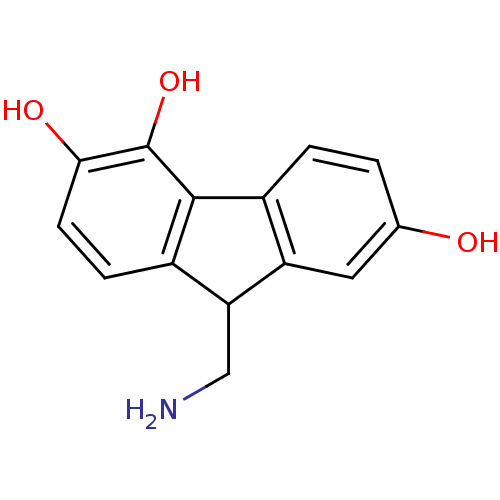
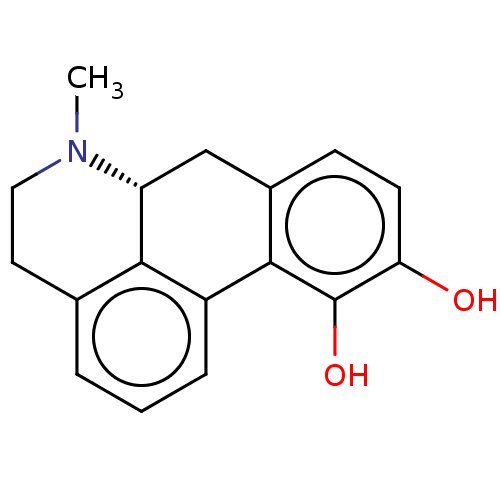
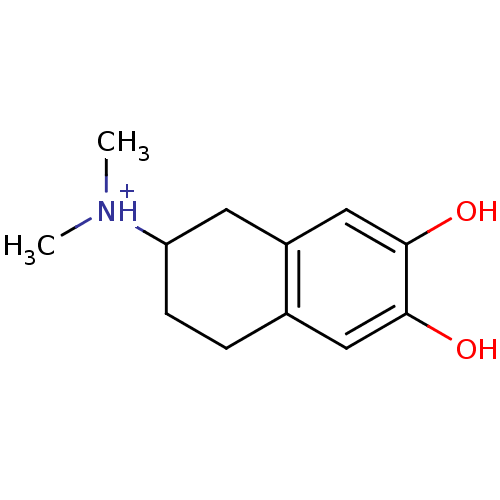
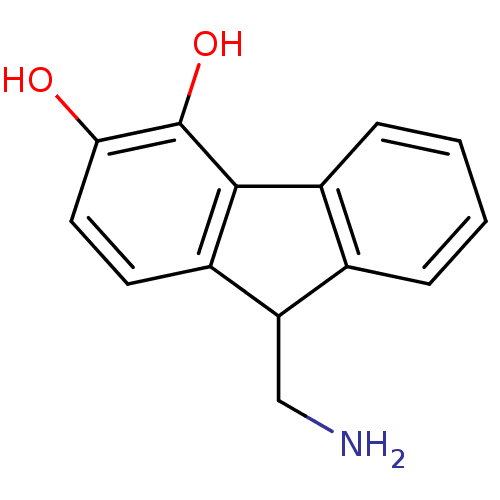
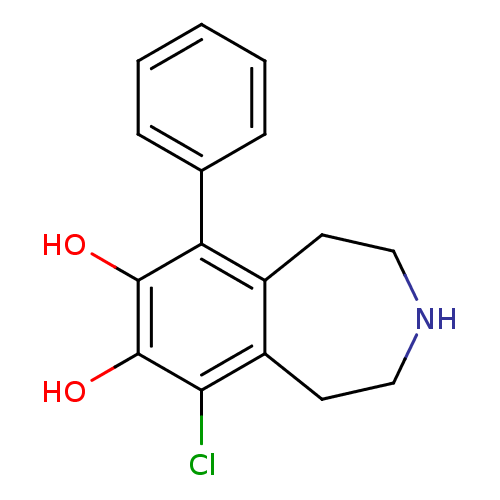
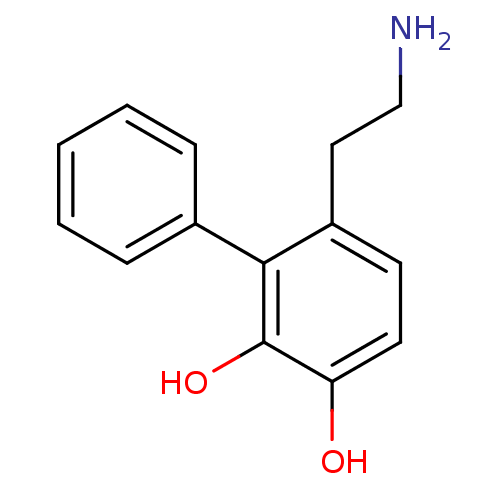
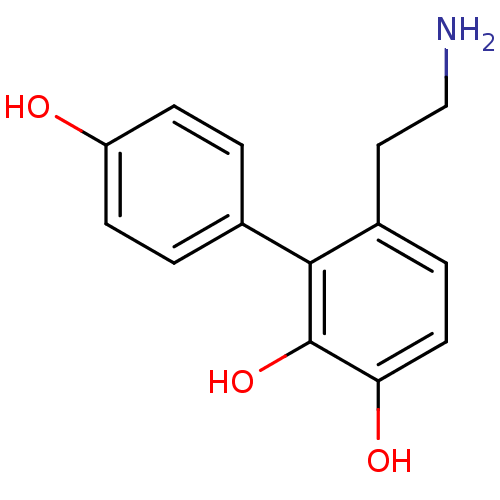
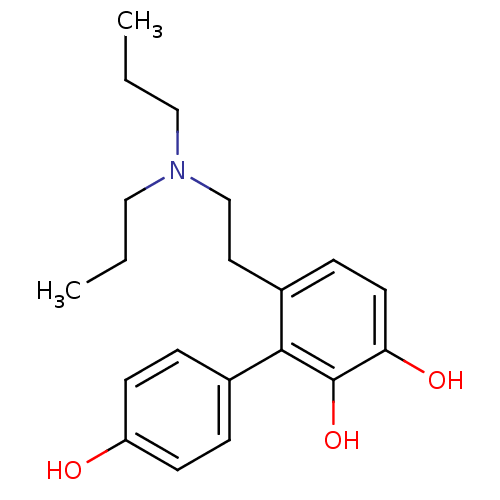
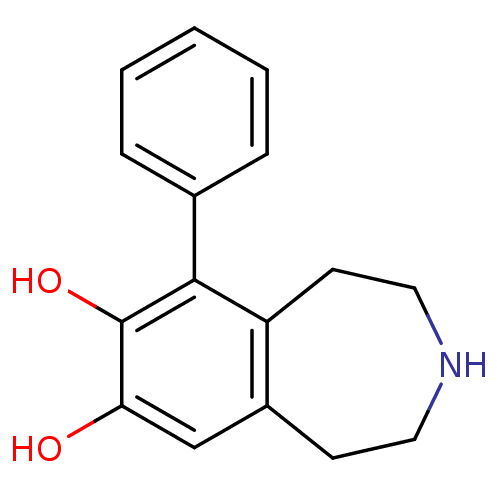
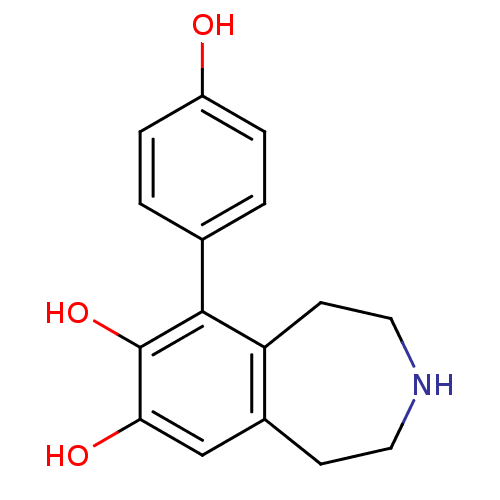
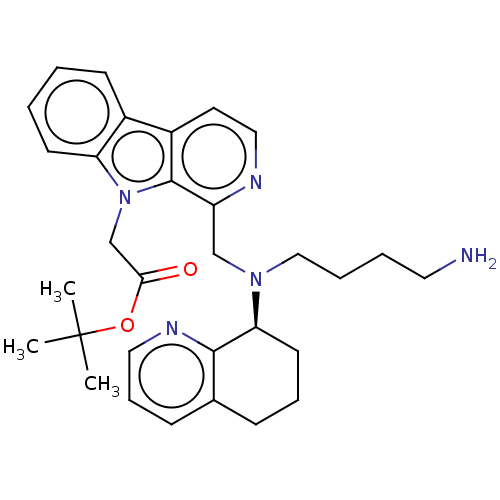
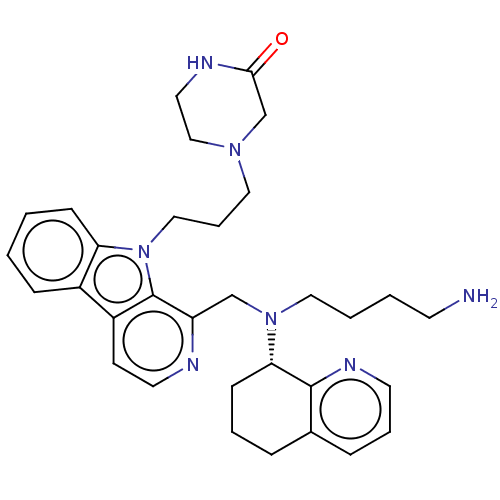
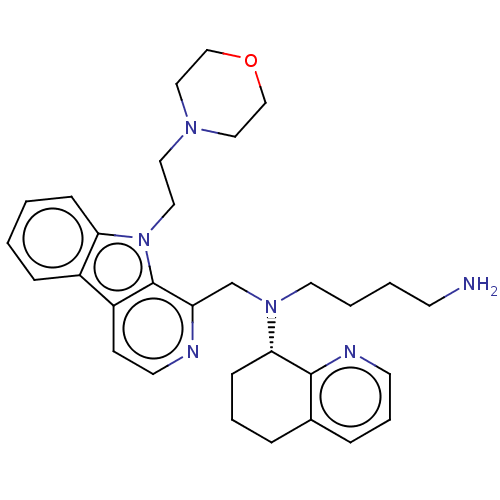
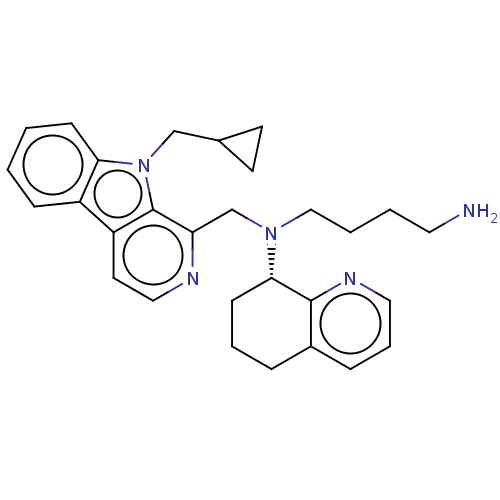
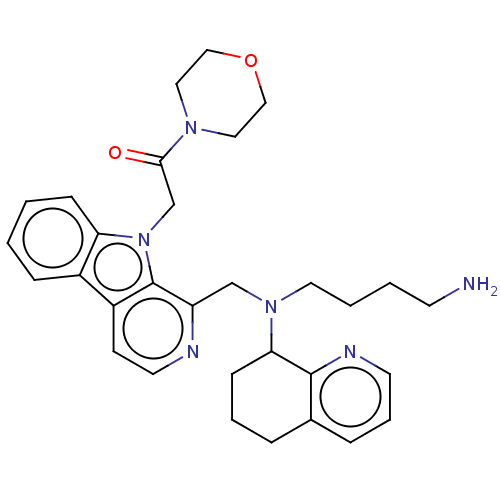
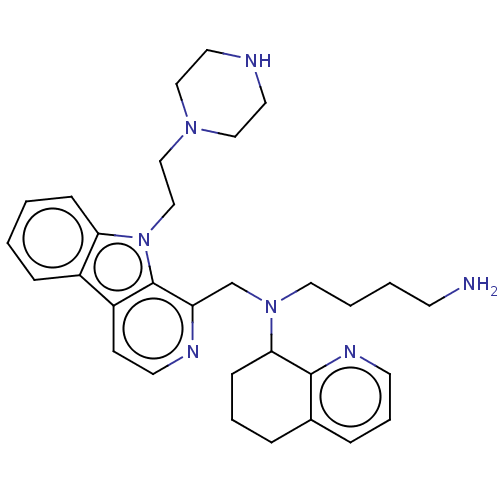
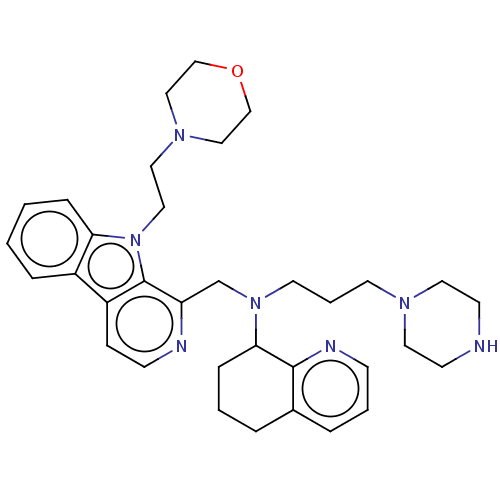
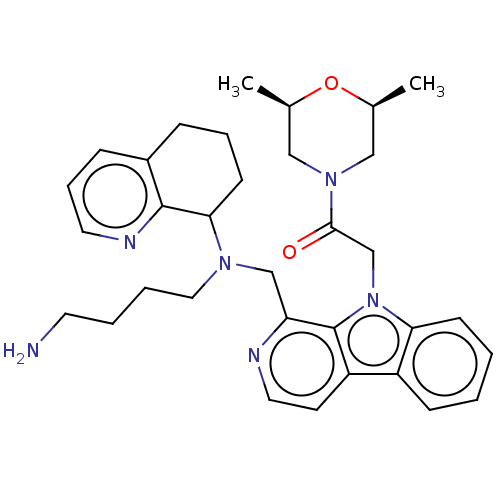
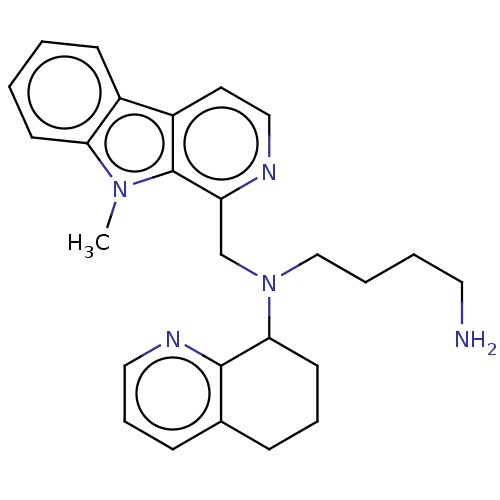
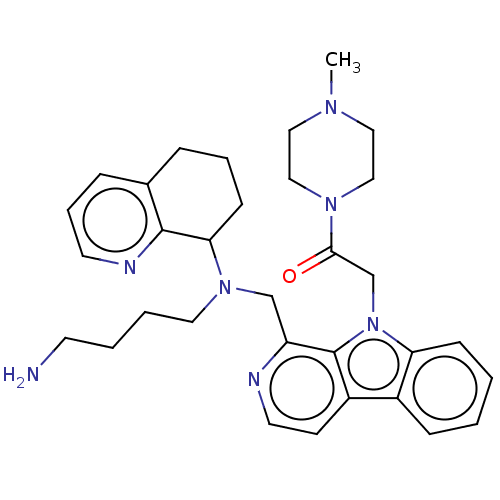
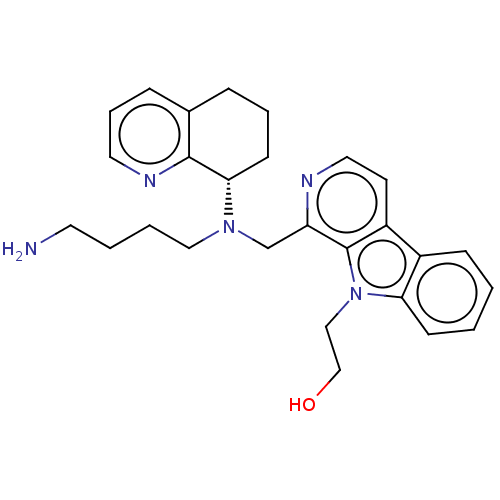
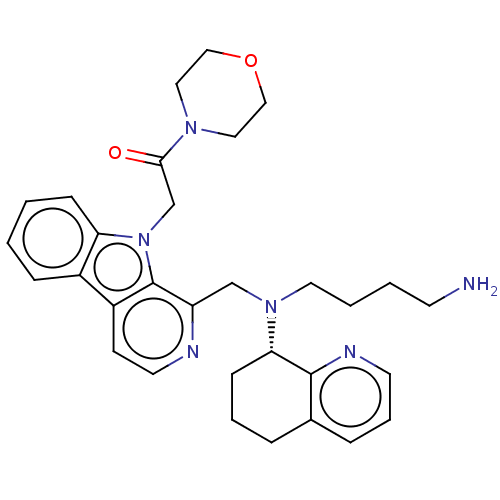
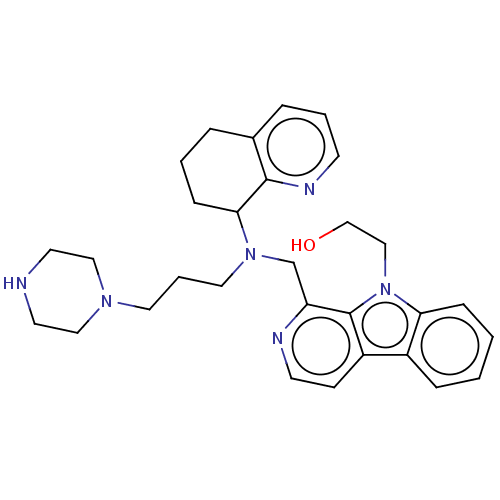
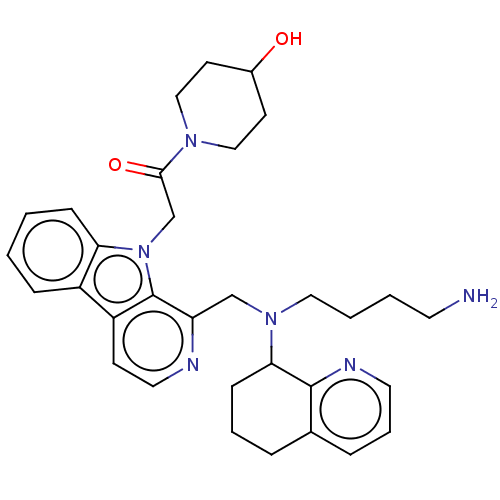
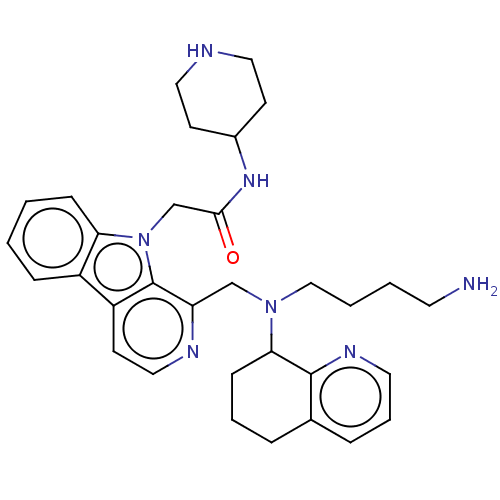
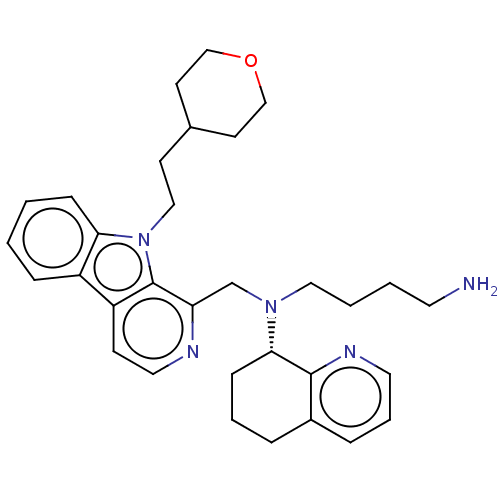
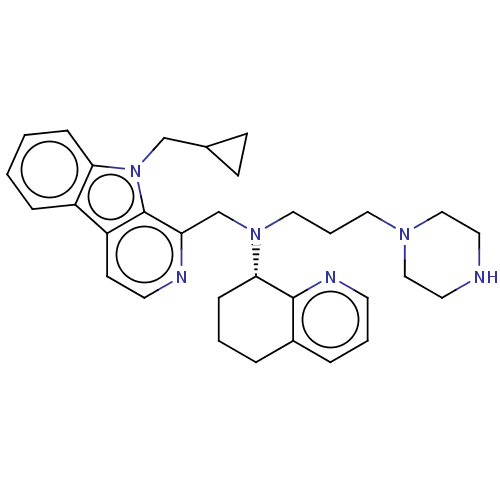
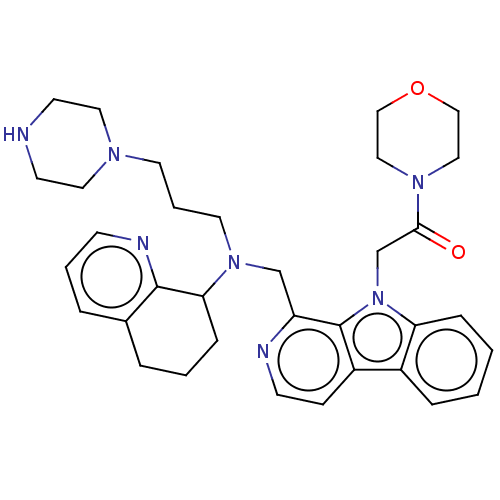
Affinity DataKi: 3nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 43nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 53nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 57nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 150nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Compound was tested for inhibition of [3H]spiroperidol binding against Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataKi: 255nMAssay Description:Compound was tested for inhibition of [3H]spiroperidol binding against Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataKi: 380nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 790nMAssay Description:Compound was tested for inhibition of [3H]spiroperidol binding against Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataKi: 970nMAssay Description:Compound was tested for binding affinity against dDopamine receptor D1 using [3H]fenoldopam as a radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.63E+3nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.65E+3nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.74E+3nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.79E+3nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nMAssay Description:Compound was tested for inhibition of [3H]spiroperidol binding against Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataKi: 1.94E+3nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 2.35E+3nMAssay Description:Compound was tested for inhibition of [3H]spiroperidol binding against Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+3nMAssay Description:Displacement of [3H]fenoldopam from Dopamine receptor D1 of rat striatum membranesMore data for this Ligand-Target Pair
Affinity DataKi: 5.74E+3nMAssay Description:Compound was tested for inhibition of [3H]spiroperidol binding against Dopamine receptor D2More data for this Ligand-Target Pair
Affinity DataIC50: 1nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 5.5nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 7.30nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 7.70nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 8.10nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 9.30nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 9.90nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair

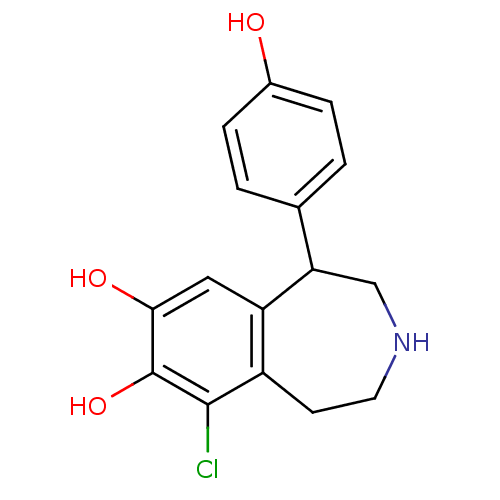
 3D Structure (crystal)
3D Structure (crystal)