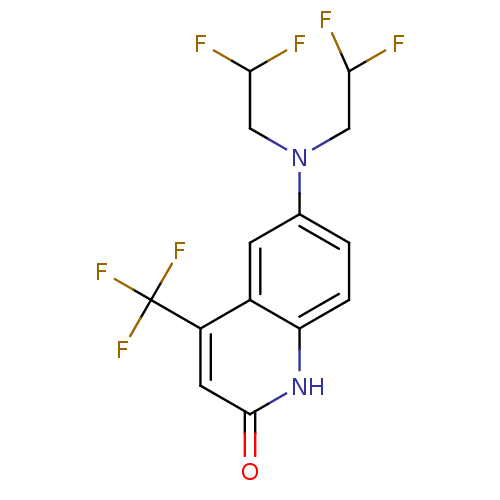
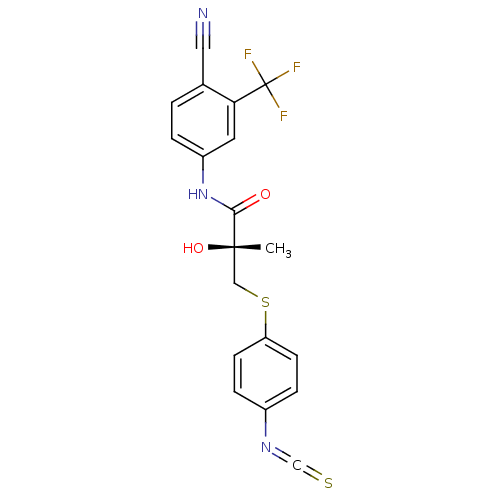
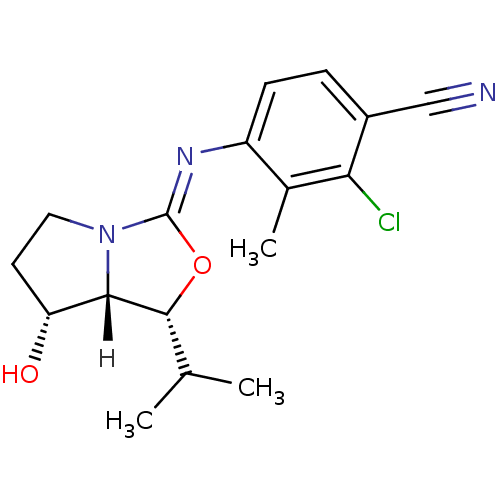
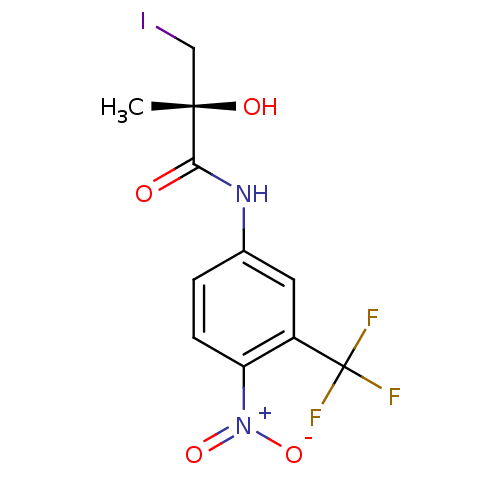
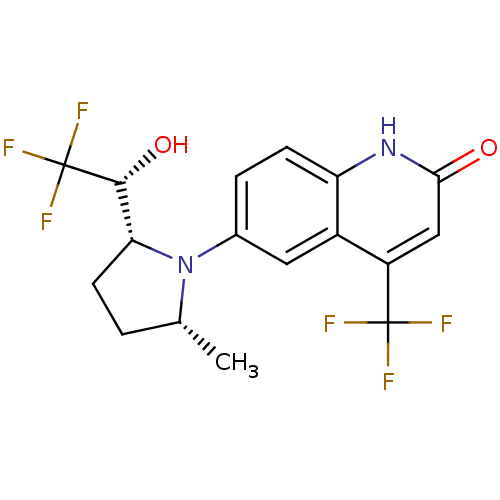
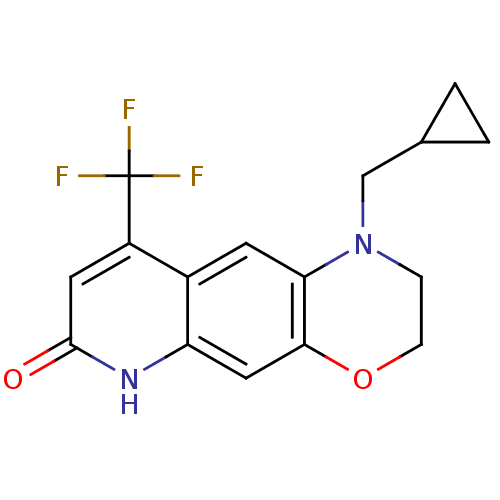
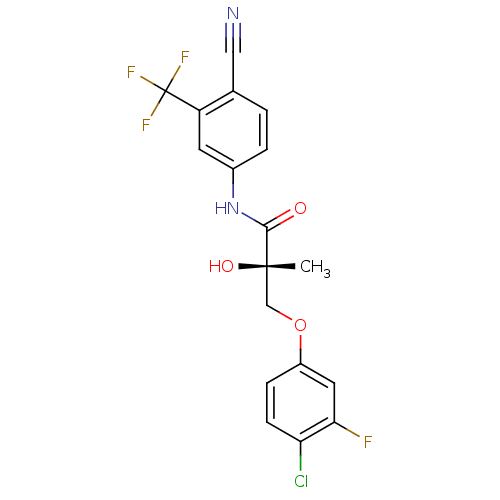
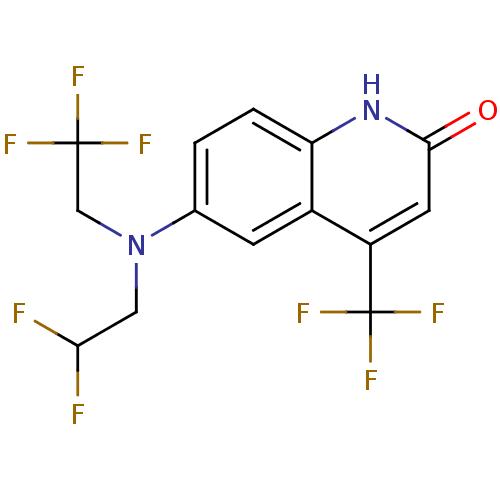
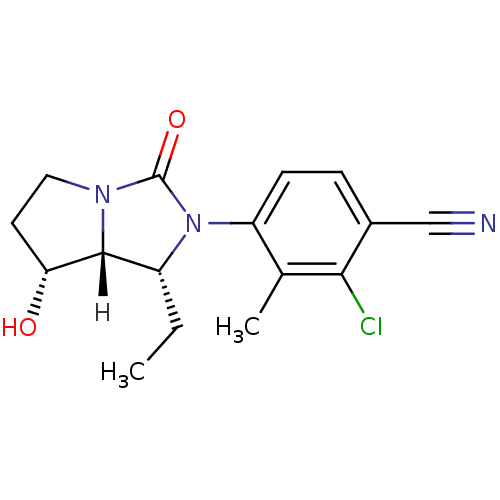
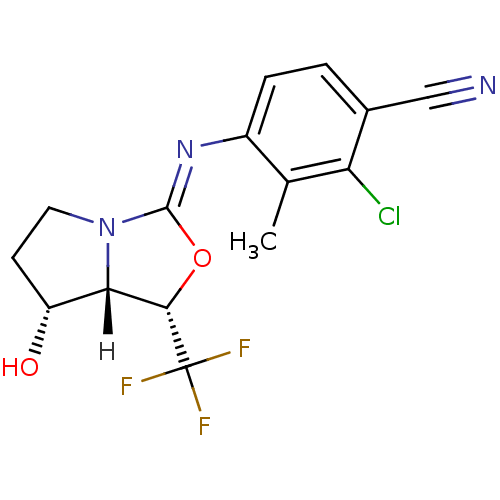
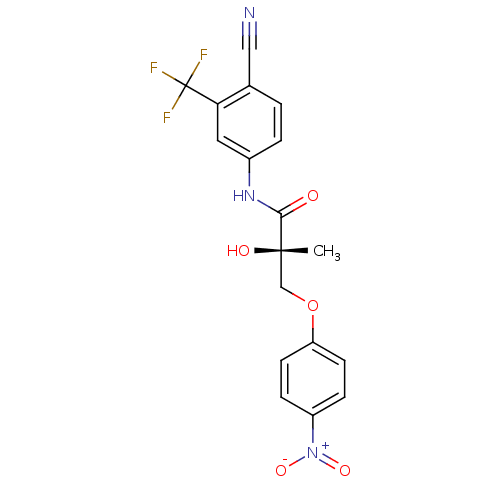
Affinity DataKi: 0.200nM ΔG°: -57.6kJ/mole EC50: 5.10nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -54.8kJ/mole EC50: 19nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -57.6kJ/mole EC50: 5.10nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 0.270nM ΔG°: -50.8kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 0.280nM ΔG°: -50.7kJ/mole EC50: 1nMpH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -50.5kJ/mole EC50: 500nMpH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -56.5kJ/mole EC50: 5.70nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -53.8kJ/mole EC50: 2.80nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -53.8kJ/mole EC50: 1.40nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -53.8kJ/mole EC50: 0.200nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -53.8kJ/mole EC50: 14nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 0.430nM ΔG°: -49.7kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nM ΔG°: -52.6kJ/mole EC50: 2nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nM ΔG°: -55.2kJ/mole EC50: 0.400nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 0.540nM ΔG°: -49.2kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -48.9kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -48.9kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 0.700nM ΔG°: -51.7kJ/mole EC50: 3.70nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 0.700nM ΔG°: -51.7kJ/mole EC50: 2.60nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nM ΔG°: -51.4kJ/mole EC50: 4.80nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 0.860nM ΔG°: -48.1kJ/mole EC50: 500nMpH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nM ΔG°: -51.1kJ/mole EC50: 2.5nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nM ΔG°: -51.1kJ/mole EC50: 1.80nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/mole EC50: 385nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/mole EC50: 2.90nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -53.0kJ/mole EC50: 6.40nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -47.0kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -50.0kJ/mole EC50: 0.700nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -49.9kJ/mole EC50: 281nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -49.9kJ/mole EC50: 5.20nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -49.9kJ/mole EC50: 320nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -52.4kJ/mole EC50: 7.10nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -49.7kJ/mole EC50: 7.10nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -52.2kJ/mole EC50: 1.40nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -49.7kJ/mole EC50: 5.40nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 1.65nM ΔG°: -46.6kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 1.65nM ΔG°: -46.6kJ/mole EC50: 100nMpH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nM ΔG°: -49.6kJ/mole EC50: 270nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nM ΔG°: -46.5kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nM ΔG°: -52.1kJ/mole EC50: 0.200nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 1.90nM ΔG°: -49.3kJ/mole EC50: 44nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 1.90nM ΔG°: -49.3kJ/mole EC50: 48nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.2kJ/mole EC50: 15nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nM ΔG°: -49.0kJ/mole EC50: 1.5nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nM ΔG°: -48.9kJ/mole EC50: 2.10nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nM ΔG°: -48.9kJ/mole EC50: 20nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 2.30nM ΔG°: -48.8kJ/mole EC50: 7.40nMpH: 7.4 T: 2°CAssay Description:Receptor Binding Assay (Ki)-Binding determined through direct displacement of ligand with [3H]-DHT in the MDA-453 cell line. Transactivation Assay (E...More data for this Ligand-Target Pair
Affinity DataKi: 2.30nM ΔG°: -51.3kJ/mole EC50: 0.400nMpH: 7.4 T: 2°CAssay Description:The compounds were evaluated in a transcriptional activation assay with hAR in a mammalian cell background (CV-1) as the primary in vitro assay. Rece...More data for this Ligand-Target Pair
Affinity DataKi: 2.30nM ΔG°: -48.8kJ/mole EC50: 1.10nMpH: 7.6 T: 2°CAssay Description:Binding assays were conducted by incubating test compound at various concentrations with [3H]DHT with human cancer epithelial breast cell lines MDAMB...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nM ΔG°: -45.6kJ/molepH: 7.4 T: 2°CAssay Description:The Ki values were determined by the application of the Cheng-Prusoff equation: Ki = (IC50 x Kd)/(Kd+[L]) where [L] is the concentration of [3H]MIB (...More data for this Ligand-Target Pair

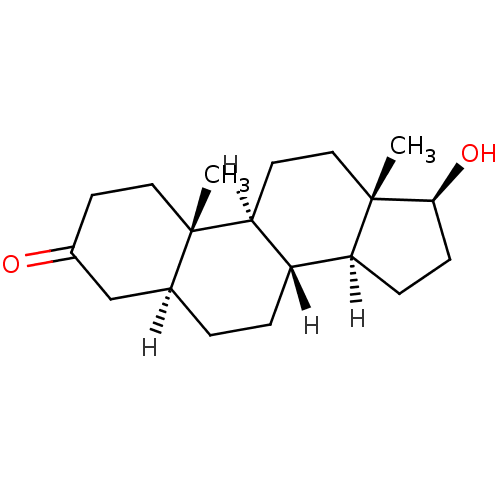
 3D Structure (crystal)
3D Structure (crystal)