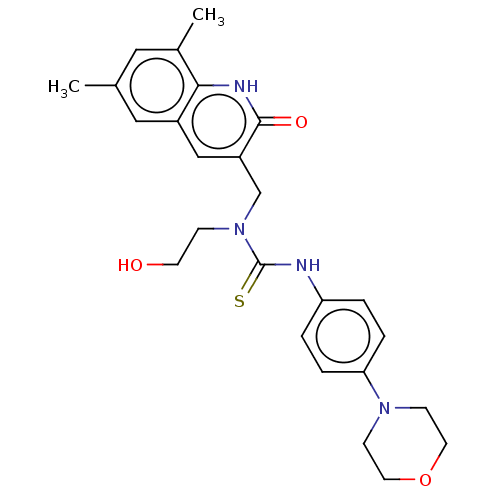
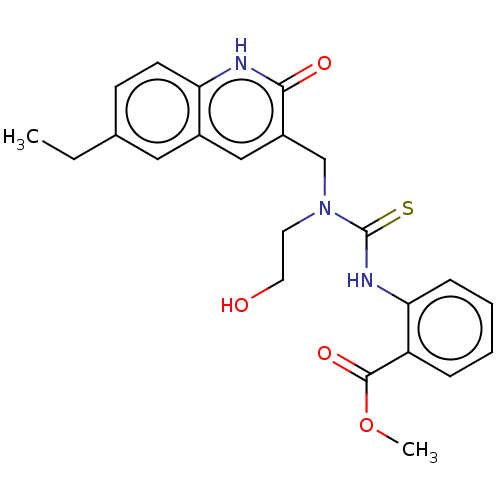
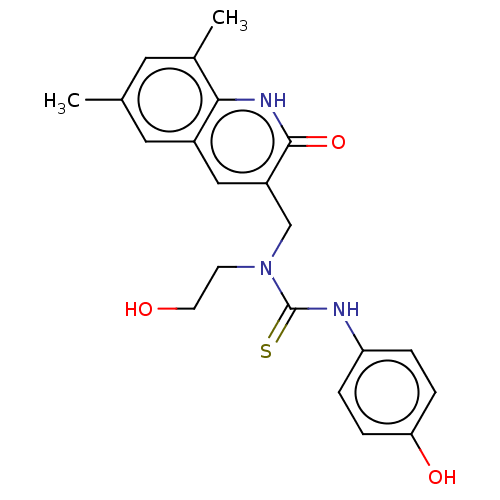
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 17 hits for monomerid = 163636,163637,163638,163639,163644,163645,163636,163637,163638,163639,163644,163645
Found 17 hits for monomerid = 163636,163637,163638,163639,163644,163645,163636,163637,163638,163639,163644,163645
Affinity DataEC50: 18nMAssay Description:Inhibition of Escherichia coli beta-glucuronidase using p-nitrophenyl-beta-D-glucuronide as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataIC50: 164nMAssay Description:In-vivo inhibition of beta-glucuronidase in BALB/CJ mouse at 10 ug administered as twice dailyMore data for this Ligand-Target Pair
Affinity DataKi: 210nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:Inhibition of beta-glucuronidase (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 610nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataKi: 680nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens)
University of North Carolina at Chapel Hill
University of North Carolina at Chapel Hill
Affinity DataKi: 970nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens)
University of North Carolina at Chapel Hill
University of North Carolina at Chapel Hill
Affinity DataKi: 1.10E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina at Chapel Hill
University of North Carolina at Chapel Hill
Affinity DataKi: 1.40E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
Affinity DataKi: 1.90E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina at Chapel Hill
University of North Carolina at Chapel Hill
Affinity DataKi: 3.00E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens)
University of North Carolina at Chapel Hill
University of North Carolina at Chapel Hill
Affinity DataKi: 7.80E+3nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina at Chapel Hill
University of North Carolina at Chapel Hill
Affinity DataKi: 1.10E+4nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase [V151I,I185V](Clostridium perfringens)
University of North Carolina at Chapel Hill
University of North Carolina at Chapel Hill
Affinity DataKi: 2.40E+4nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
TargetBeta-galactosidase(Streptococcus agalactiae (Firmicutes))
University of North Carolina at Chapel Hill
University of North Carolina at Chapel Hill
Affinity DataKi: 3.60E+4nMAssay Description:Reactions were conducted similarly to the kinetic assays but the reaction consisted of 10 µL assay buffer, 5 µL inhibitor solution, 5 µL of 10 nM enz...More data for this Ligand-Target Pair
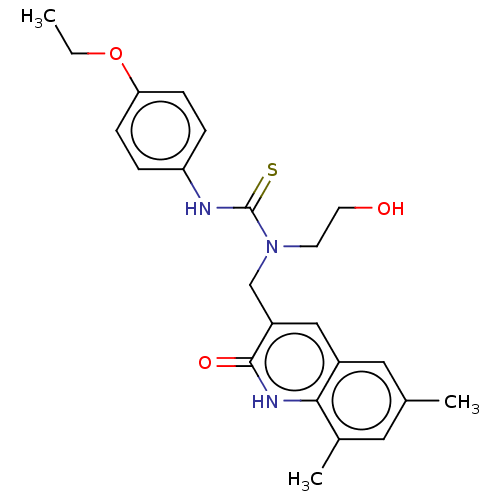
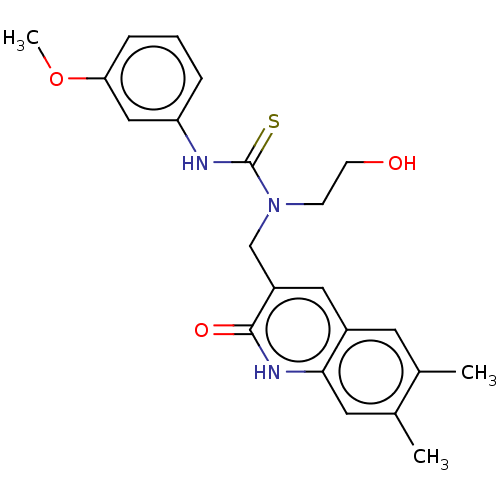
 3D Structure (crystal)
3D Structure (crystal)