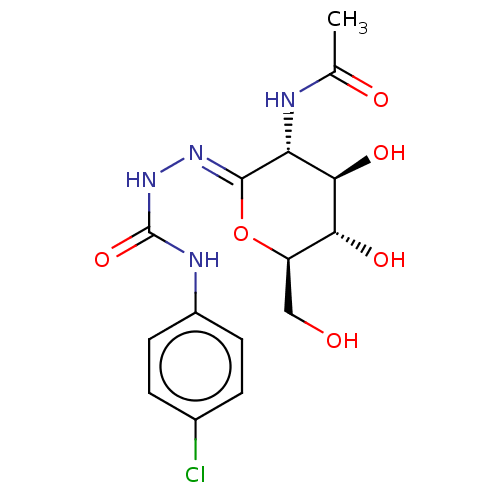
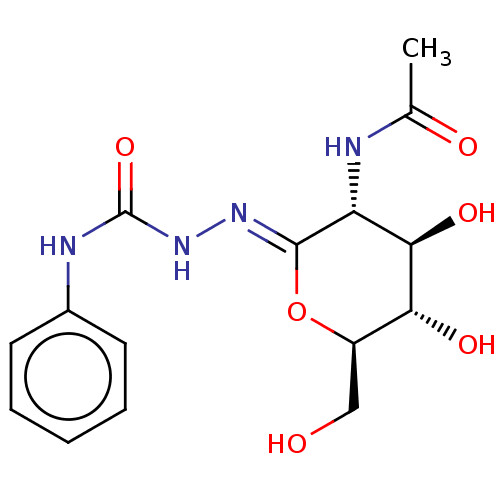
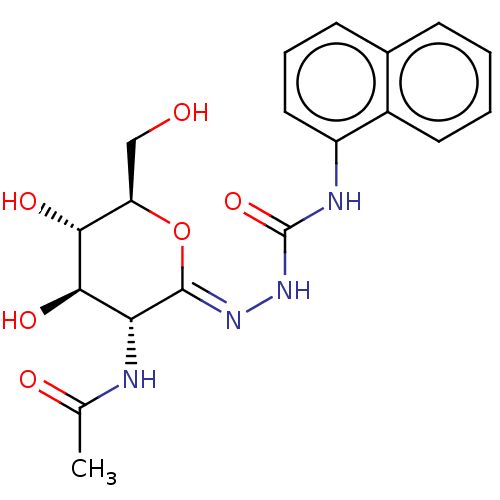
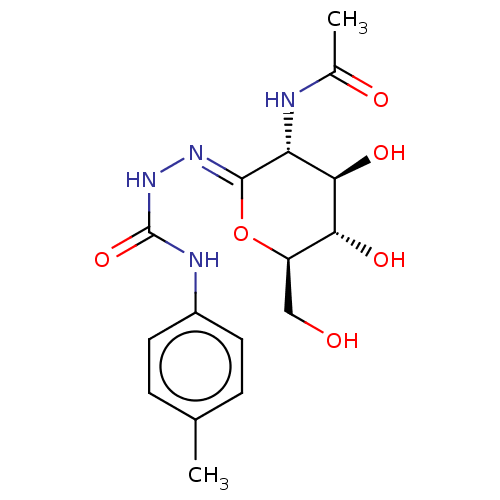
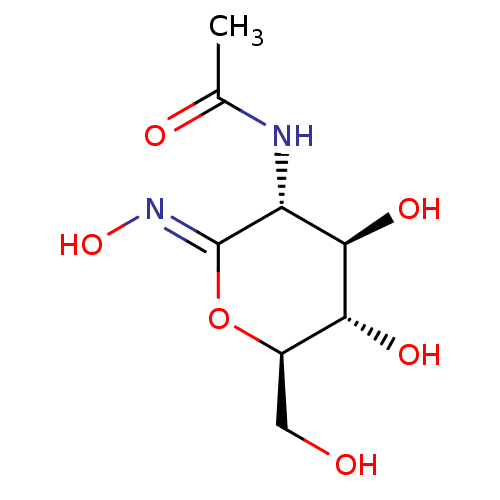
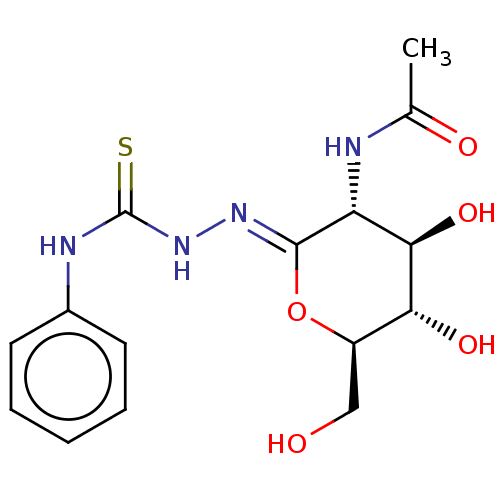
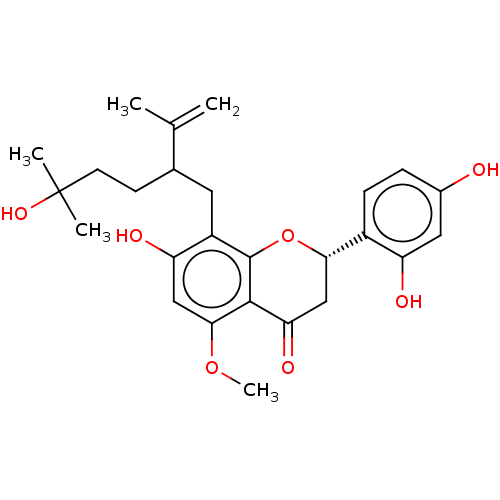
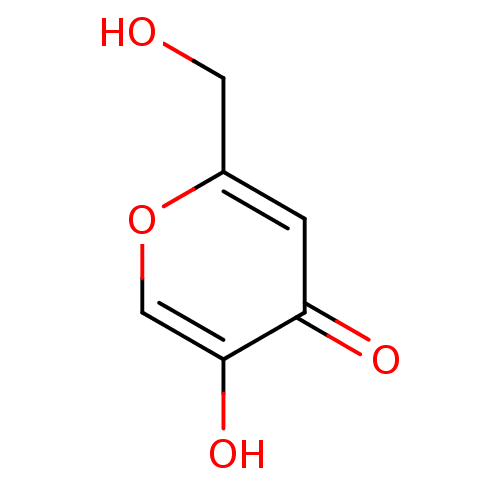
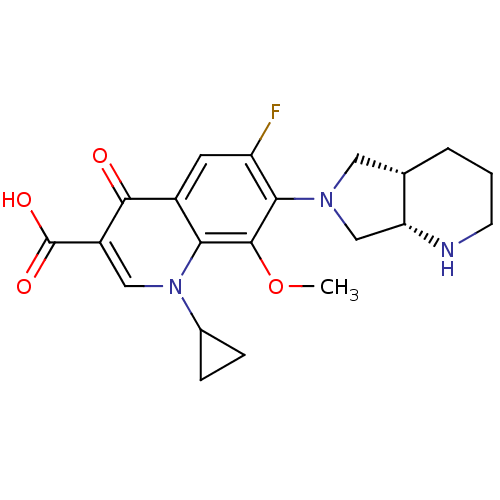
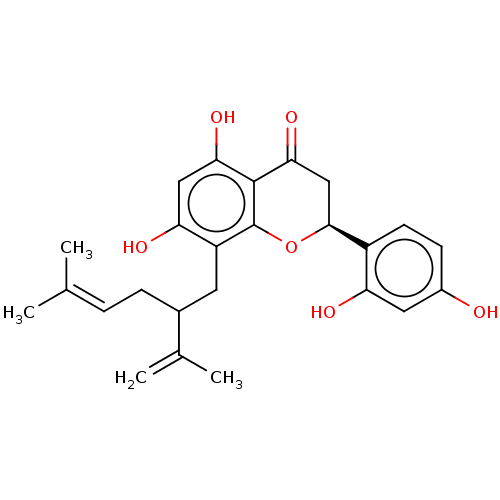
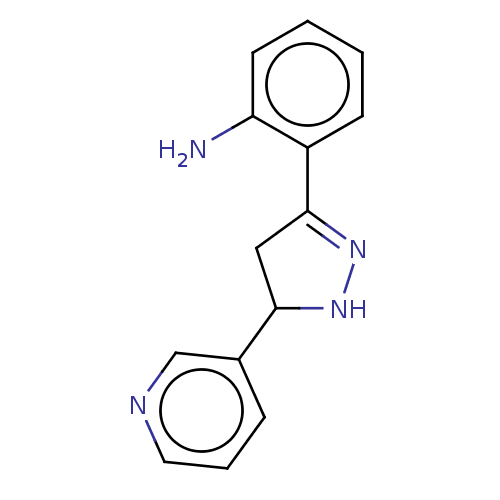
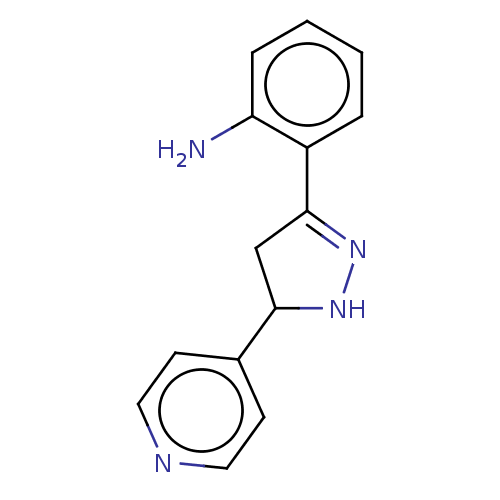
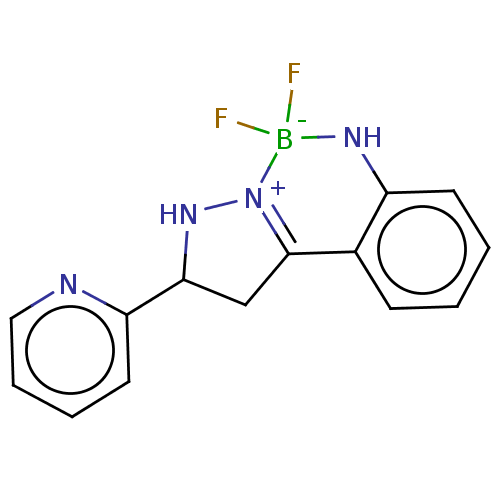
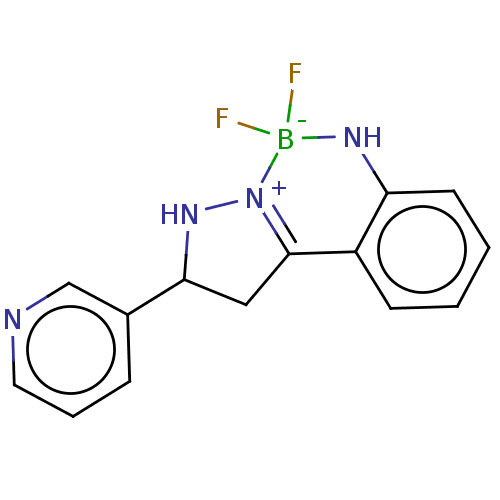
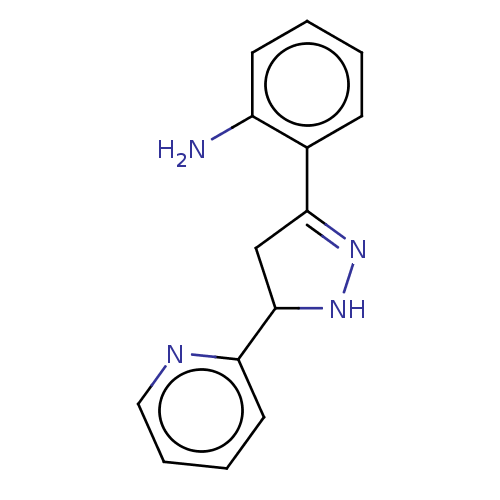
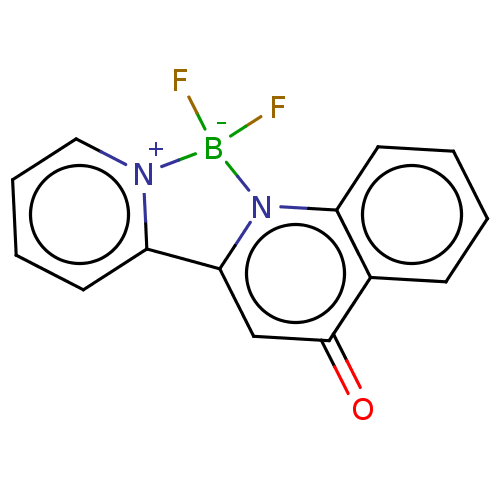
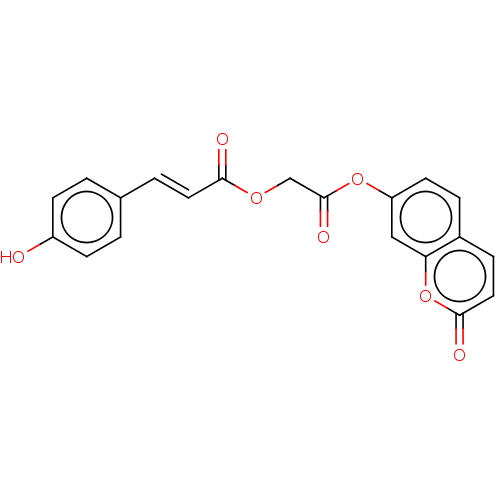
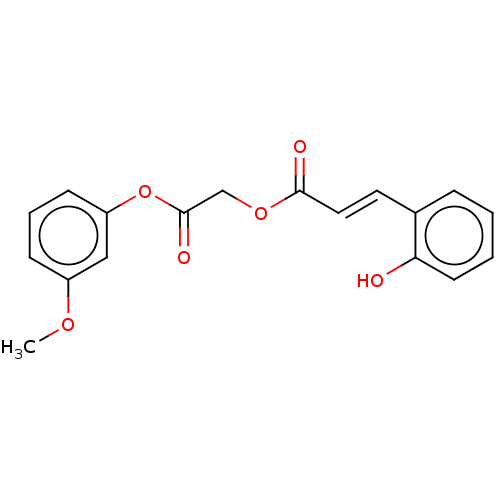
Affinity DataKi: 36nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using 4-Mu-GlcNAc as substrate assessed as inhibition constant by measuring liberation of n...More data for this Ligand-Target Pair
Affinity DataKi: 47nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Inhibition of OGA (unknown origin) using pNP-O-GlcNAc as substrate assessed as inhibition constant incubated for 30 minsMore data for this Ligand-Target Pair
Affinity DataKi: 83nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using 4-Mu-GlcNAc as substrate assessed as inhibition constant by measuring liberation of n...More data for this Ligand-Target Pair
Affinity DataKi: 125nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibition of HexA/HexB (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 154nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 155nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using pNP-GlcNAc as chromogenic substrate assessed as inhibition constant by measuring libe...More data for this Ligand-Target Pair
Affinity DataKi: 167nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using pNP-GlcNAc as chromogenic substrate assessed as inhibition constant by measuring libe...More data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 190nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using pNP-GlcNAc as chromogenic substrate assessed as inhibition constant by measuring libe...More data for this Ligand-Target Pair
Affinity DataKi: 205nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 270nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using pNP-GlcNAc as chromogenic substrate assessed as inhibition constant by measuring libe...More data for this Ligand-Target Pair
Affinity DataKi: 332nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 413nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 506nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using pNP-GlcNAc as chromogenic substrate assessed as inhibition constant by measuring libe...More data for this Ligand-Target Pair
Affinity DataKi: 509nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using 4-Mu-GlcNAc as substrate assessed as inhibition constant by measuring liberation of n...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nMAssay Description:Inhibition of OGA (unknown origin) using pNP-O-GlcNAc as substrate assessed as inhibition constant incubated for 30 minsMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataKi: 1.10E+4nMAssay Description:Non-competitive inhibition of mushroom tyrosinase using L-DOPA as substrate preincubated for 10 mins followed by substrate addition and further incub...More data for this Ligand-Target Pair
Affinity DataKi: 2.70E+4nMAssay Description:Competitive inhibition of N-terminal His-tagged human OGA using pNP-GlcNAc as chromogenic substrate assessed as inhibition constant by measuring libe...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataKi: 1.30E+5nMAssay Description:Non-competitive inhibition of mushroom tyrosinase using L-DOPA as substrate preincubated for 10 mins followed by substrate addition and further incub...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 10 mins by spectrophotometer assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.260nMAssay Description:Inhibition of tyrosinase (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.613nMAssay Description:Inhibition of tyrosinase (unknown origin) using DOPA as substrate incubated for 15 min by spectrophotometer assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 0.700nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.762nMAssay Description:Inhibition of tyrosinase (unknown origin) using DOPA as substrate incubated for 15 min by spectrophotometer assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.856nMAssay Description:Inhibition of tyrosinase (unknown origin) using DOPA as substrate incubated for 15 min by spectrophotometer assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.908nMAssay Description:Inhibition of tyrosinase (unknown origin) using DOPA as substrate incubated for 15 min by spectrophotometer assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 0.960nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 10 mins by spectrophotometer assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 1.20nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 10 mins by spectrophotometer assayMore data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of tyrosinase (unknown origin) using DOPA as substrate incubated for 15 min by spectrophotometer assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: >200nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: >200nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate incubated for 20 mins by microplate reader assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: >200nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate pre-incubated for 10 mins followed by substrate addition and measured after 20 mins by mi...More data for this Ligand-Target Pair
Affinity DataIC50: 315nMAssay Description:Inhibition of tyrosinase (unknown origin) using DOPA as substrate incubated for 15 min by spectrophotometer assayMore data for this Ligand-Target Pair
Affinity DataIC50: 635nMAssay Description:Inhibition of tyrosinase (unknown origin) using DOPA as substrate incubated for 15 min by spectrophotometer assayMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 5.70E+3nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in dopachrome formation using L-DOPA as substrate preincubated for 10 mins followed by substr...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 1.67E+4nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in dopachrome formation using L-DOPA as substrate preincubated for 10 mins followed by substr...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 2.38E+4nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in dopachrome formation using L-DOPA as substrate preincubated for 10 mins followed by substr...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 3.94E+4nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in dopachrome formation using L-DOPA as substrate preincubated for 10 mins followed by substr...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 1.24E+5nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in dopachrome formation using L-DOPA as substrate preincubated for 10 mins followed by substr...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 1.73E+5nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in dopachrome formation using L-DOPA as substrate preincubated for 10 mins followed by substr...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
The University Of Queensland (Uq)
Curated by ChEMBL
The University Of Queensland (Uq)
Curated by ChEMBL
Affinity DataIC50: 1.93E+5nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in dopachrome formation using L-DOPA as substrate preincubated for 10 mins followed by substr...More data for this Ligand-Target Pair

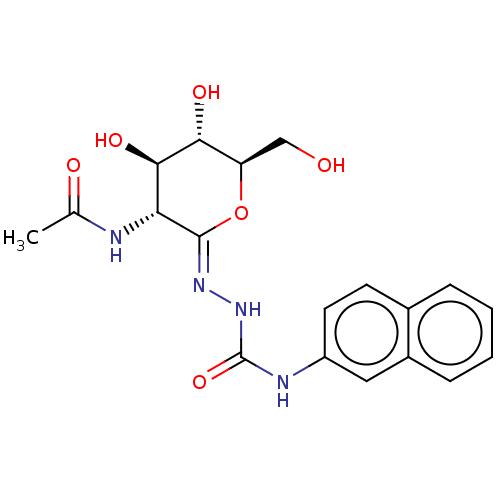

 3D Structure (crystal)
3D Structure (crystal)