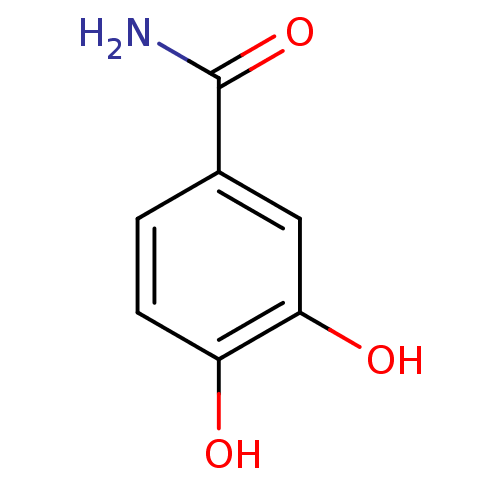
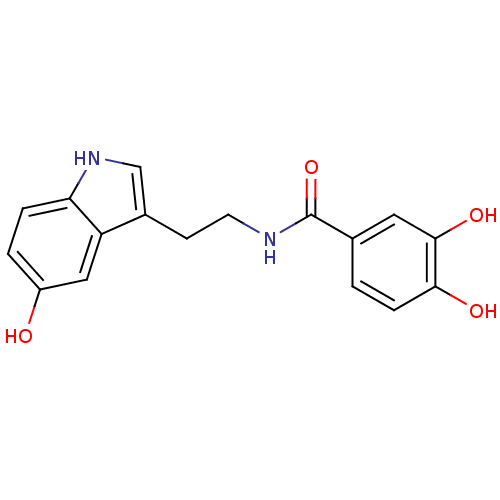
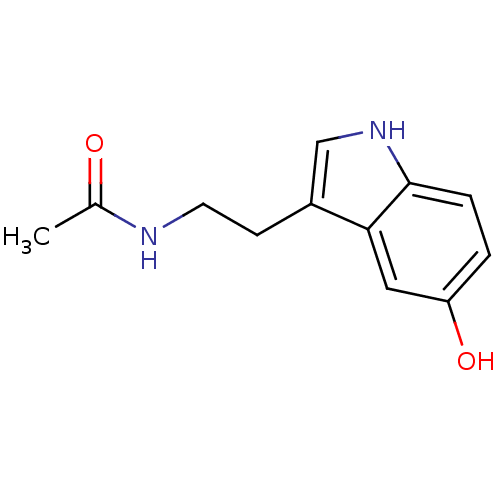
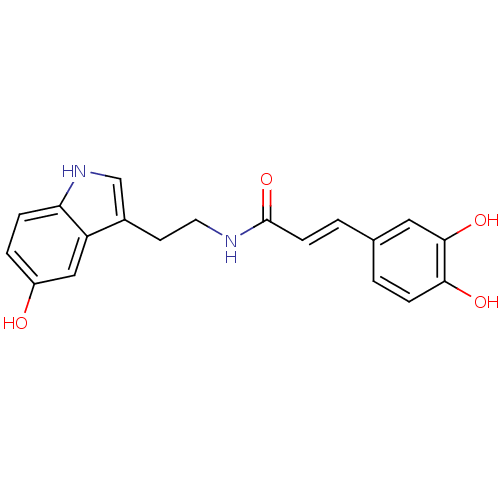
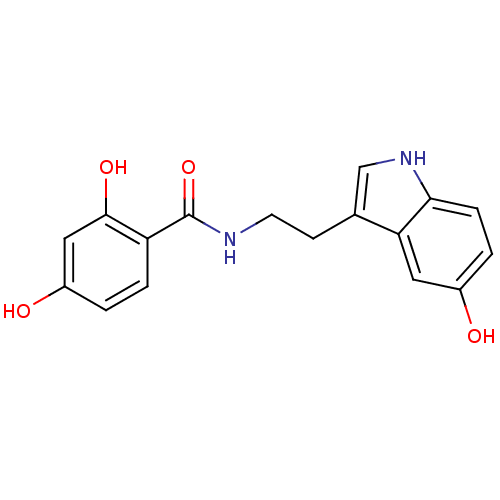
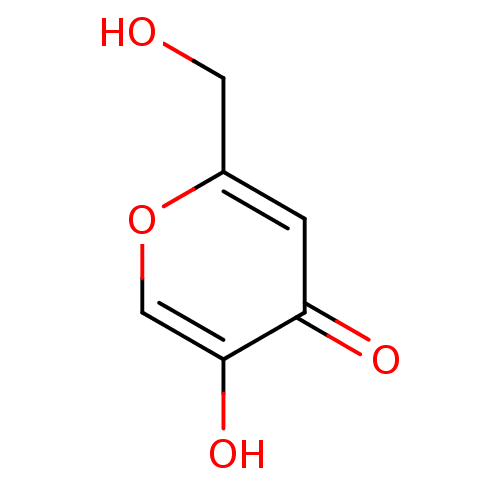
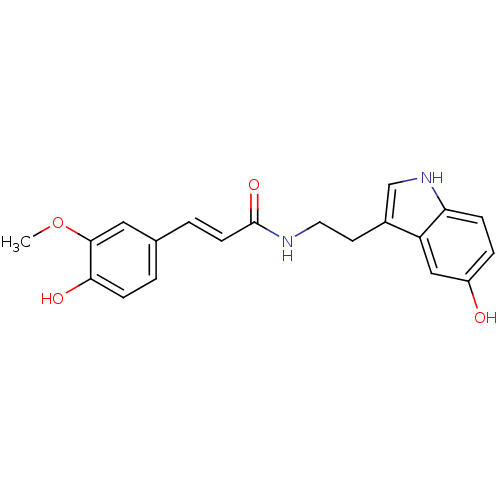
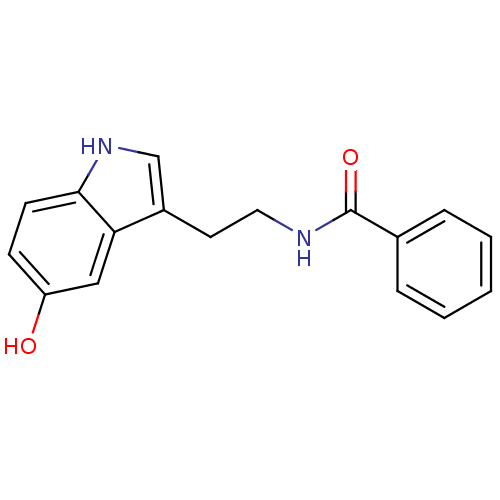
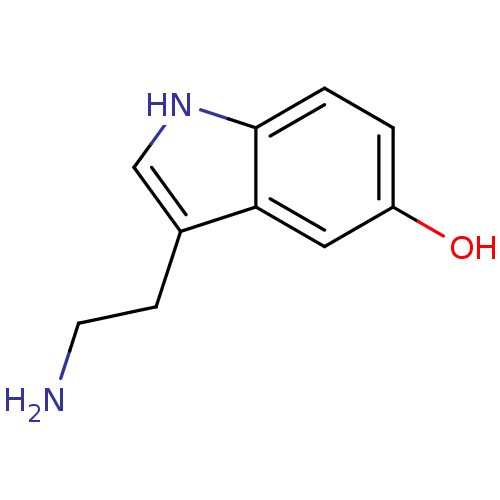
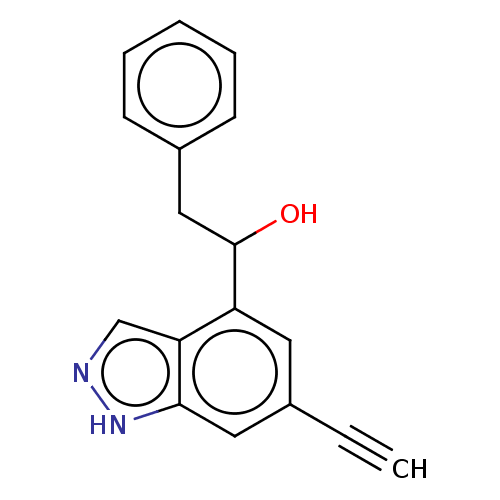
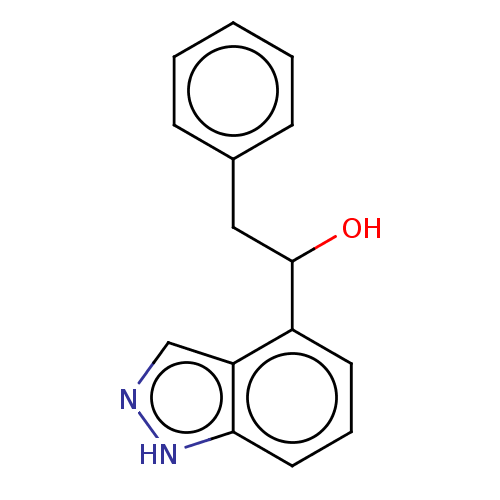
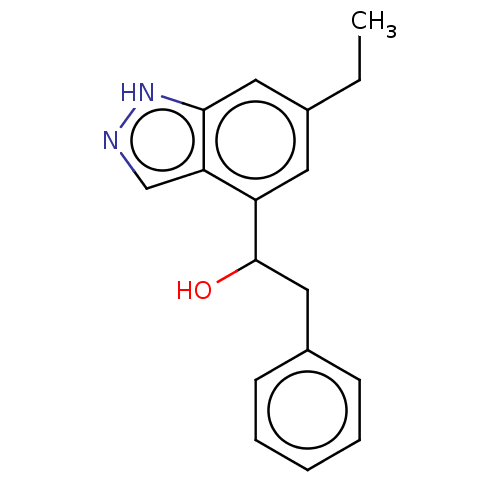
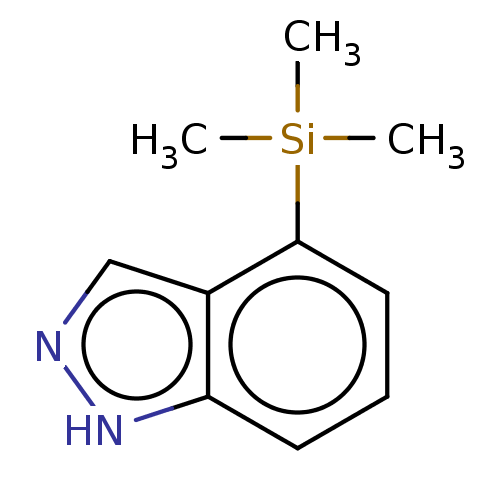
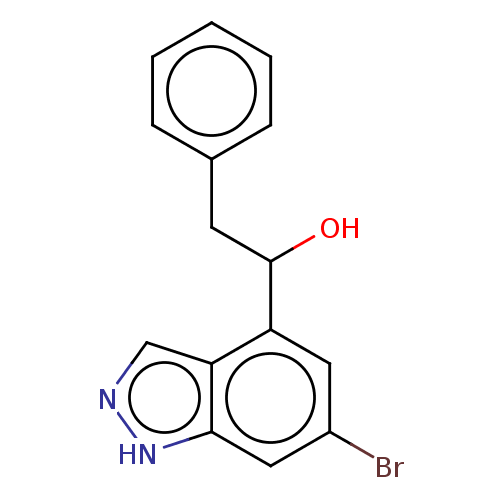
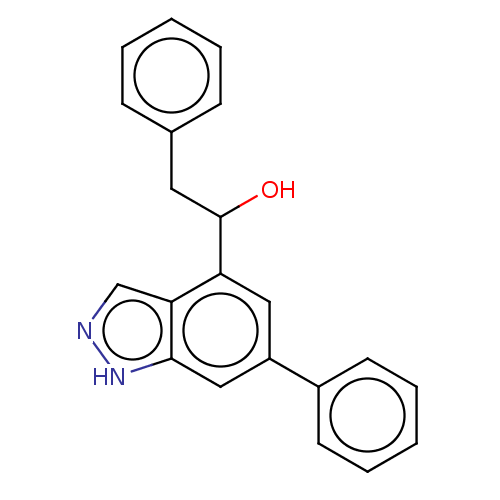
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataKi: 3.70E+3nMAssay Description:Apparent noncompetitive inhibition of catecholase activity of tyrosinase in mouse B16 cells by Lineweaver-Burke plot analysisMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataKi: 6.40E+4nMAssay Description:Apparent noncompetitive inhibition of catecholase activity of tyrosinase in mouse B16 cells by Lineweaver-Burke plot analysisMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataKi: 2.40E+5nMAssay Description:Apparent noncompetitive inhibition of catecholase activity of tyrosinase in mouse B16 cells by Lineweaver-Burke plot analysisMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 4.20E+4nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 6.10E+4nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 6.80E+4nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 7.30E+4nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 2.11E+5nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 2.16E+5nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 2.33E+5nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 3.11E+5nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 3.33E+5nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
TargetTyrosinase(Mus musculus (Mouse))
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
National Institute Of Advanced Industrial Science And Technology
Curated by ChEMBL
Affinity DataIC50: 5.50E+5nMAssay Description:Inhibition of catecholase activity of tyrosinase in mouse B16 cells assessed as dopachrome formationMore data for this Ligand-Target Pair
Affinity DataKd: 2.80E+4nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 C129Y mutant (unknown origin) expressed in Escherichia coli assessed as dissociation constant i...More data for this Ligand-Target Pair
Affinity DataKd: 8.10E+4nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 3.60E+5nMAssay Description:Binding affinity to recombinant ferric state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for 3...More data for this Ligand-Target Pair
Affinity DataKd: 7.40E+3nMAssay Description:Binding affinity to recombinant ferric state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for 3...More data for this Ligand-Target Pair
Affinity DataKd: 7.90E+3nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 740nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 4.40E+4nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 C129Y mutant (unknown origin) expressed in Escherichia coli assessed as dissociation constant i...More data for this Ligand-Target Pair
Affinity DataKd: 3.10E+4nMAssay Description:Binding affinity to recombinant ferric state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for 3...More data for this Ligand-Target Pair
Affinity DataKd: 470nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 1.50E+5nMAssay Description:Binding affinity to recombinant ferric state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for 3...More data for this Ligand-Target Pair
Affinity DataKd: 2.20E+3nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 C129Y mutant (unknown origin) expressed in Escherichia coli assessed as dissociation constant i...More data for this Ligand-Target Pair
Affinity DataKd: 600nMAssay Description:Binding affinity to recombinant ferric state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for 3...More data for this Ligand-Target Pair
Affinity DataKd: 2.70E+4nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 C129Y mutant (unknown origin) expressed in Escherichia coli assessed as dissociation constant i...More data for this Ligand-Target Pair
Affinity DataKd: 6.60E+4nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 C129Y mutant (unknown origin) expressed in Escherichia coli assessed as dissociation constant i...More data for this Ligand-Target Pair
Affinity DataKd: 1.20E+3nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 5.30E+4nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 1.30E+3nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 140nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 9.40E+4nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 C129Y mutant (unknown origin) expressed in Escherichia coli assessed as dissociation constant i...More data for this Ligand-Target Pair
Affinity DataKd: 1.70E+3nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair
Affinity DataKd: 1.10E+3nMAssay Description:Binding affinity to recombinant ferric state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for 3...More data for this Ligand-Target Pair
Affinity DataKd: 1.20E+3nMAssay Description:Binding affinity to recombinant ferric state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for 3...More data for this Ligand-Target Pair
Affinity DataKd: 2.50E+3nMAssay Description:Binding affinity to recombinant ferric state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for 3...More data for this Ligand-Target Pair
Affinity DataKd: 4.30E+4nMAssay Description:Binding affinity to recombinant ferrous state of IDO1 (unknown origin) expressed in Escherichia coli assessed as dissociation constant incubated for ...More data for this Ligand-Target Pair