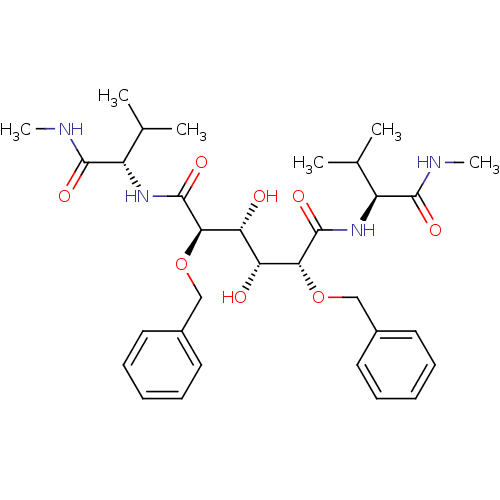
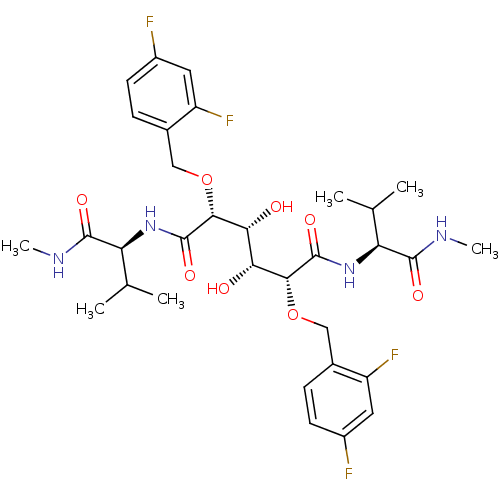
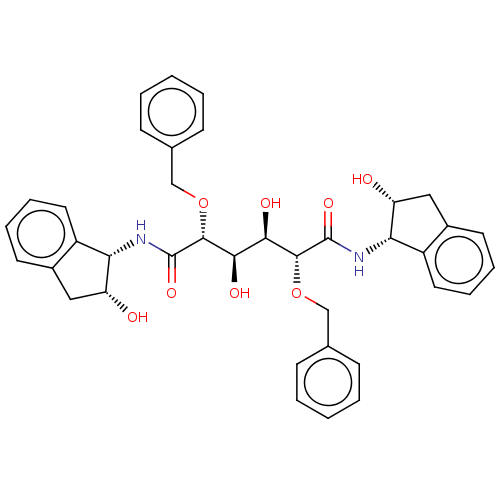
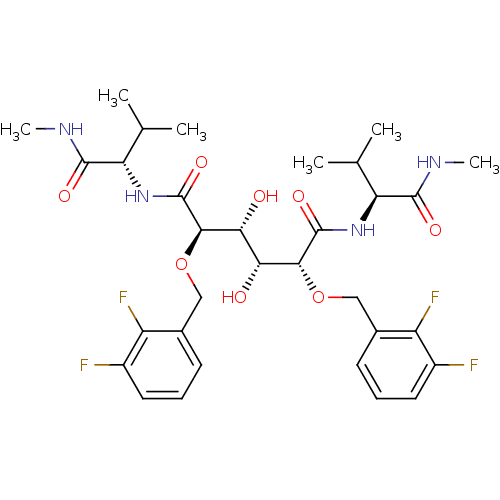
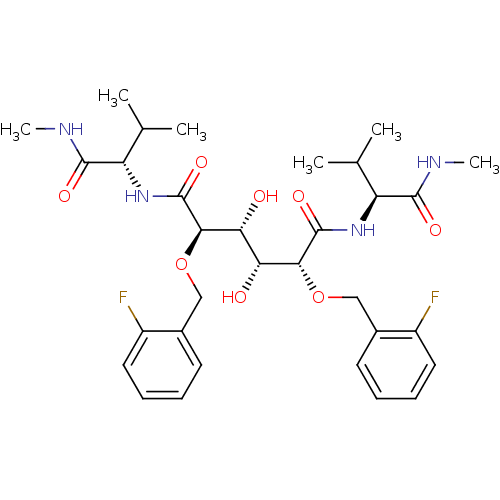
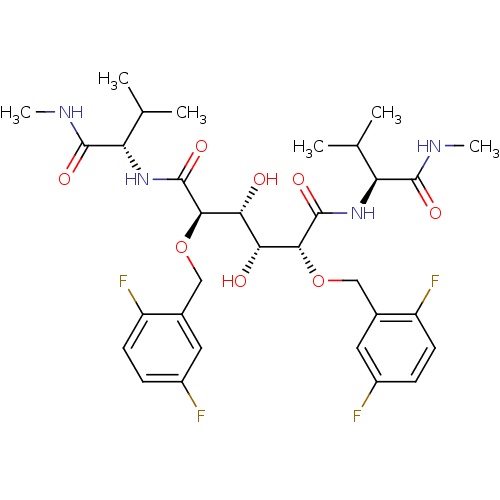
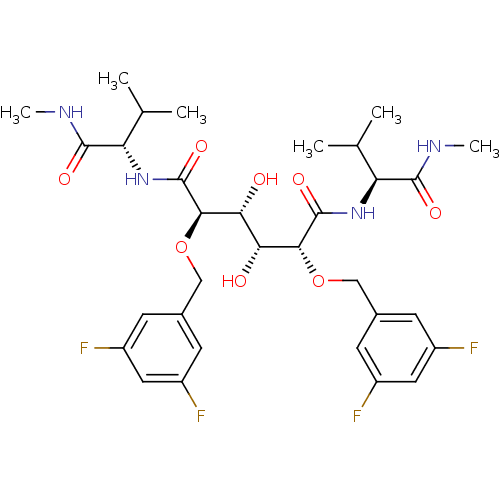
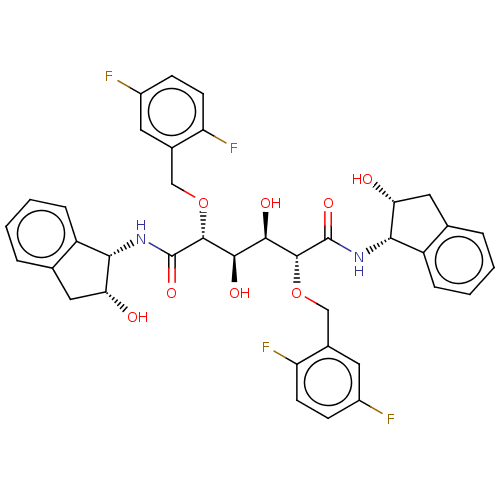
Report error Found 20 Enz. Inhib. hit(s) with all data for entry = 313
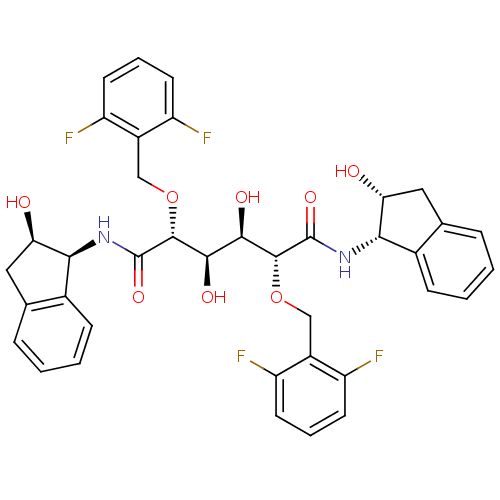
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 0.800nM ΔG°: -52.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
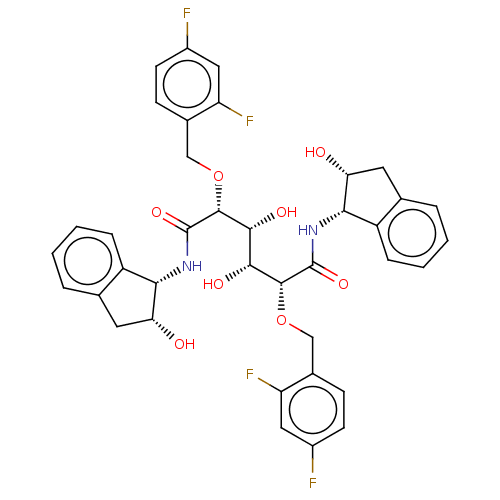
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.12nM ΔG°: -51.9kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
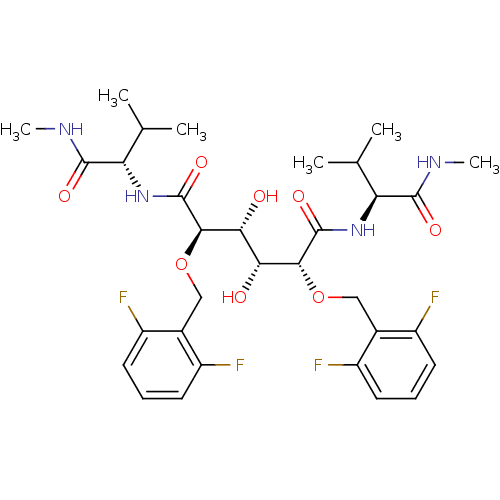
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.22nM ΔG°: -51.7kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
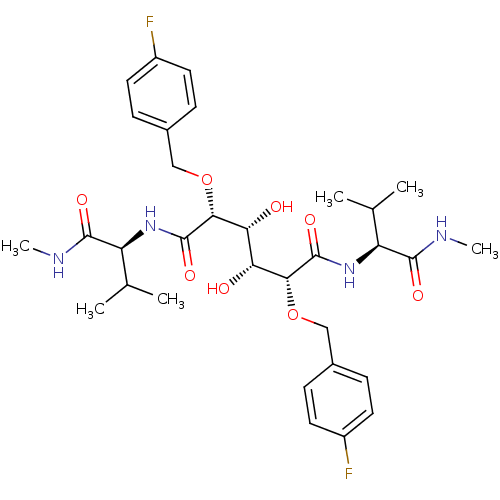
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.28nM ΔG°: -51.6kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.64nM ΔG°: -51.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.65nM ΔG°: -51.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.73nM ΔG°: -50.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.79nM ΔG°: -50.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.83nM ΔG°: -50.7kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.92nM ΔG°: -50.6kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.98nM ΔG°: -50.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.98nM ΔG°: -50.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 1.98nM ΔG°: -50.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 3.26nM ΔG°: -49.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 3.29nM ΔG°: -49.2kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 3.90nM ΔG°: -48.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 4nM ΔG°: -48.7kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 5nM ΔG°: -48.2kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 5.39nM ΔG°: -48.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Linkoping University
Linkoping University
Affinity DataKi: 7.13nM ΔG°: -47.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair