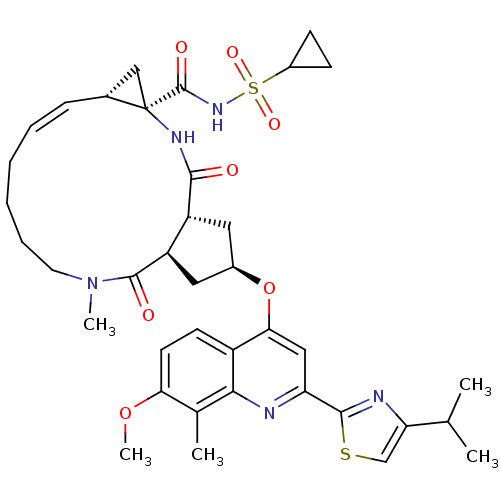
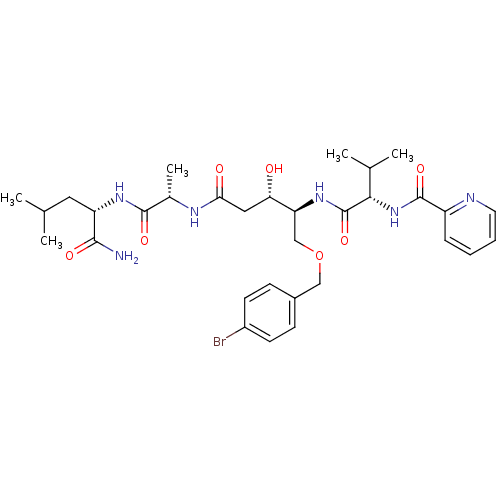
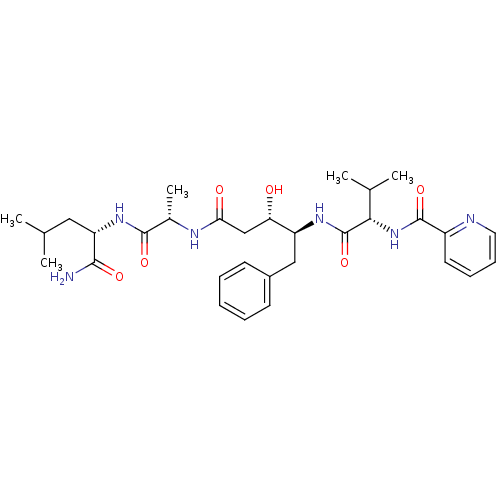
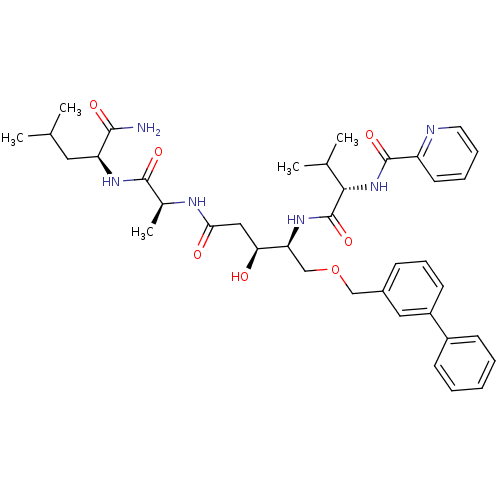
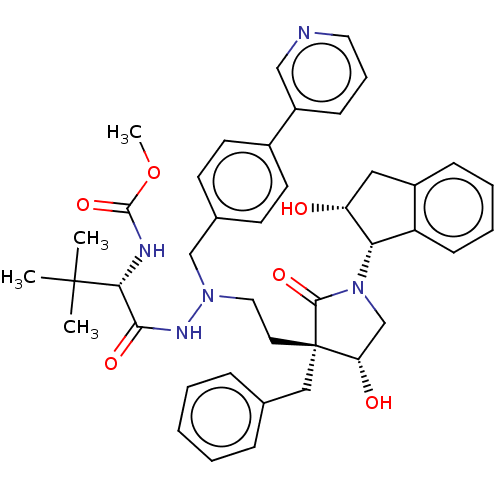
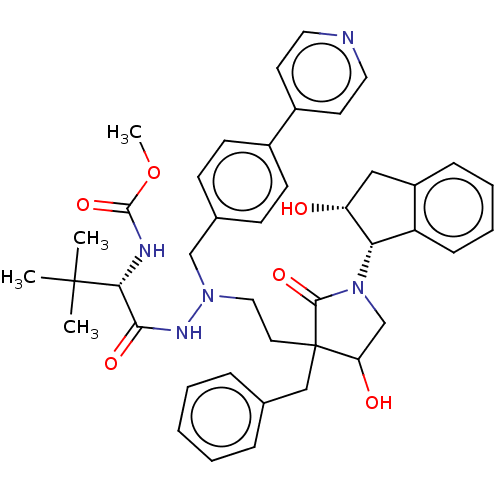
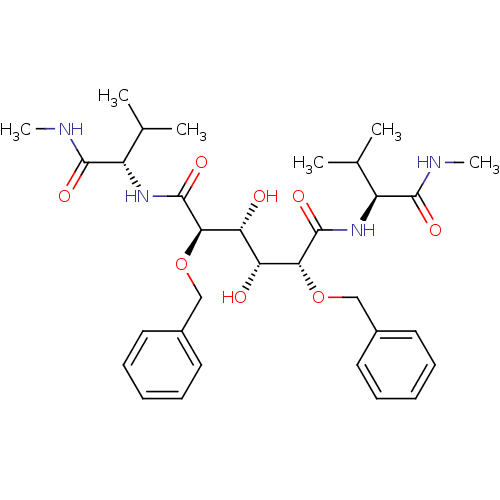
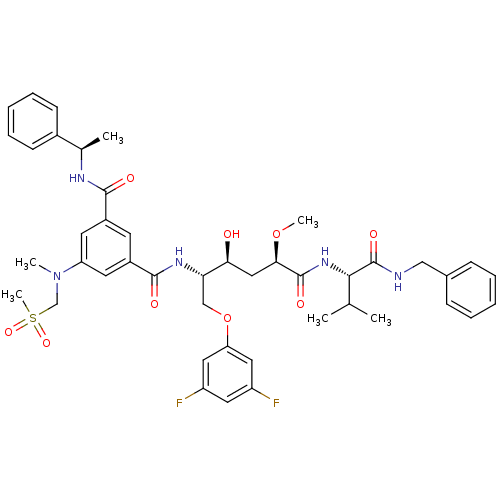
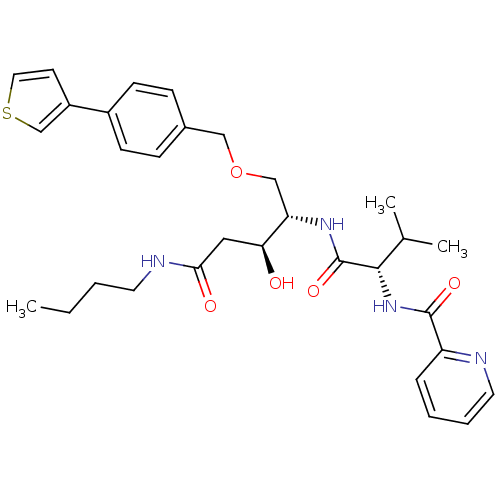
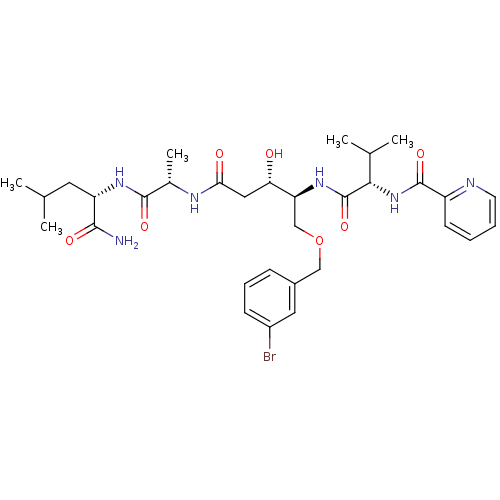
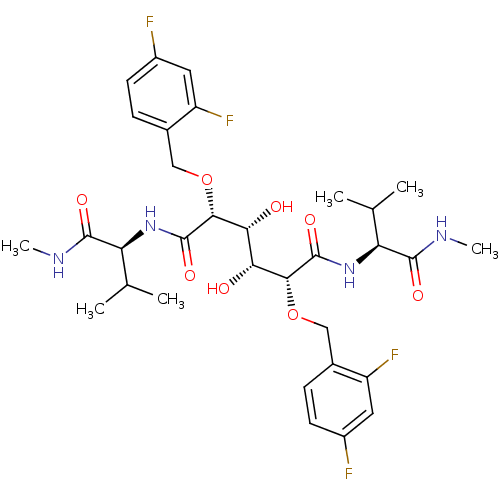
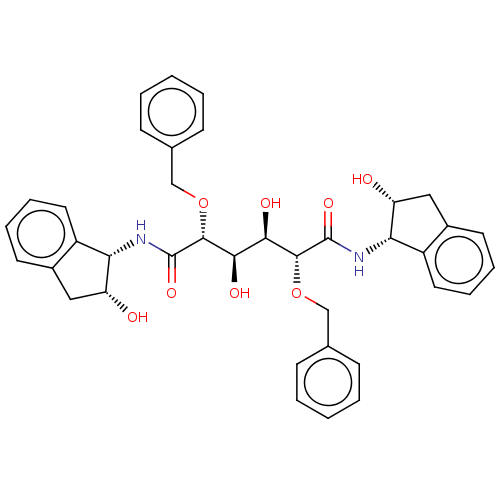
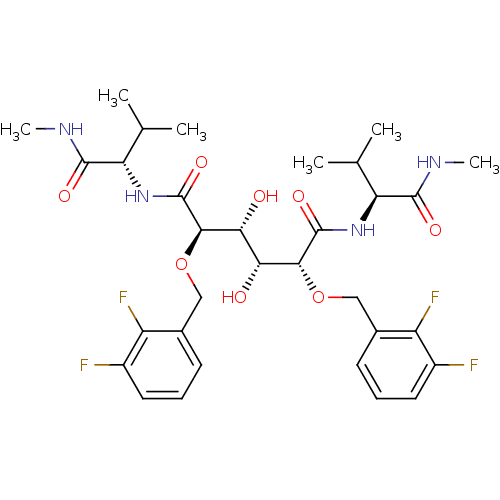
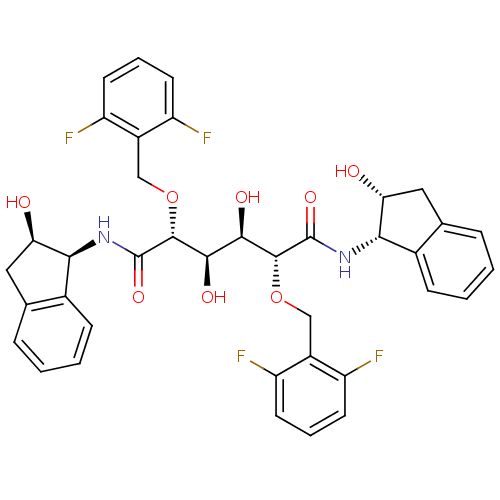
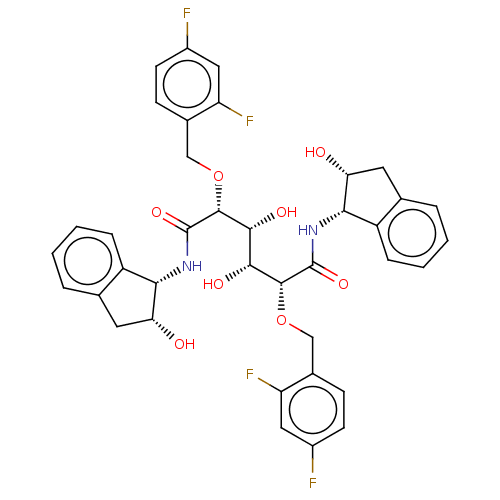
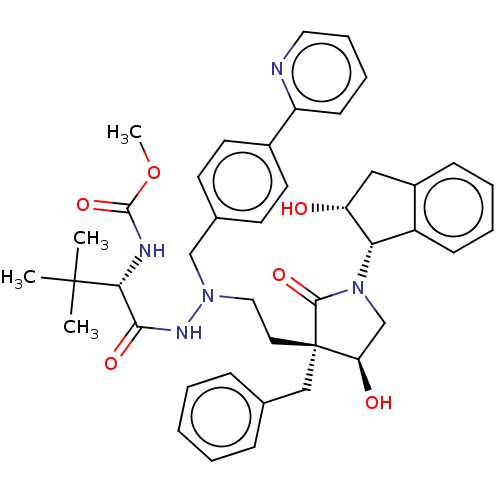
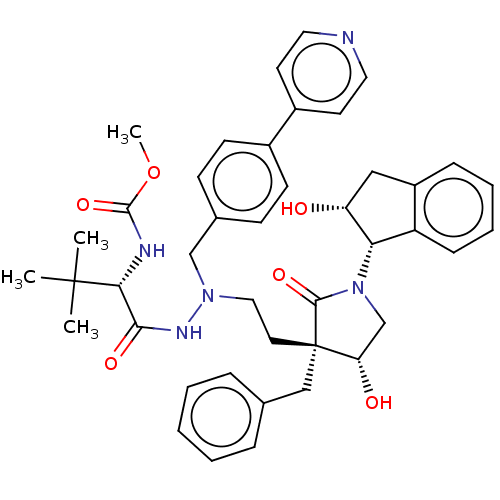
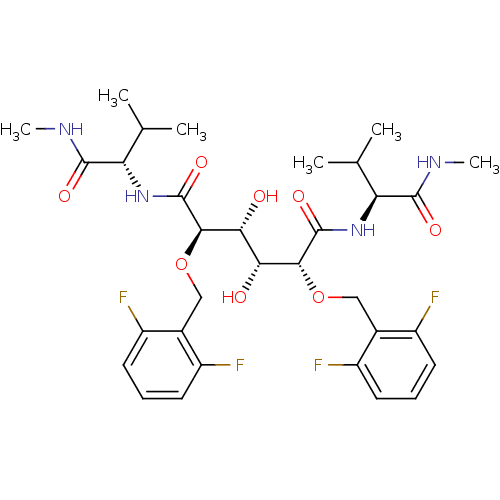
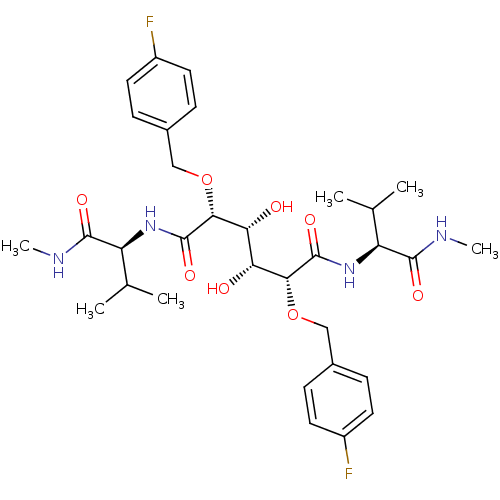
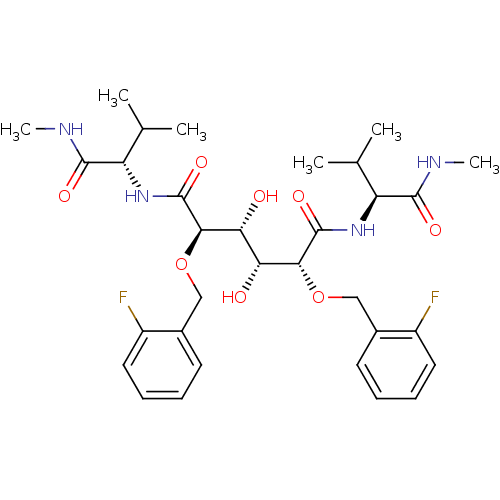
Affinity DataKi: 0.0500nM ΔG°: -59.8kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -56.3kJ/molepH: 7.5 T: 2°CAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nM ΔG°: -55.7kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -55.3kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nM ΔG°: -54.0kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.560nMAssay Description:Inhibitory concentration against the Plasmepsin II of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 0.560nM ΔG°: -52.3kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Inhibitory concentration against the human Cathepsin DMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 0.800nM ΔG°: -52.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Inhibitory concentration against the Plasmepsin II of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: <1nMAssay Description:Inhibition of human BACE1 expressed in Escherichia coli cells (BL21(DE3) by TRF assayMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Inhibitory concentration against the Plasmepsin II of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Inhibitory concentration against the Plasmepsin II of Plasmodium falciparumMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.12nM ΔG°: -51.9kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.22nM ΔG°: -51.7kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.28nM ΔG°: -51.6kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Inhibitory concentration against the human Cathepsin DMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.64nM ΔG°: -51.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.65nM ΔG°: -51.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using DABCYL-Abu-Ser-Gln-ASN-Tyr-Pro-Ile-Val-Gln-EDANS as substrate preincubated for 20 min...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.73nM ΔG°: -50.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.79nM ΔG°: -50.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.83nM ΔG°: -50.7kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.90nMAssay Description:Inhibitory concentration against the Plasmepsin I of Plasmodium falciparumMore data for this Ligand-Target Pair
Affinity DataKi: 1.90nMAssay Description:Inhibitory concentration against the Plasmepsin II of Plasmodium falciparumMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.92nM ΔG°: -50.6kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair


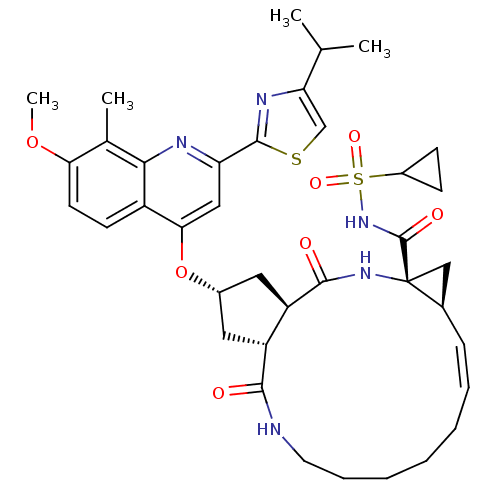

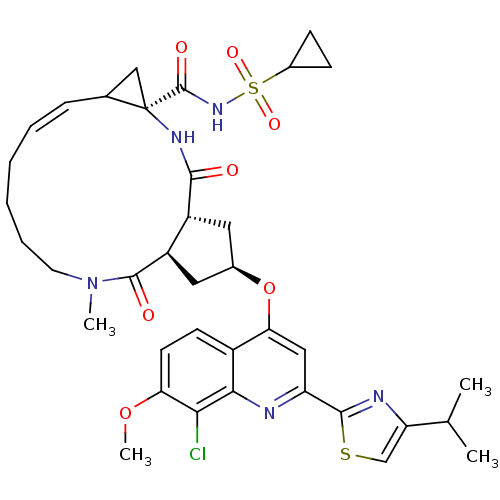
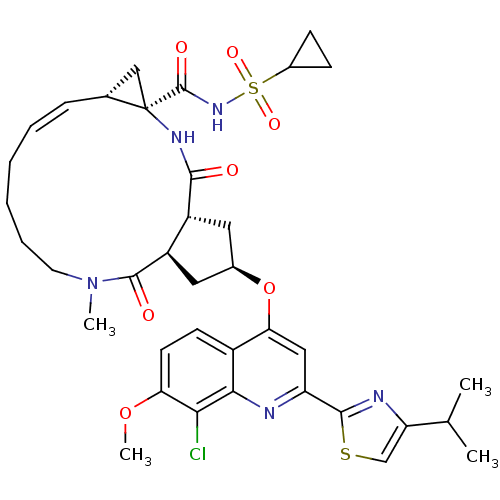

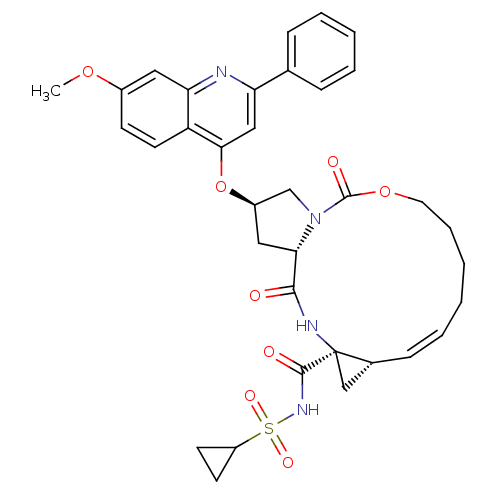
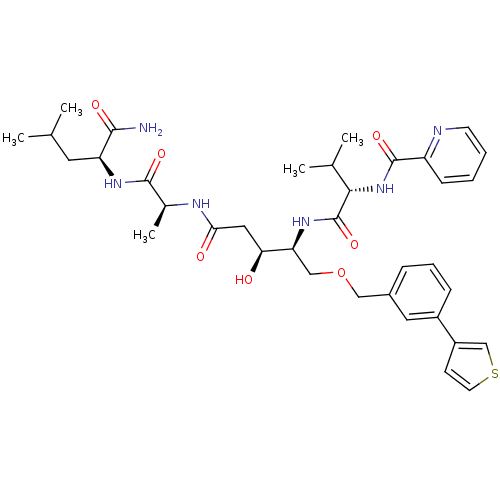
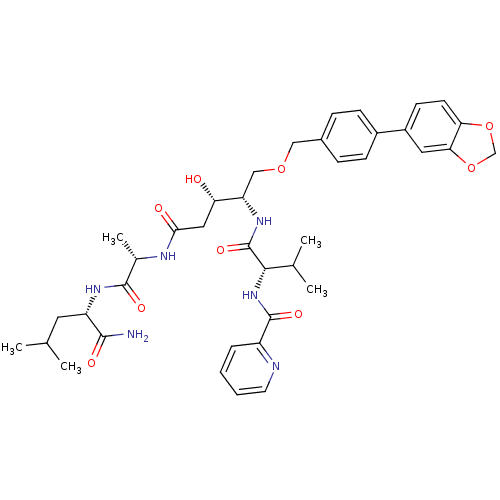
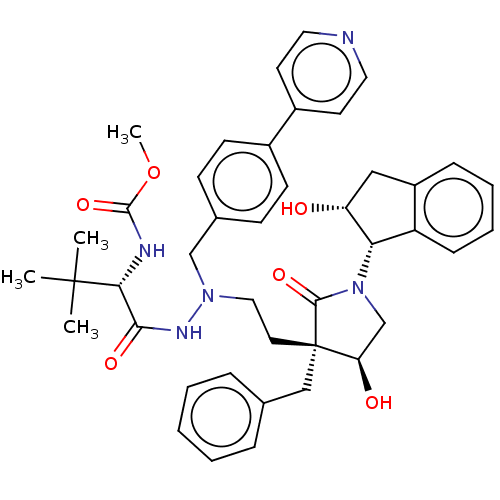
 3D Structure (crystal)
3D Structure (crystal)