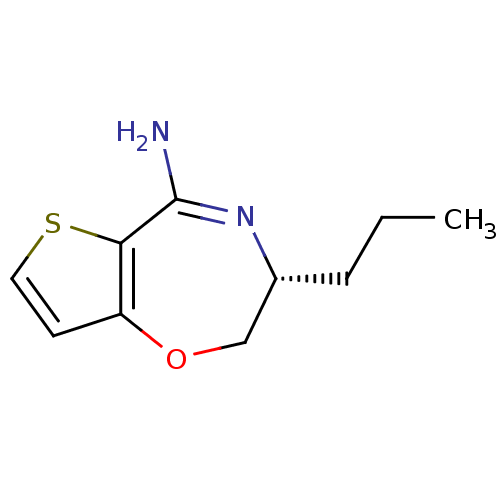
Affinity DataKi: 0.100nM ΔG°: -56.5kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.280nM ΔG°: -54.0kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.290nM ΔG°: -53.9kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.330nM ΔG°: -53.6kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.350nM ΔG°: -53.4kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.420nM ΔG°: -53.0kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.680nM ΔG°: -51.8kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.790nM ΔG°: -51.4kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nM ΔG°: -50.6kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nM ΔG°: -48.9kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 2.70nM ΔG°: -48.4kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 3.20nM ΔG°: -48.0kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 3.5nM ΔG°: -47.8kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 3.90nM ΔG°: -47.5kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 4.70nM ΔG°: -47.1kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 4.80nM ΔG°: -47.0kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 4.90nM ΔG°: -47.0kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 5.30nM ΔG°: -46.8kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 5.5nM ΔG°: -46.7kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.5kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.5kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 6.5nM ΔG°: -46.3kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 6.60nM ΔG°: -46.2kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 8.70nM ΔG°: -45.5kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 9.20nM ΔG°: -45.4kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 12nM ΔG°: -44.8kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 13nM ΔG°: -44.6kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 16nM ΔG°: -44.0kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 26nM ΔG°: -42.9kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 37nM ΔG°: -42.0kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human recombinant iNOSMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human recombinant iNOSMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human recombinant iNOSMore data for this Ligand-Target Pair
Affinity DataIC50: 5.40nMAssay Description:Inhibition of human recombinant iNOSMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of human recombinant iNOSMore data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of human recombinant iNOSMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro inhibition of human Inducible nitric oxide synthase.More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.0 T: 2°CAssay Description:Nitric oxide formation from NOS was monitored by the hemoglobin capture assay. The assay was initiated by addition of enzyme and was monitored at 401...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro inhibition of human neuronal nitric oxide synthase.More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 7.0 T: 2°CAssay Description:Nitric oxide formation from NOS was monitored by the hemoglobin capture assay. The assay was initiated by addition of enzyme and was monitored at 401...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of human recombinant iNOSMore data for this Ligand-Target Pair
Affinity DataIC50: 23nMpH: 7.0 T: 2°CAssay Description:Nitric oxide formation from NOS was monitored by the hemoglobin capture assay. The assay was initiated by addition of enzyme and was monitored at 401...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:In vitro inhibition of human Inducible nitric oxide synthase.More data for this Ligand-Target Pair
Affinity DataIC50: 31nMpH: 7.0 T: 2°CAssay Description:Nitric oxide formation from NOS was monitored by the hemoglobin capture assay. The assay was initiated by addition of enzyme and was monitored at 401...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMpH: 7.0 T: 2°CAssay Description:Nitric oxide formation from NOS was monitored by the hemoglobin capture assay. The assay was initiated by addition of enzyme and was monitored at 401...More data for this Ligand-Target Pair
Affinity DataIC50: 35nMpH: 7.0 T: 2°CAssay Description:Nitric oxide formation from NOS was monitored by the hemoglobin capture assay. The assay was initiated by addition of enzyme and was monitored at 401...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:In vitro inhibition of human Inducible nitric oxide synthase.More data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 7.0 T: 2°CAssay Description:Nitric oxide formation from NOS was monitored by the hemoglobin capture assay. The assay was initiated by addition of enzyme and was monitored at 401...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 7.0 T: 2°CAssay Description:Nitric oxide formation from NOS was monitored by the hemoglobin capture assay. The assay was initiated by addition of enzyme and was monitored at 401...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMAssay Description:Inhibition of human recombinant iNOSMore data for this Ligand-Target Pair



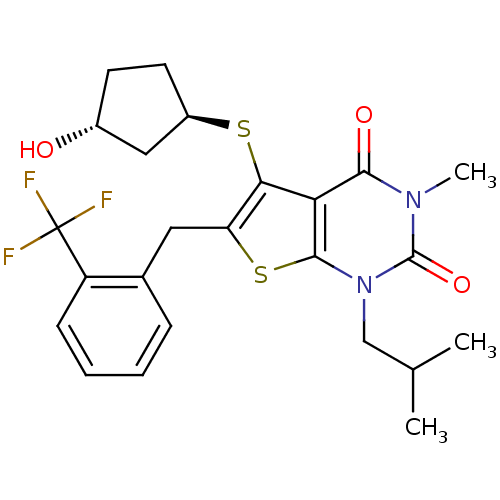
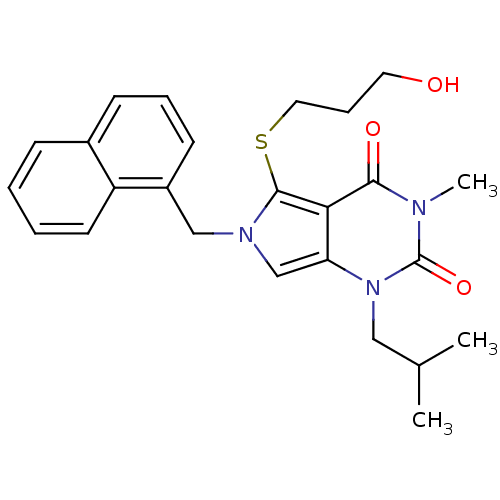
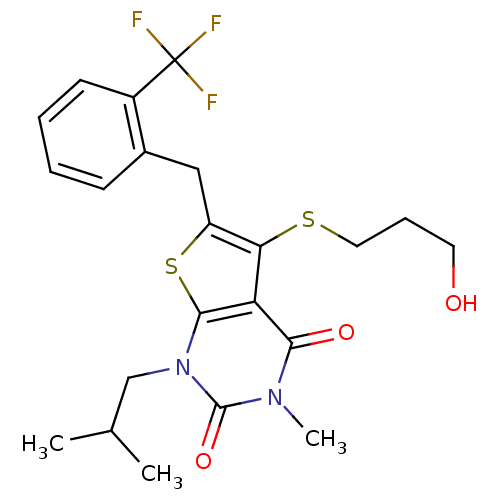
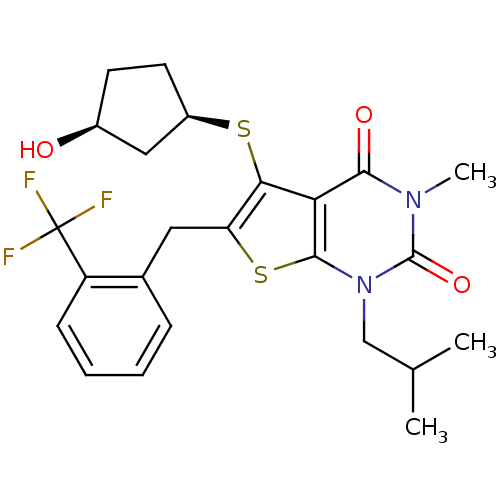



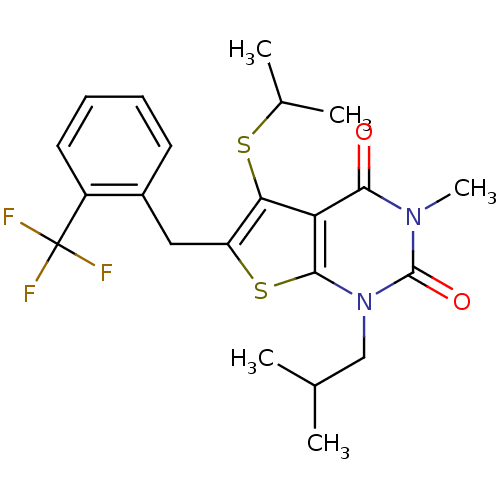
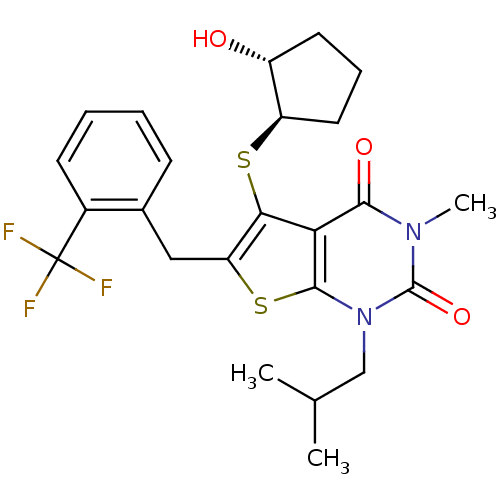
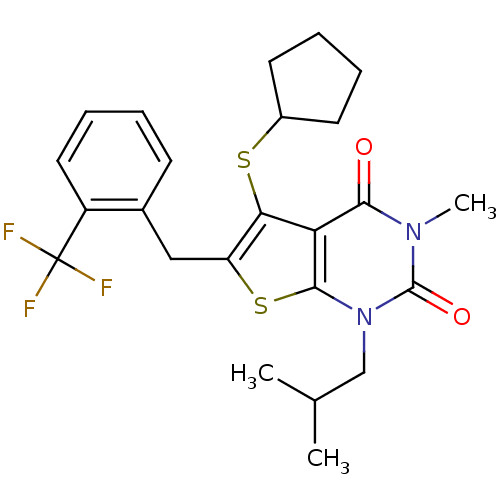
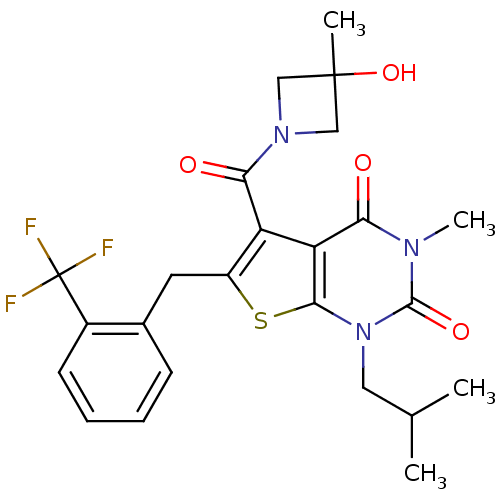

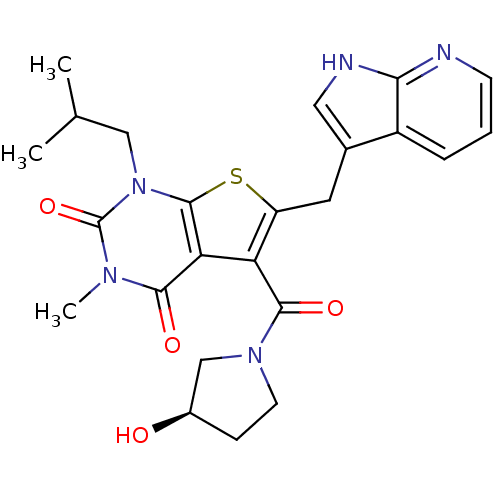


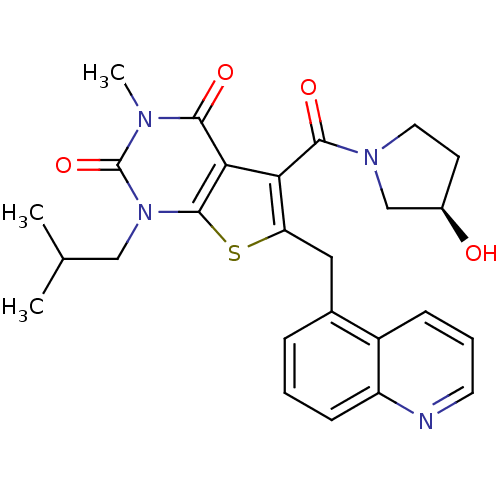
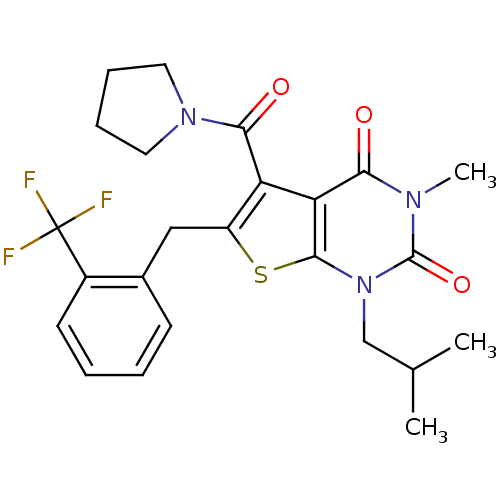



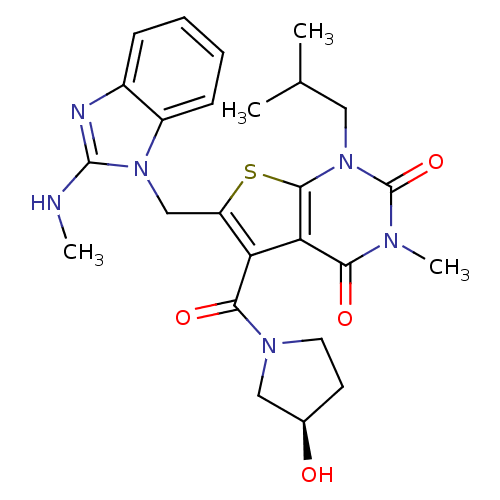
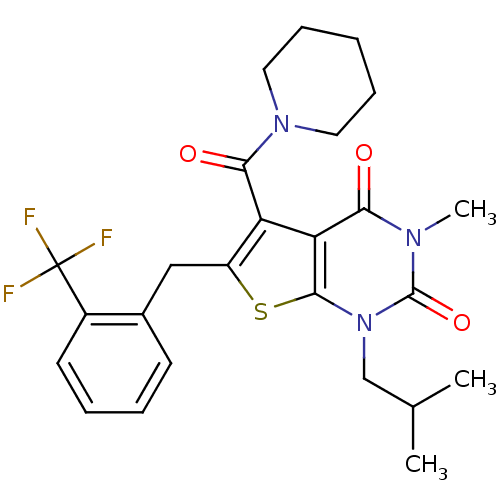
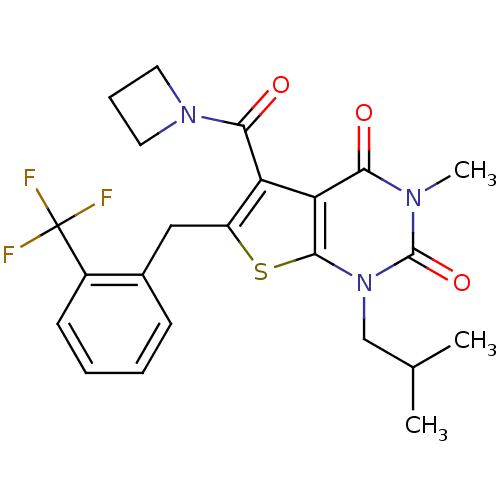
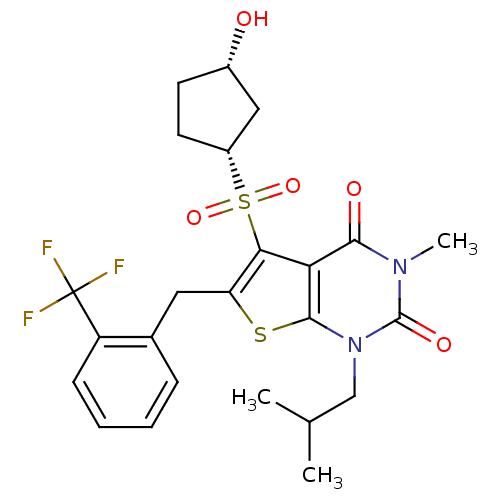


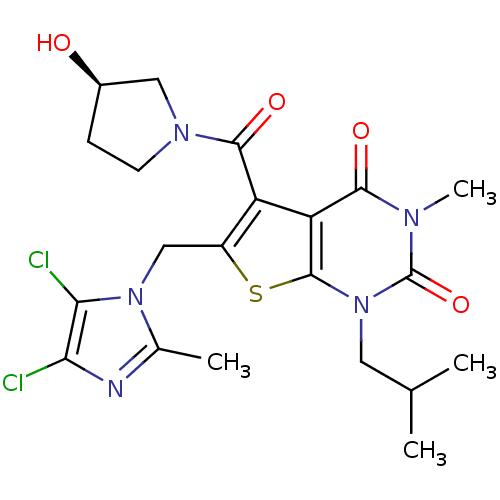
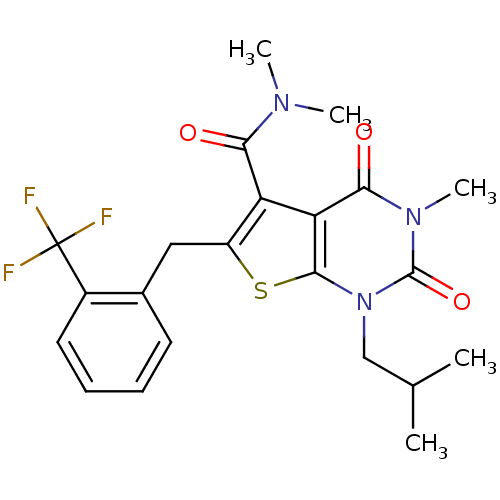

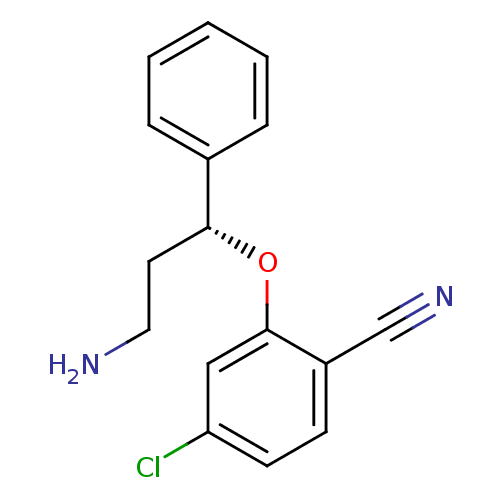
 3D Structure (crystal)
3D Structure (crystal)