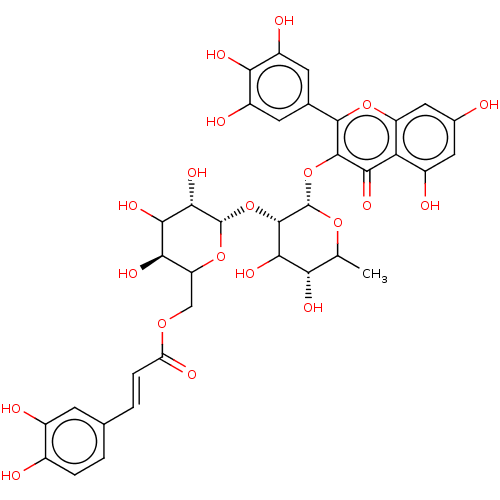
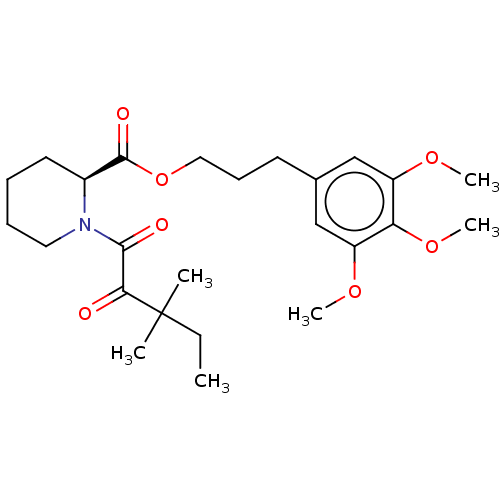
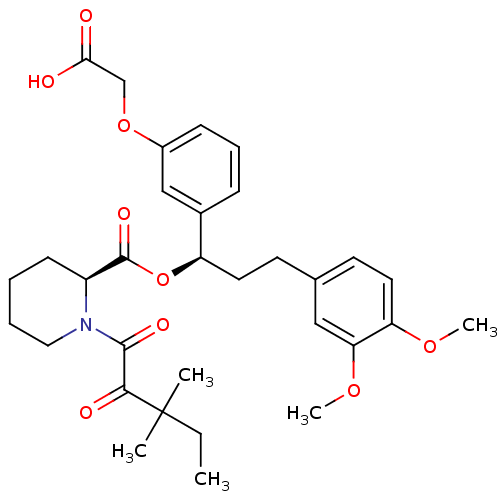
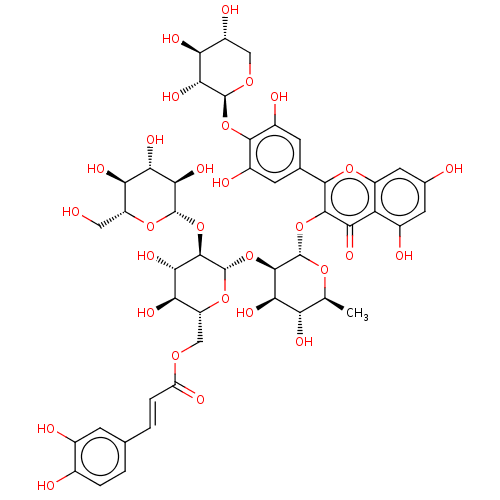
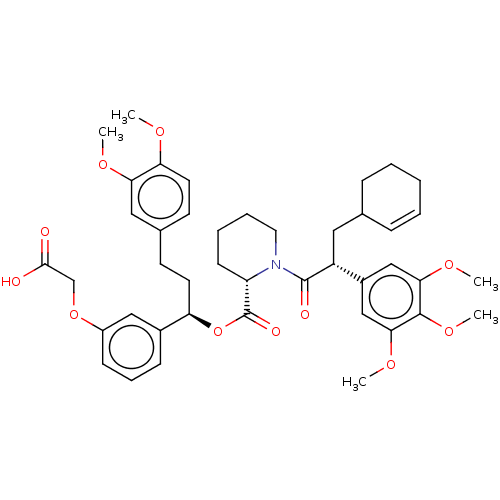
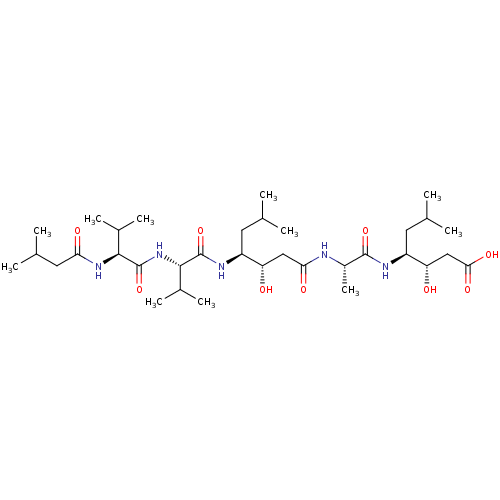
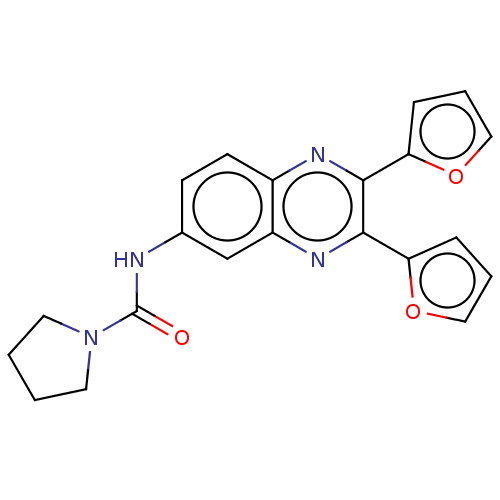
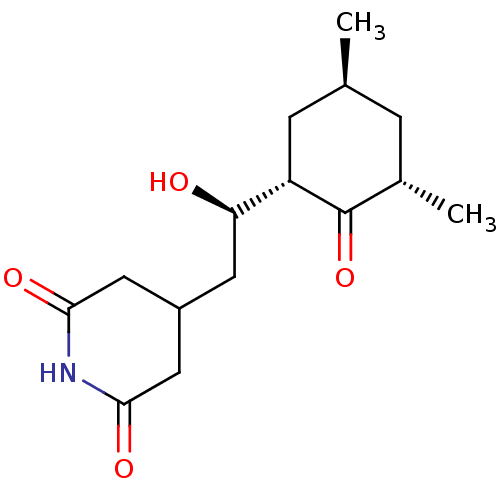
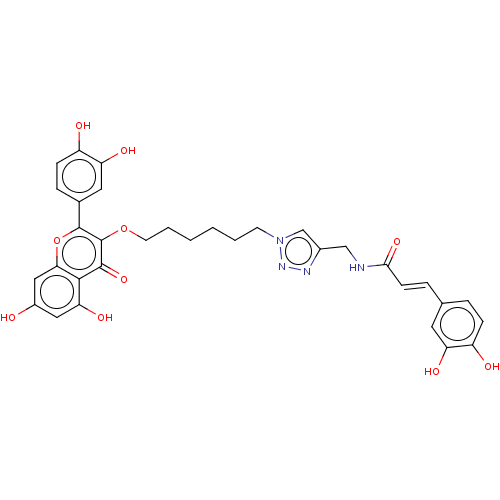
Affinity DataKi: 4nMAssay Description:Binding affinity to wild type FKBP51 (unknown origin) by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Ligand Info
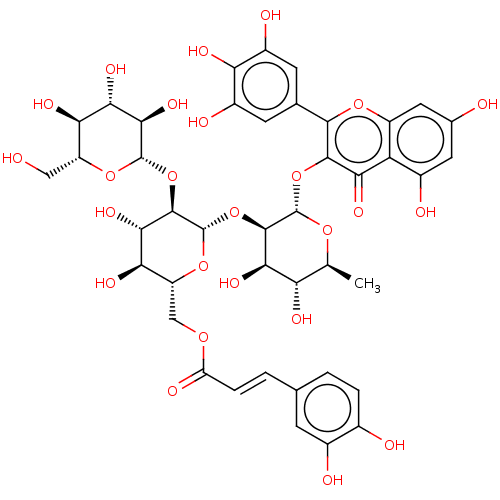
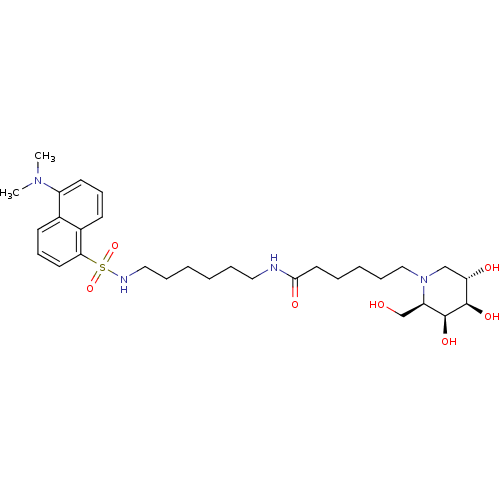
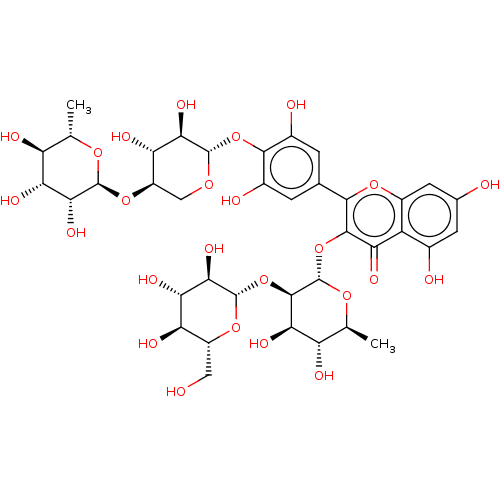
Affinity DataKi: 6nMAssay Description:Binding affinity to wild type FKBP51 (unknown origin) by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Ligand Info
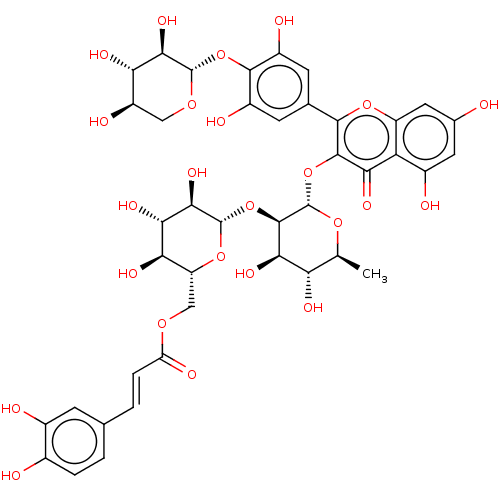
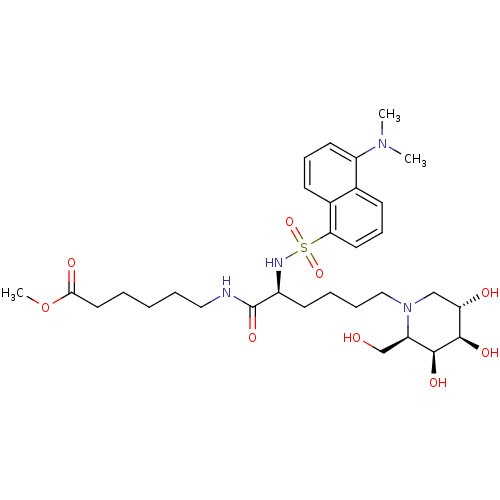
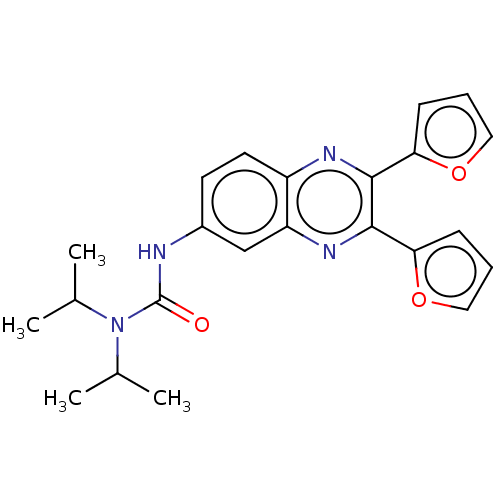
Affinity DataKi: 7nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Target InfoPDBMMDBNCI pathwayReactome pathwayKEGG
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
Ligand Info
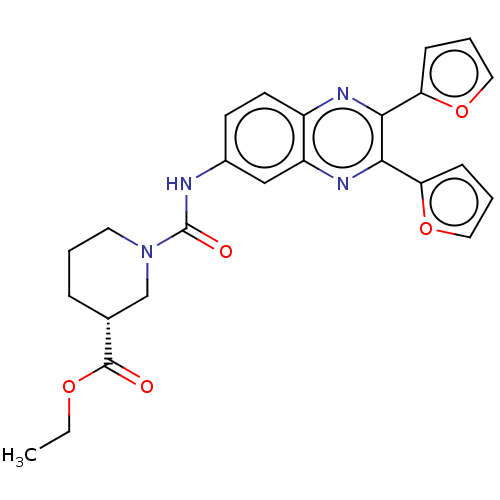
Affinity DataKi: 7.5nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 8.10nM ΔG°: -47.0kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 9.10nM ΔG°: -46.7kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Target InfoPDBMMDBNCI pathwayReactome pathwayKEGG
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
Ligand Info
Affinity DataKi: 20nMAssay Description:Binding affinity to FKBP52 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 21.3nM ΔG°: -44.5kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Binding affinity to wild type FKBP51 (unknown origin) by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 26nMAssay Description:Binding affinity to wild type FKBP51 (unknown origin) by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 42.4nM ΔG°: -42.8kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 79.3nM ΔG°: -41.2kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 93nMAssay Description:Binding affinity to FKBP51 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 93.3nM ΔG°: -40.8kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 94nMAssay Description:Inhibition of wild-type HIV-1 protease using fluorogenic peptide substrate incubated for 30 mins prior to substrate addition measured after 10 mins b...More data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 220nMAssay Description:Inhibition of Escherichia coli beta-galactosidase assessed as inhibition of nitrophenolate release by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 230nMAssay Description:Inhibition of multidrug resistant HIV-1 protease L10I/L63P/A71V/G73S/I84V/L90M mutant using fluorogenic peptide substrate incubated for 30 mins prior...More data for this Ligand-Target Pair
Affinity DataKi: 270nMAssay Description:Inhibition of Escherichia coli beta-galactosidase assessed as inhibition of nitrophenolate release by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 300nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Target InfoPDBMMDBNCI pathwayReactome pathwayKEGG
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
Ligand InfoPDB
Affinity DataKi: 300nMAssay Description:Inhibition of human lysosomal beta-galactosidase assessed as inhibition of hydrolyzed 4-methylumbelliferone production after 30 mins by luminescence ...More data for this Ligand-Target Pair
Affinity DataKi: 306nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 447nMAssay Description:Binding affinity to FKBP51 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 600nMAssay Description:Inhibition of human lysosomal beta-galactosidase assessed as inhibition of hydrolyzed 4-methylumbelliferone production after 30 mins by luminescence ...More data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:Inhibition of human lysosomal beta-galactosidase assessed as inhibition of hydrolyzed 4-methylumbelliferone production after 30 mins by luminescence ...More data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:Inhibition of human lysosomal beta-galactosidase assessed as inhibition of hydrolyzed 4-methylumbelliferone production after 30 mins by luminescence ...More data for this Ligand-Target Pair
Affinity DataKi: 730nM ΔG°: -35.6kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 936nMAssay Description:Binding affinity to FKBP52 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 958nMAssay Description:Binding affinity to FKBP51 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Binding affinity to CypA (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nMAssay Description:Inhibition of green coffee bean alpha-galactosidase assessed as inhibition of nitrophenolate release by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 1.69E+3nMAssay Description:Binding affinity to FKBP52 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:Binding affinity to CypD (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+3nMAssay Description:Binding affinity to CypB (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.24E+3nM ΔG°: -32.8kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nMAssay Description:Binding affinity to CypA (unknown origin)More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 3.00E+3nMAssay Description:Binding affinity to CypD (unknown origin)More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 3.00E+3nMAssay Description:Binding affinity to CypA (unknown origin)More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 3.00E+3nMAssay Description:Binding affinity to CypD (unknown origin)More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 3.40E+3nMAssay Description:Inhibition of green coffee bean alpha-galactosidase assessed as inhibition of nitrophenolate release by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 3.40E+3nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity to FKBP52 (unknown origin)More data for this Ligand-Target Pair
Target InfoPDBMMDBNCI pathwayReactome pathwayKEGG
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
Ligand Info
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity to FKBP12 (unknown origin)More data for this Ligand-Target Pair
Target InfoPDBMMDBNCI pathwayReactome pathwayKEGG
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
Ligand Info
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity to FKBP51 (unknown origin)More data for this Ligand-Target Pair
Ligand Info
Affinity DataKi: 1.39E+4nM ΔG°: -28.2kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 �C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nMAssay Description:Binding affinity to CypA (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.30E+4nMAssay Description:Binding affinity to wild type FKBP51 (unknown origin) by competitive fluorescence polarization assayMore data for this Ligand-Target Pair
Ligand Info


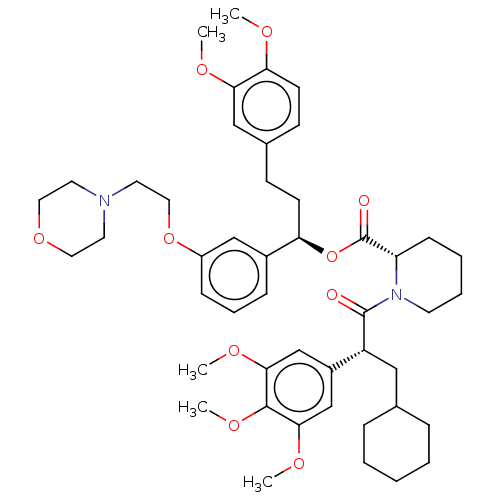
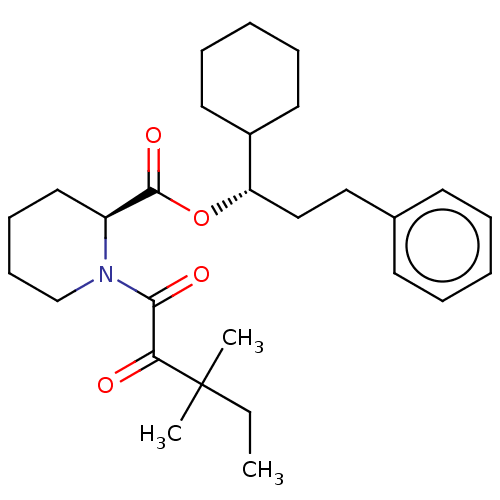

 3D Structure (crystal)
3D Structure (crystal)