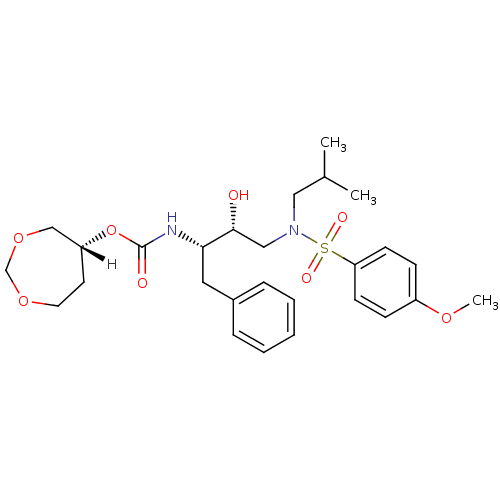
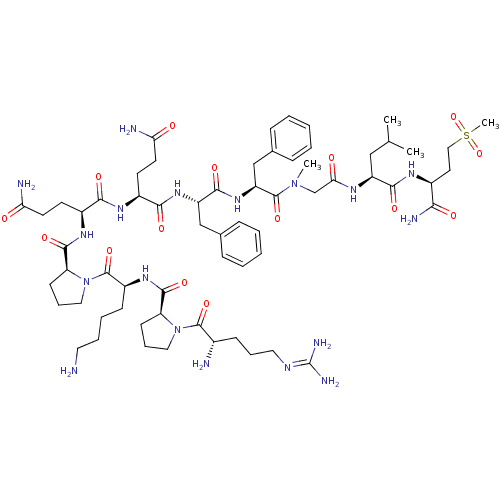
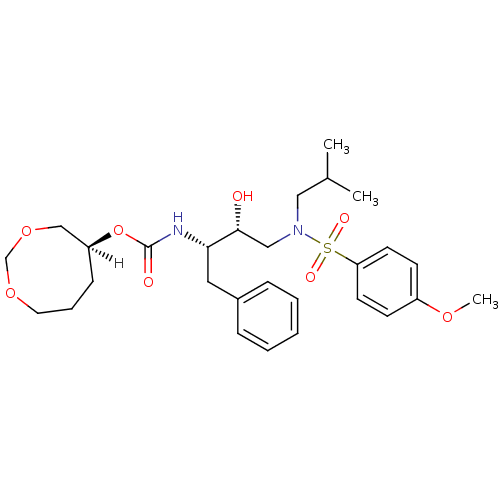
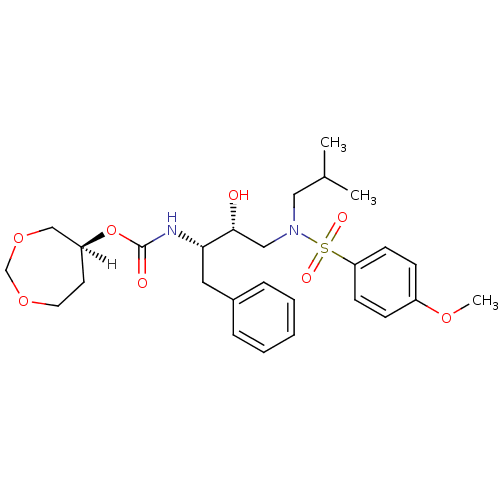
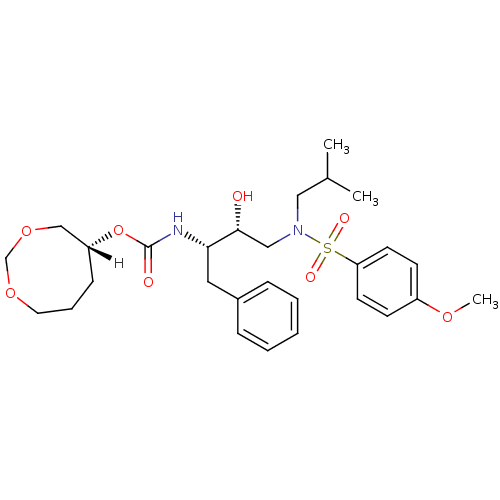
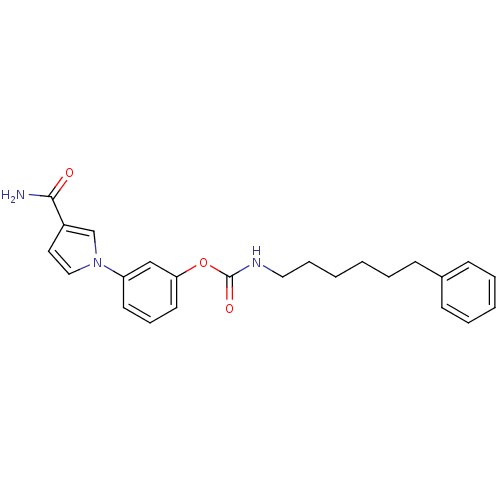
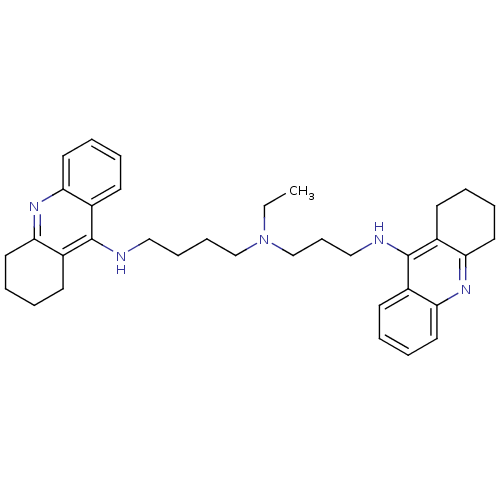
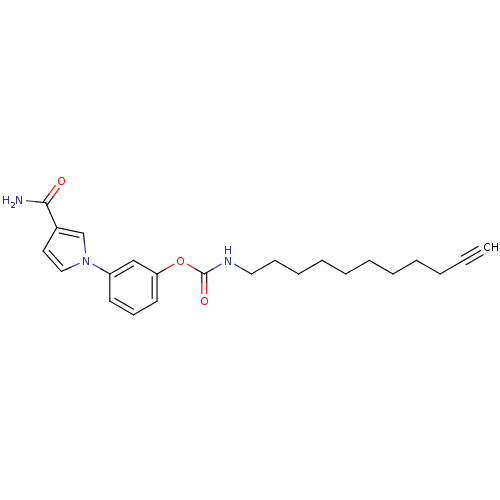
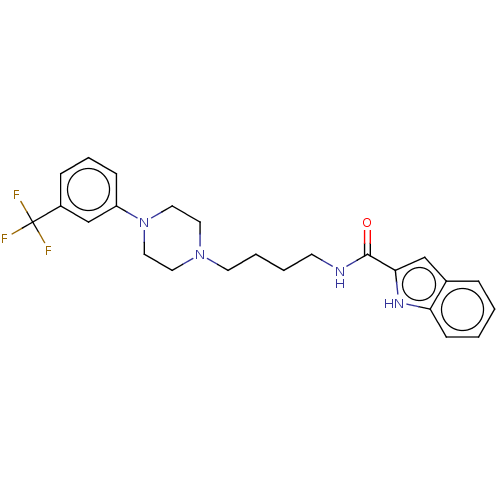
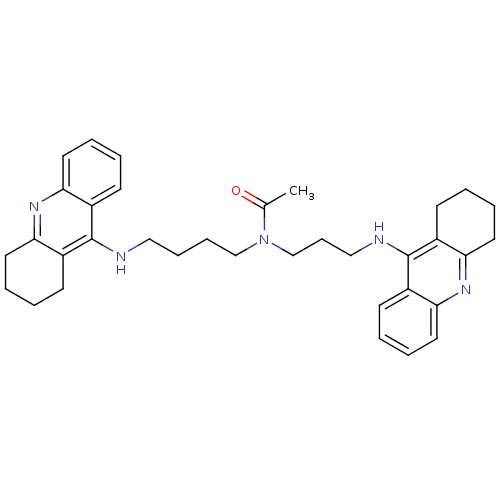
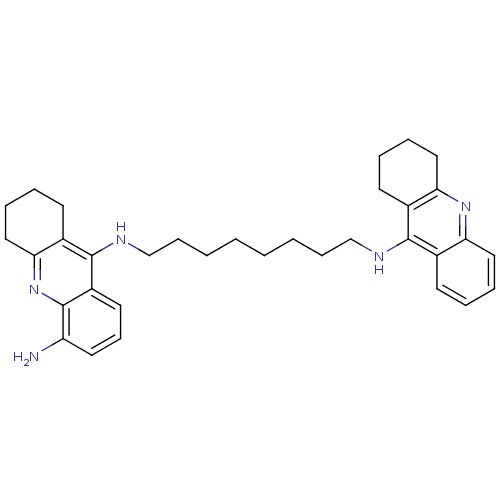
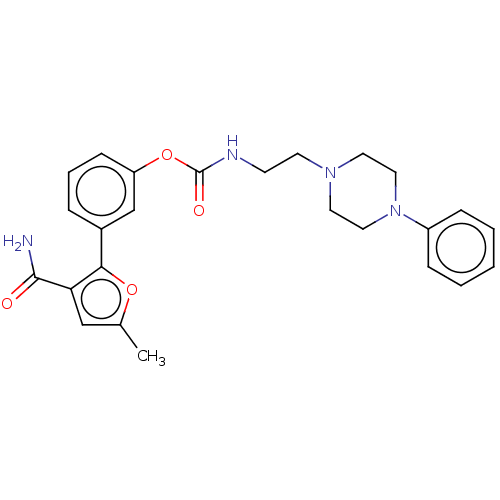
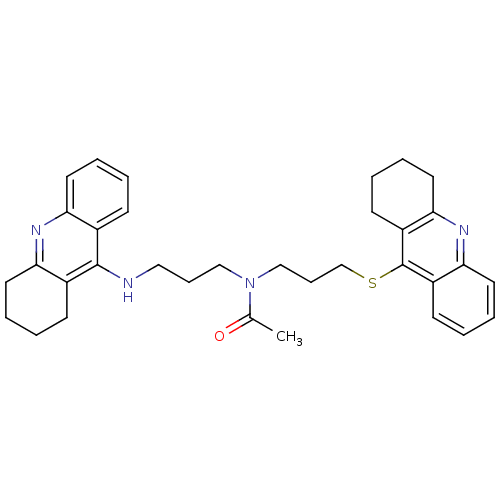
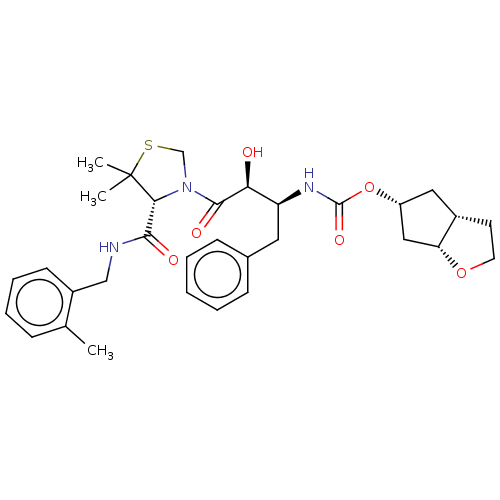
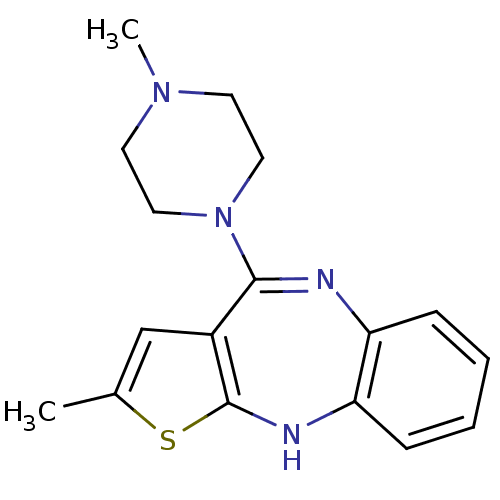
Affinity DataKi: 0.00520nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
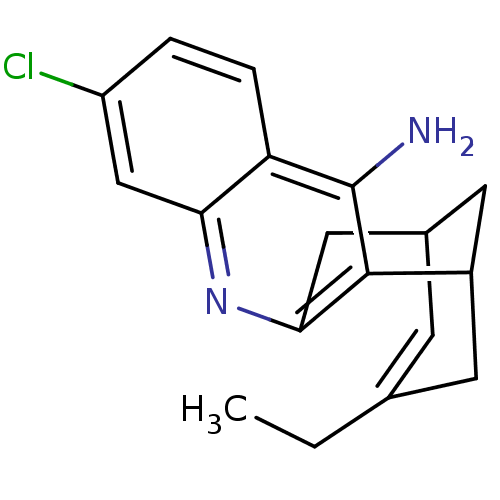
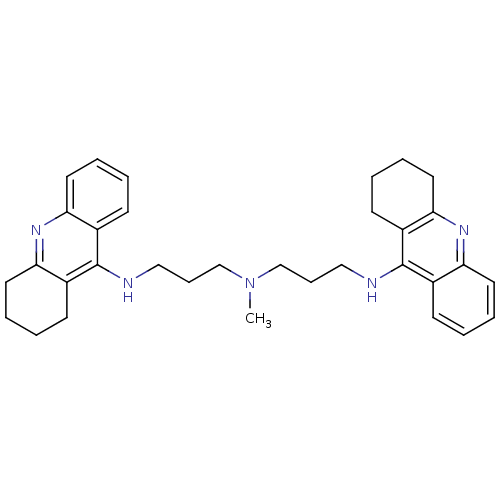
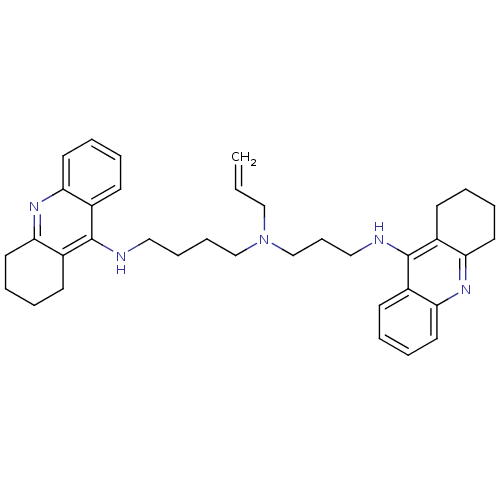
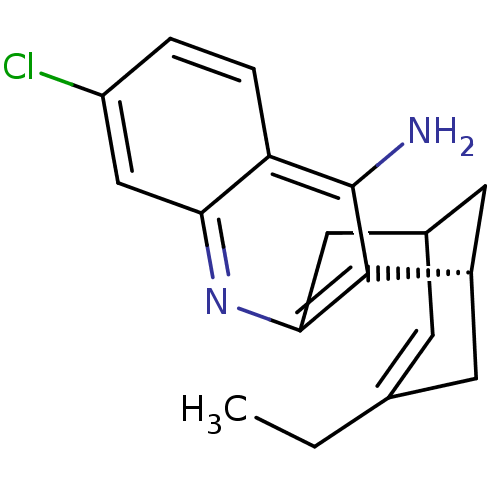
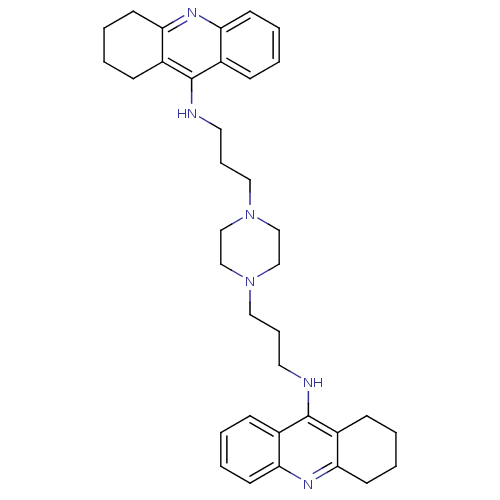
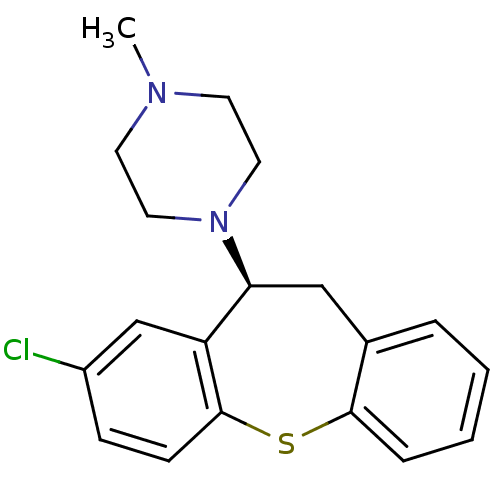
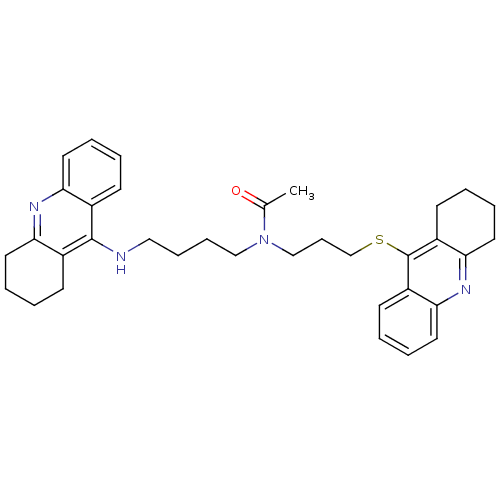
Affinity DataKi: 0.0120nMAssay Description:Inhibition of human acetylcholine esteraseMore data for this Ligand-Target Pair
Affinity DataKi: 0.0120nMAssay Description:Inhibition of human recombinant AChEMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universit£
Curated by ChEMBL
Universit£
Curated by ChEMBL
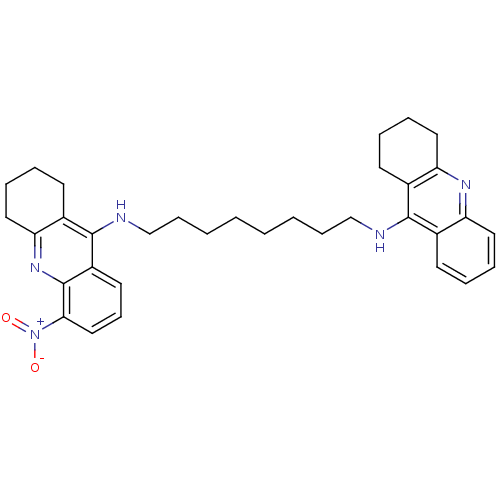
Affinity DataKi: 0.0150nMAssay Description:Inhibition of Torpedo californica AChEMore data for this Ligand-Target Pair
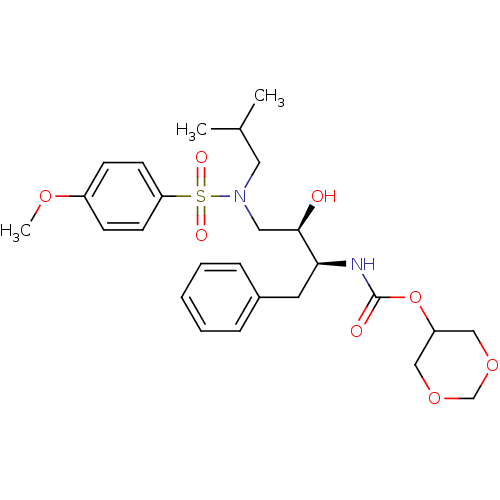
Affinity DataKi: 0.0160nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Purdue University
Purdue University
Affinity DataKi: 0.0260nM ΔG°: -60.4kJ/mole IC50: 4.90nMpH: 6.4 T: 2°CAssay Description:The Ki values were determined by substrate cleavage assay using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-Arg-NH2. A standar...More data for this Ligand-Target Pair
Affinity DataKi: 0.0260nM ΔG°: -60.4kJ/molepH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by E llman. Inhibition of enzyme activity was measured over a substrate c...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Purdue University
Purdue University
Affinity DataKi: 0.0410nM ΔG°: -59.3kJ/mole IC50: 3.40nMpH: 6.4 T: 2°CAssay Description:The Ki values were determined by substrate cleavage assay using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-Arg-NH2. A standar...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:Inhibition of human butyrylcholine esteraseMore data for this Ligand-Target Pair
Affinity DataKi: 0.0600nM ΔG°: -58.3kJ/molepH: 8.0 T: 2°CAssay Description:Inhibition of enzyme activity was measured over a substrate concentration range of 0.01-30 mM and at least six inhibitor concentrations to determine ...More data for this Ligand-Target Pair
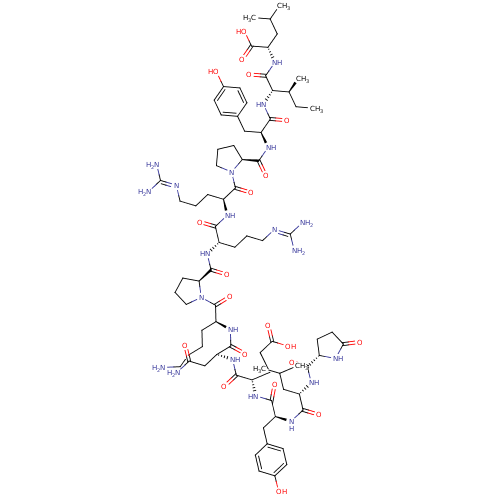
Affinity DataKi: 0.0670nMAssay Description:Binding affinity to human VPAC1 receptor by radioligand displacement assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0760nMAssay Description:Binding affinity to human neuropeptide Y receptor type 2 by radioligand displacement assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universit£
Curated by ChEMBL
Universit£
Curated by ChEMBL
Affinity DataKi: 0.0770nMAssay Description:Inhibition of Torpedo californica AChEMore data for this Ligand-Target Pair
Affinity DataKi: 0.0850nMAssay Description:Inhibition of human recombinant AChEMore data for this Ligand-Target Pair
Affinity DataKi: 0.0960nMAssay Description:Binding affinity to human neuropeptide Y receptor type 1 by radioligand displacement assayMore data for this Ligand-Target Pair
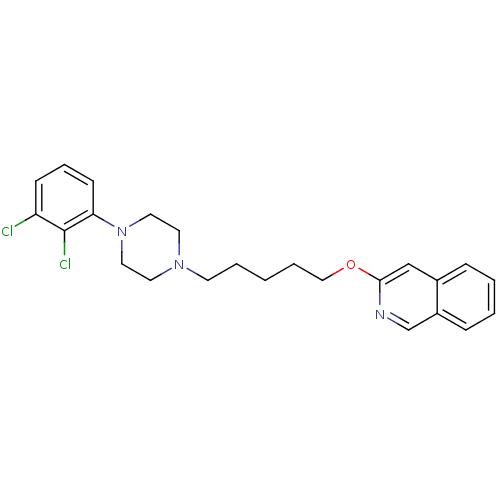
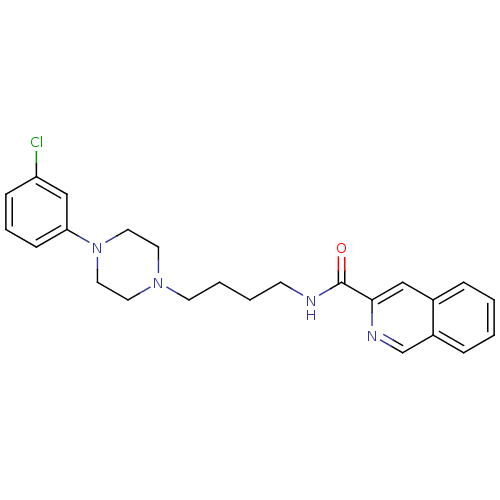
Affinity DataKi: 0.0990nMAssay Description:Displacement of [3H]7OH-DPAT from dopamine D3 receptor (unknown origin) expressed in Sf9 cells by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 0.105nMAssay Description:Displacement of [3H]7OH-DPAT from dopamine D3 receptor (unknown origin) expressed in Sf9 cells by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 0.110nMAssay Description:Inhibition of enzyme activity was measured over a substrate concentration range of 0.01-30 mM and at least six inhibitor concentrations to determine ...More data for this Ligand-Target Pair
Affinity DataKi: 0.119nMAssay Description:Inhibition of human recombinant AChEMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Universit£
Curated by ChEMBL
Universit£
Curated by ChEMBL
Affinity DataKi: 0.130nMAssay Description:Inhibition of Torpedo californica AChEMore data for this Ligand-Target Pair
Affinity DataKi: 0.136nMAssay Description:Inhibition of human recombinant AChEMore data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Binding affinity to human NK1 receptor by radioligand displacement assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Displacement of [3H]7OH-DPAT from dopamine D3 receptor (unknown origin) expressed in Sf9 cells by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Half-maximal inhibition of [3H]ketanserin binding to 5-hydroxytryptamine 2A receptor in rat cerebral cortex homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:In vitro binding affinity towards 5-hydroxytryptamine 2A receptor in rat tissue homogenate using [3H]ketanserin as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Binding affinity to 5HT2A receptor (unknown origin)More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Purdue University
Purdue University
Affinity DataKi: 0.150nM ΔG°: -56.1kJ/molepH: 6.4 T: 2°CAssay Description:The Ki values were determined by substrate cleavage assay using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-Arg-NH2. A standar...More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Displacement of [3H]ketanserin from 5HT2A receptor in CRL:CD(SD)BR-COBS rat cortex by scintillation spectrometryMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Purdue University
Purdue University
Affinity DataKi: 0.160nM ΔG°: -55.9kJ/mole IC50: 30nMpH: 6.4 T: 2°CAssay Description:The Ki values were determined by substrate cleavage assay using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-Arg-NH2. A standar...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Purdue University
Purdue University
Affinity DataKi: 0.160nM ΔG°: -55.9kJ/molepH: 6.4 T: 2°CAssay Description:The Ki values were determined by substrate cleavage assay using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-Arg-NH2. A standar...More data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Inhibition of mouse FAAH isolated from brain homogenate using [3H-ethanolamine]AEA as substrate incubated for 20 mins prior to substrate addition mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.162nMAssay Description:Inhibition of human recombinant AChEMore data for this Ligand-Target Pair
Affinity DataKi: 0.180nMAssay Description:Displacement of [3H]-7-OH-DPAT from rat brain membrane D3 receptor expressed in Sf9 cells incubated for 60 mins by liquid scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataKi: 0.180nMAssay Description:Inhibition of mouse FAAH isolated from brain homogenate using [3H-ethanolamine]AEA as substrate incubated for 20 mins prior to substrate addition mea...More data for this Ligand-Target Pair
Affinity DataKi: 0.180nMAssay Description:Displacement of [3H]7OH-DPAT from dopamine D3 receptor (unknown origin) expressed in Sf9 cells by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 0.180nMAssay Description:Binding affinity for rat striatum Dopamine receptor D3 (sf9 cells) by [3H]7-OH-DPAT displacement.More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Binding affinity to dopamine D3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.210nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.223nMAssay Description:Inhibition of human recombinant AChEMore data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Inhibition of human acetylcholine esteraseMore data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Inhibition of FAAH in mouse brain membranes assessed as inhibitory constant using [14C]-AEA as substrate incubated for 15 mins by scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Inhibition of enzyme activity was measured over a substrate concentration range of 0.01-30 mM and at least six inhibitor concentrations to determine ...More data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Half-maximal inhibition of [3H]ketanserin binding to 5-hydroxytryptamine 2A receptor in rat cerebral cortex homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:In vitro binding affinity towards Dopamine receptor D3 in Sf9 cell membranes using [3H]7-OH-DPAT as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 0.270nMAssay Description:Displacement of [3H]7OH-DPAT from dopamine D3 receptor (unknown origin) expressed in Sf9 cells by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 0.270nMAssay Description:Inhibition of human recombinant BuChEMore data for this Ligand-Target Pair
Affinity DataKi: 0.270nMAssay Description:Binding affinity to human NTS1 receptor by radioligand displacement assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.290nMAssay Description:Inhibition of HIV1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity to dopamine D3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Half-maximal inhibition of [3H]7-OH-DPAT binding to Histamine H1 receptor in rat tissue homogenateMore data for this Ligand-Target Pair

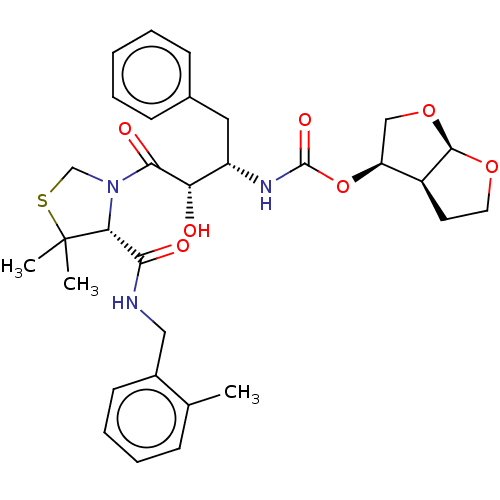
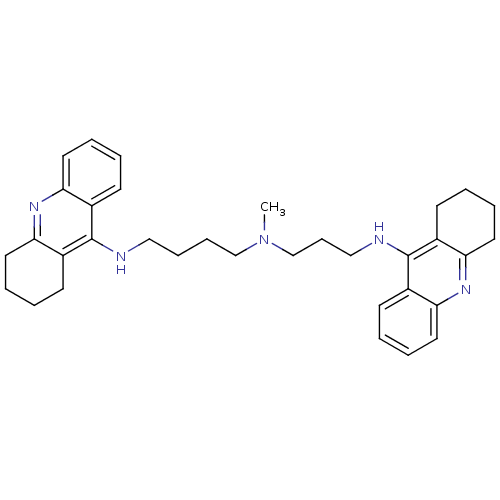

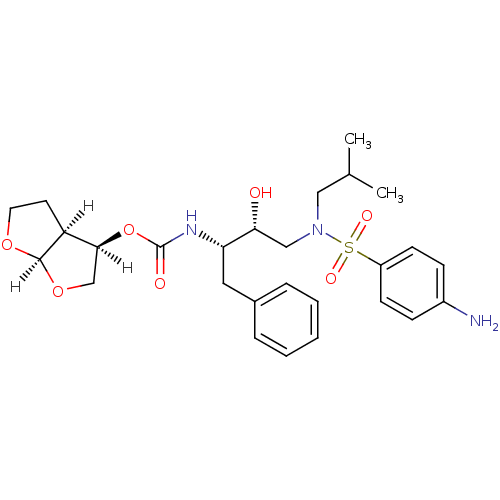
 3D Structure (crystal)
3D Structure (crystal)