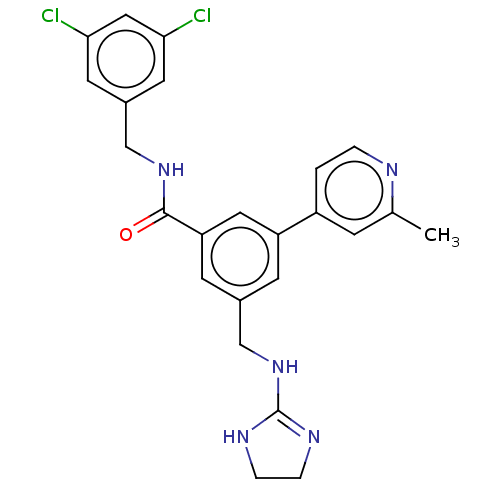
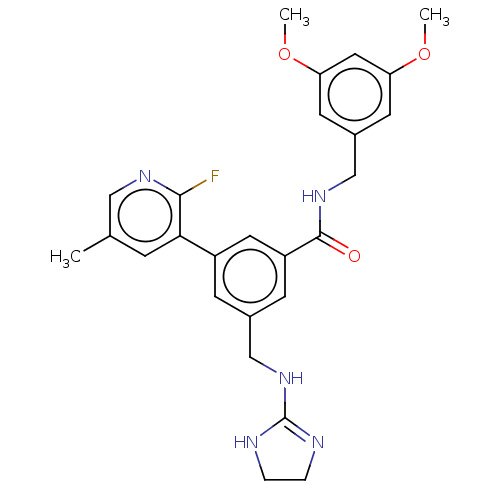
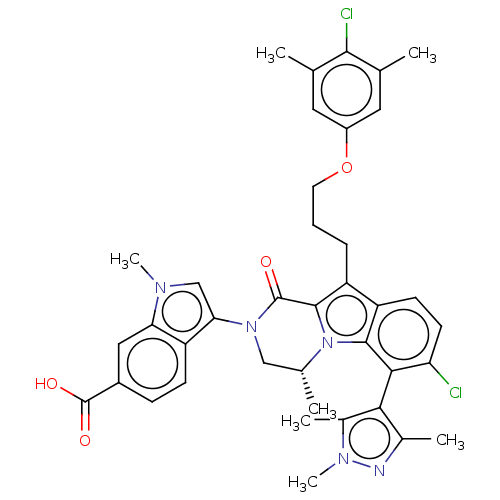
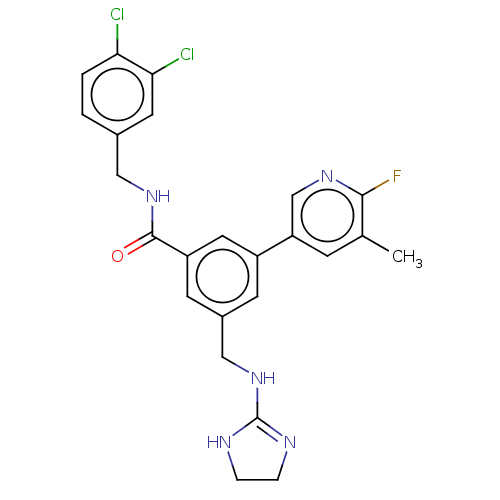
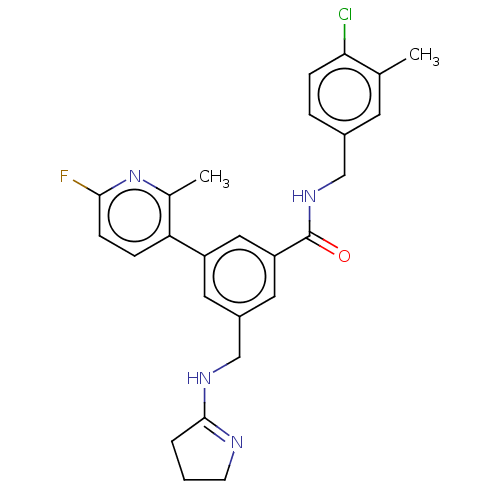
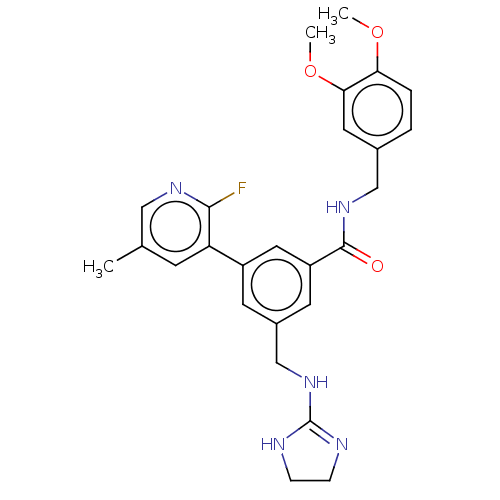
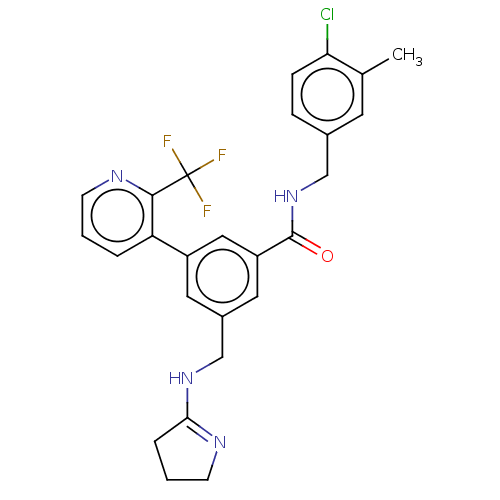
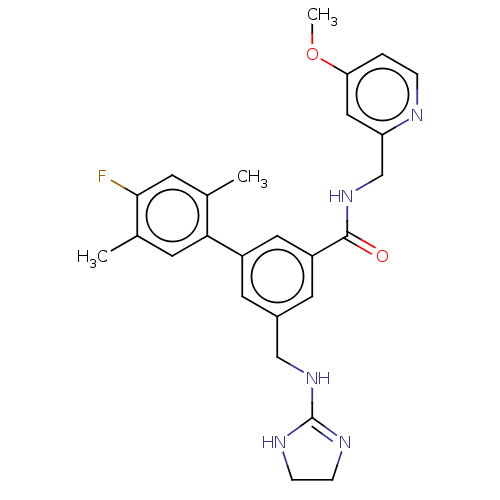
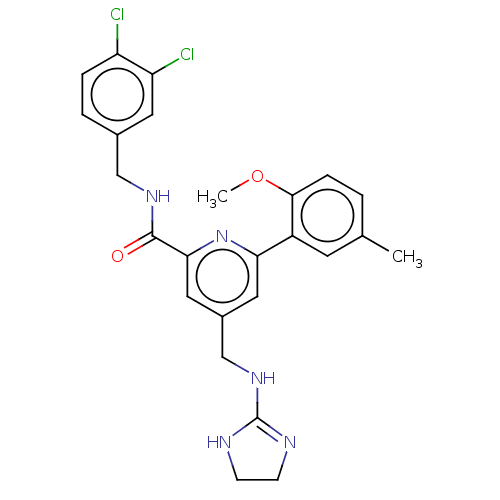
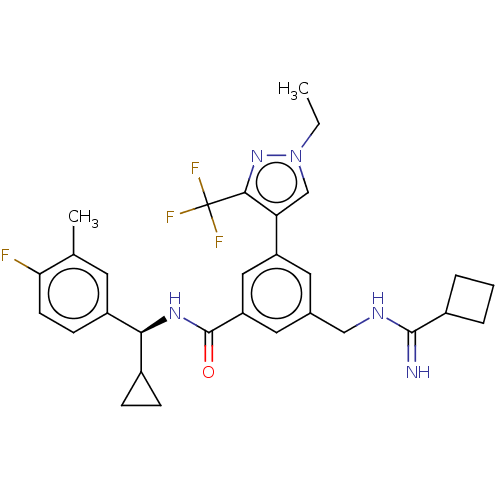
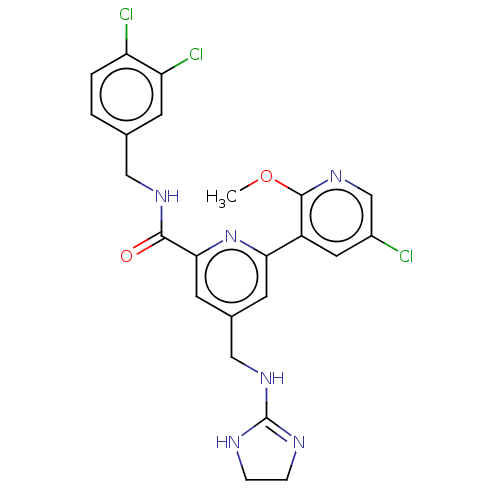
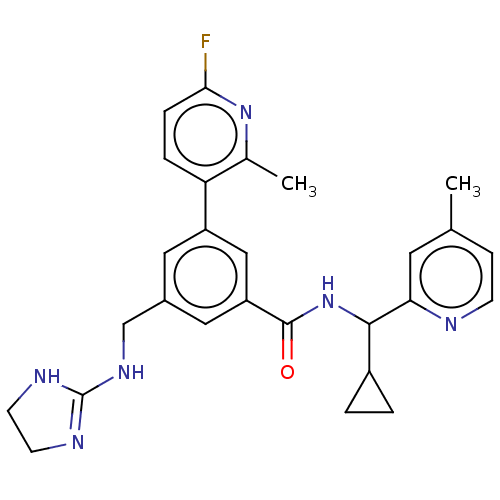
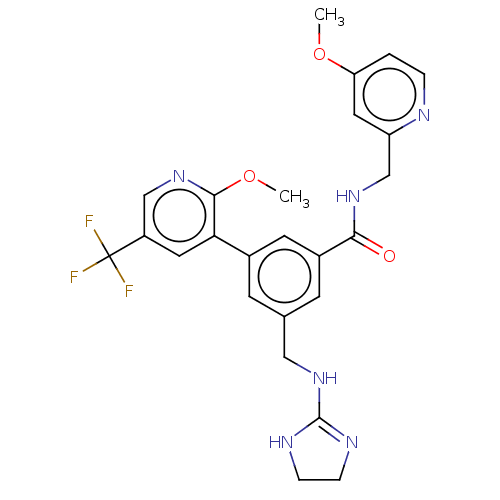
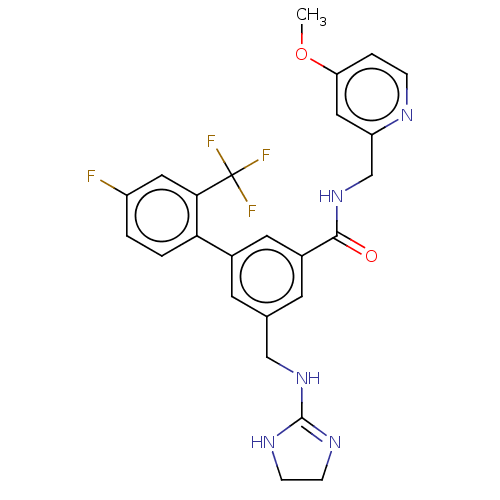
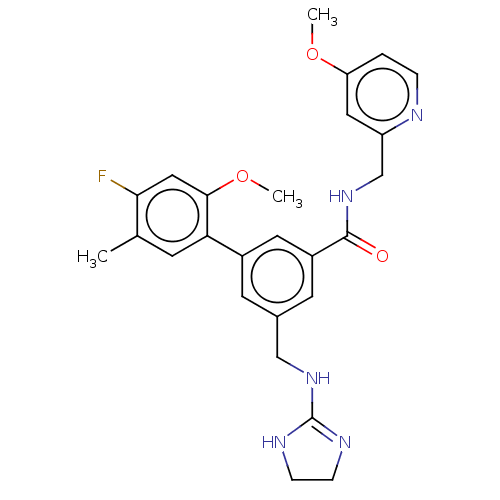
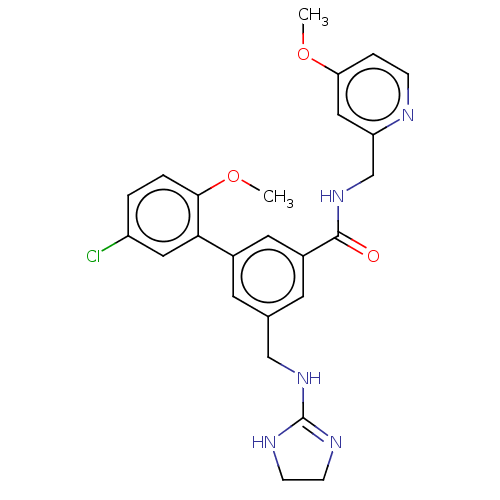
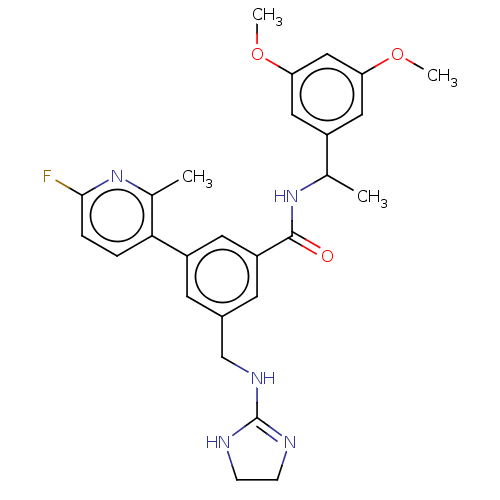
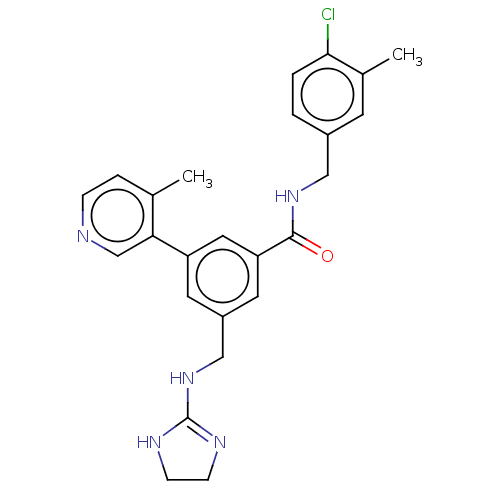
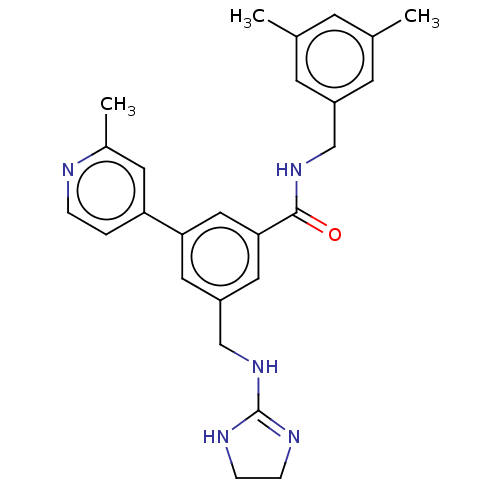
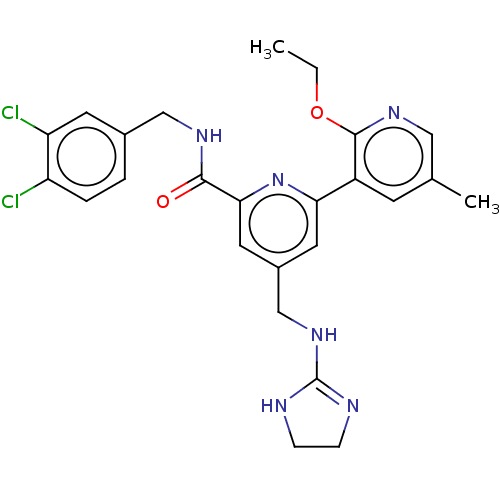
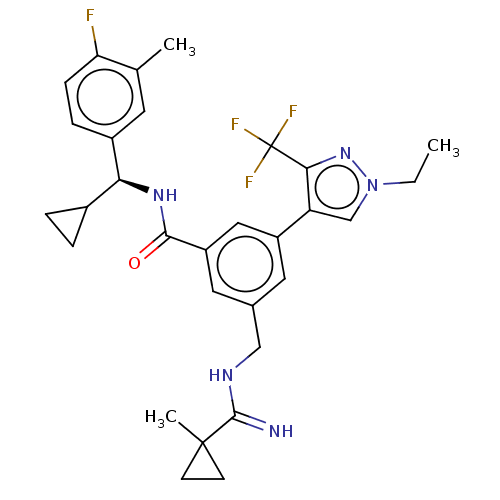
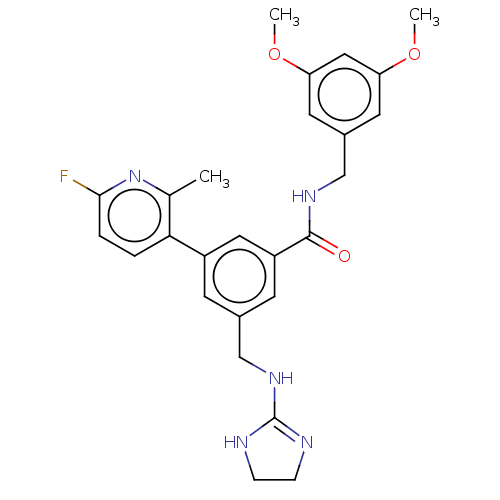
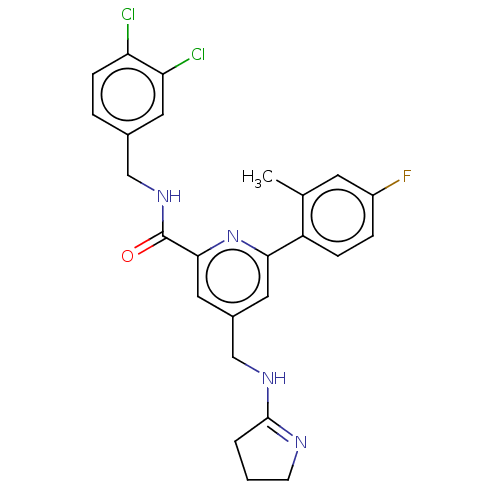
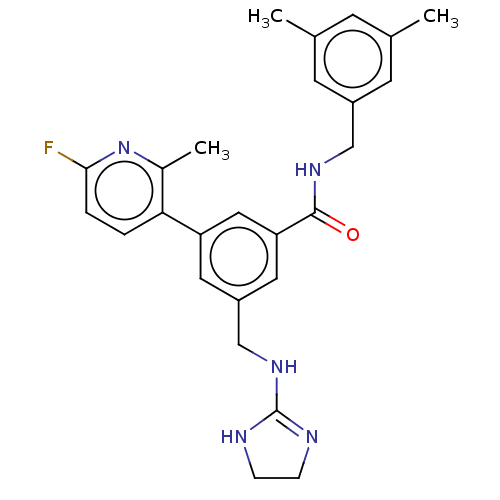
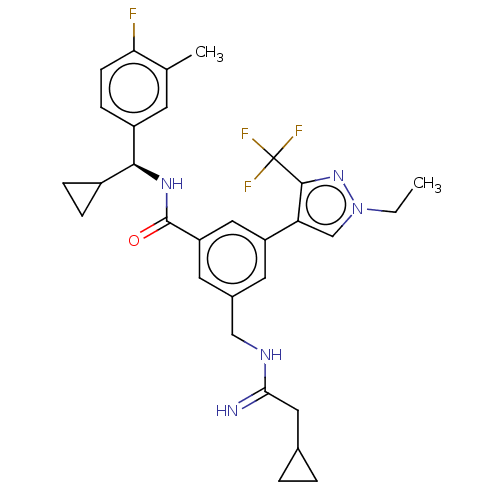
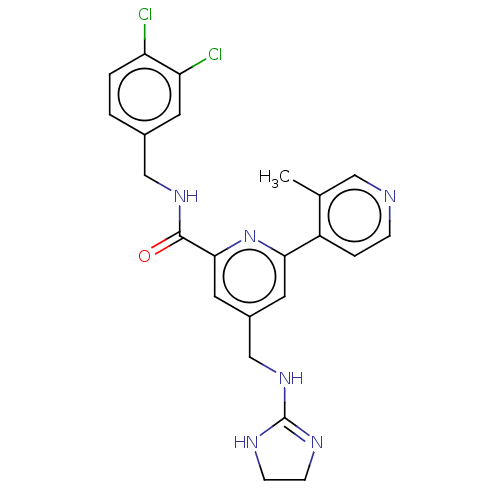
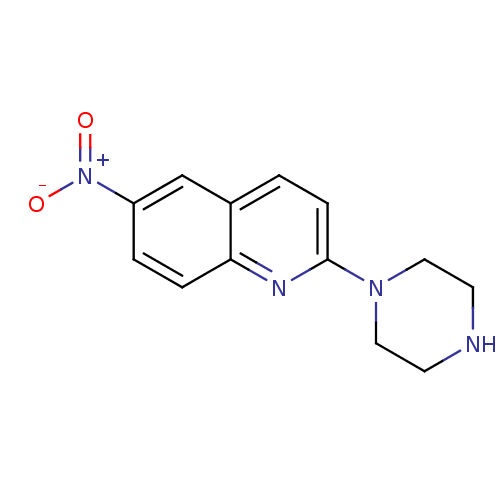
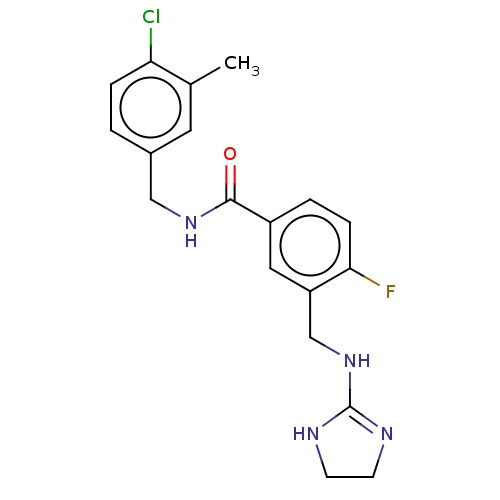
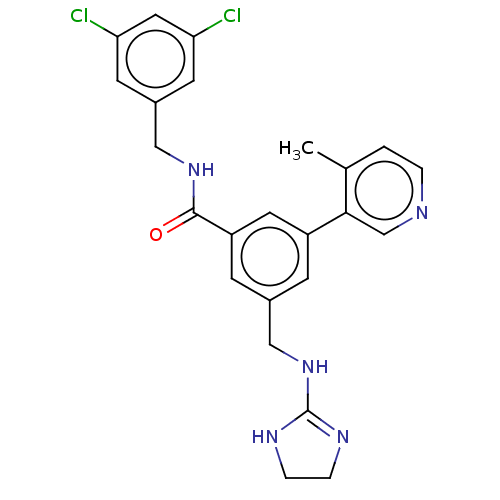
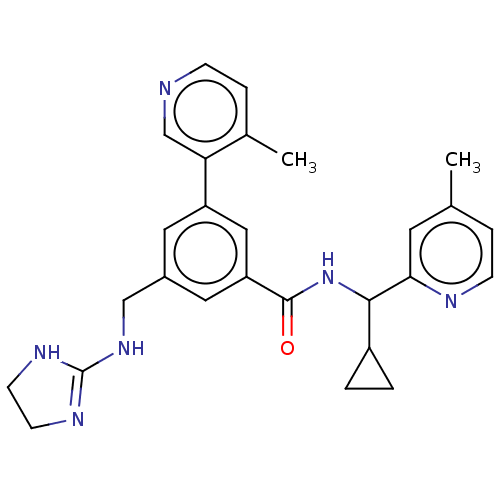
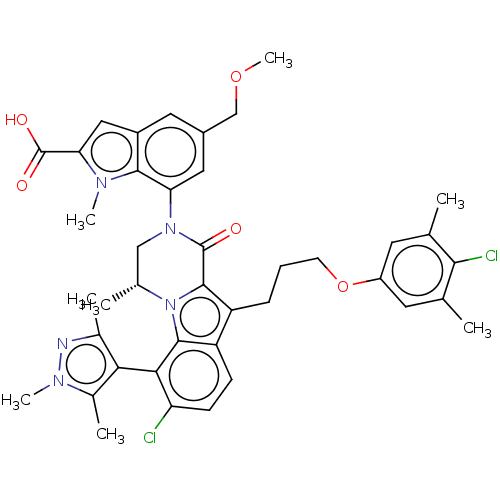
Affinity DataKi: 0.0200nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.0210nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 0.0300nMAssay Description:Inhibition of FITC-labeled Bak peptide binding to MBP-fused Mcl-1 (unknown origin) measured after 3 hrs by TR-FRET assayMore data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 0.0300nMAssay Description:Inhibition of FITC-labeled Bak peptide binding to MBP-fused Mcl-1 (unknown origin) measured after 3 hrs by TR-FRET assayMore data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 0.0300nMAssay Description:Inhibition of FITC-labeled Bak peptide binding to MBP-fused Mcl-1 (unknown origin) measured after 3 hrs by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0310nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.0330nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.0380nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 0.0810nMAssay Description:Inhibition of FITC-labeled Bak peptide binding to MBP-fused Mcl-1 (unknown origin) measured after 3 hrs by TR-FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0870nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.0870nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 0.0890nMAssay Description:Inhibition of FITC-Bak-BH3 binding to MBP-fused MCL1 (172 to 327 residues) (unknown origin) expressed in Escherichia coli BL21 CodonPlus (DE3) RIL af...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 0.0890nMAssay Description:Inhibition of FITC-Bak-BH3 binding to MBP-fused MCL1 (172 to 327 residues) (unknown origin) expressed in Escherichia coli BL21 CodonPlus (DE3) RIL af...More data for this Ligand-Target Pair
Affinity DataKi: 0.0890nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 0.0890nMAssay Description:Inhibition of FITC-Bak-BH3 binding to MBP-fused MCL1 (172 to 327 residues) (unknown origin) expressed in Escherichia coli BL21 CodonPlus (DE3) RIL af...More data for this Ligand-Target Pair
Affinity DataKi: 0.0970nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: <0.100nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.106nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.110nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.115nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.115nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.118nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.130nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.131nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.132nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rattus norvegicus (rat))
Inha University
Curated by ChEMBL
Inha University
Curated by ChEMBL
Affinity DataKi: 0.170nMAssay Description:Inhibition of [3H]citalopram binding to Serotonin transporter of rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.189nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.198nMAssay Description:FPA Assay protocol adopted from Karatas et al. (J. Med. Chem. 2010, 5179.; J. Amer. Chem. Soc. 2013, 669.): WDR5 (Δ23, residues 24-334), is expr...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:A Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) assay that measures the displacement of the 10mer-Thr-FAM probe in response to compo...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: <0.200nMAssay Description:Measurements were carried out in OptiPlate-384 White Opaque 384-well plates (Perkin Elmer. Shelton, Conn.). The assay was run using a fluorescein iso...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: <0.200nMAssay Description:Measurements were carried out in OptiPlate-384 White Opaque 384-well plates (Perkin Elmer. Shelton, Conn.). The assay was run using a fluorescein iso...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Vanderbilt University School Of Medicine
Curated by ChEMBL
Vanderbilt University School Of Medicine
Curated by ChEMBL
Affinity DataKi: <0.200nMAssay Description:Measurements were carried out in OptiPlate-384 White Opaque 384-well plates (Perkin Elmer. Shelton, Conn.). The assay was run using a fluorescein iso...More data for this Ligand-Target Pair

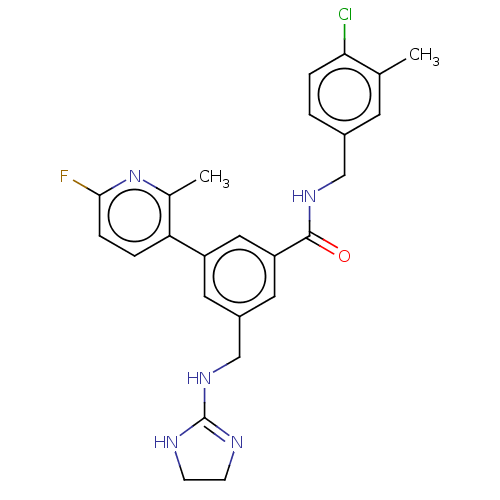
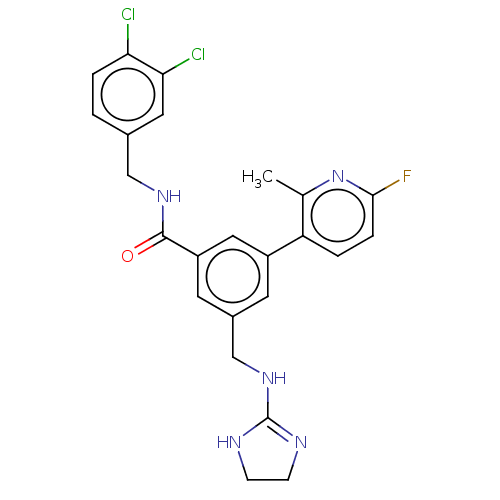
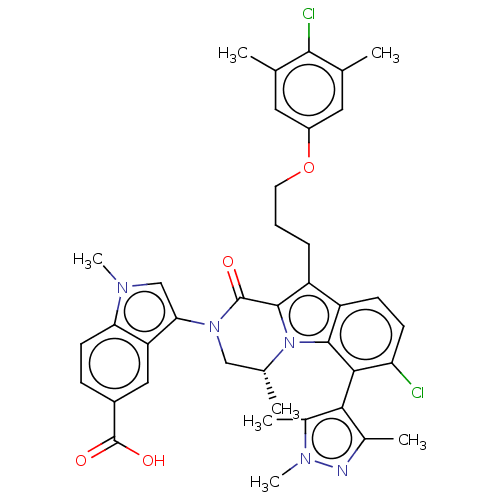
 3D Structure (crystal)
3D Structure (crystal)