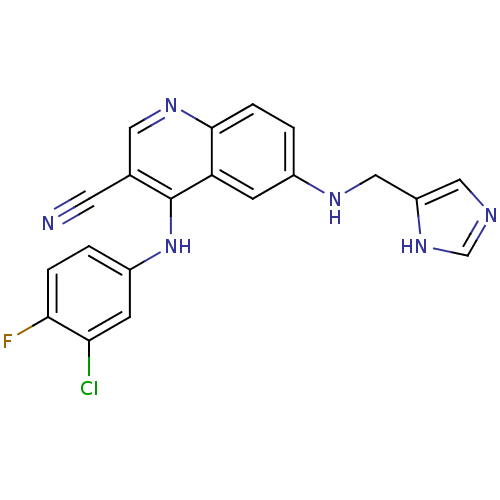
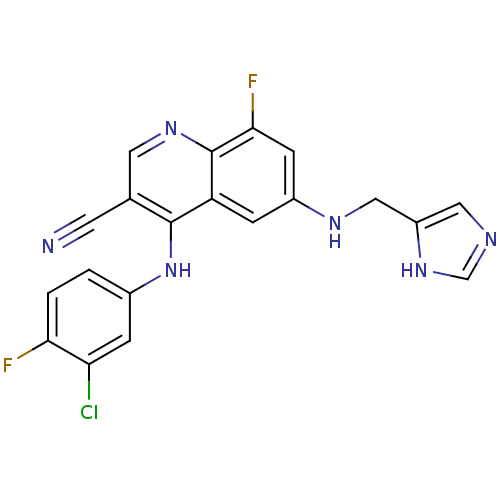
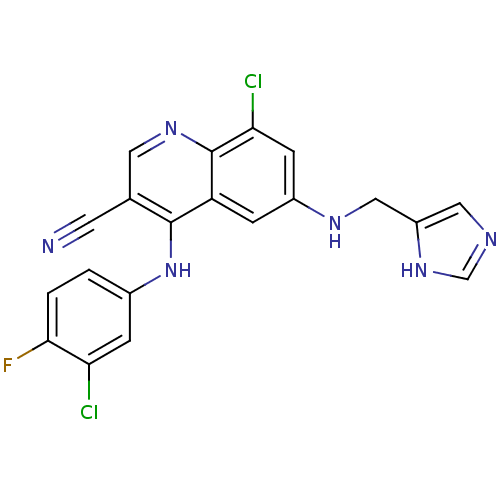
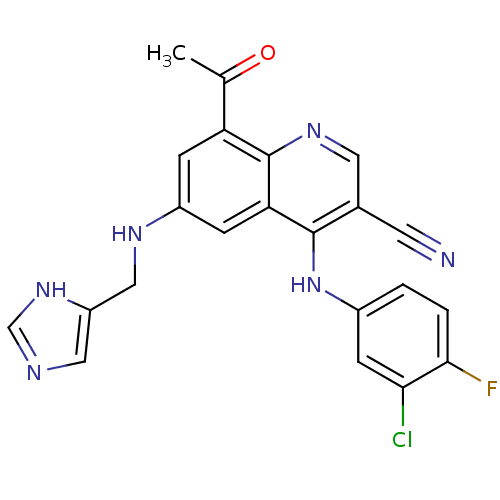
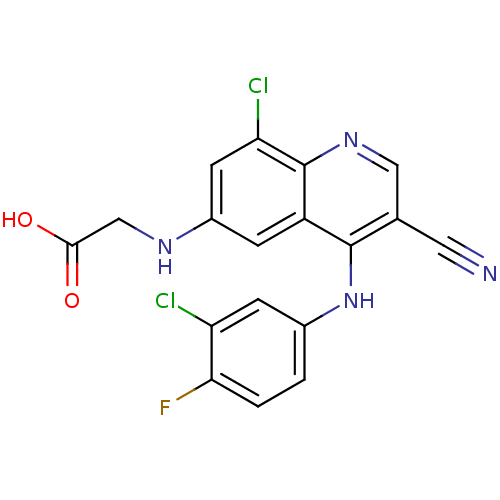
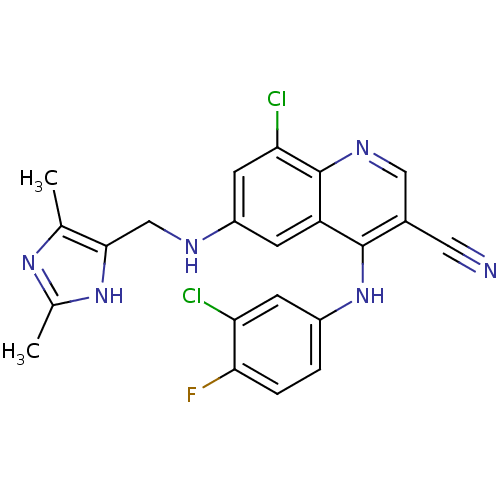
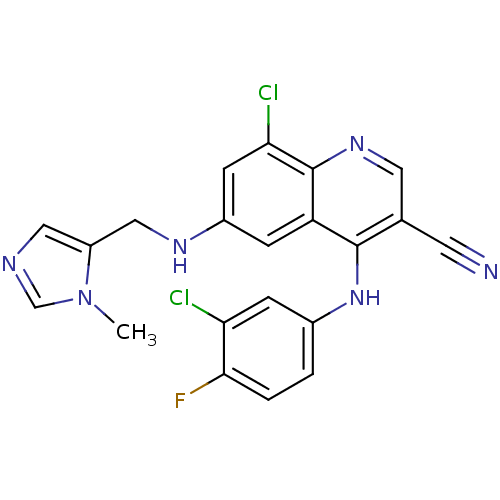
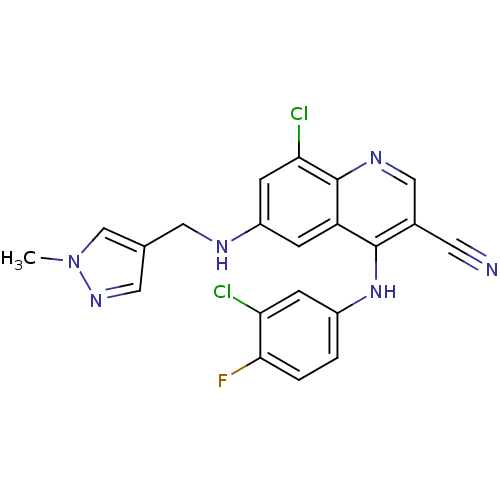
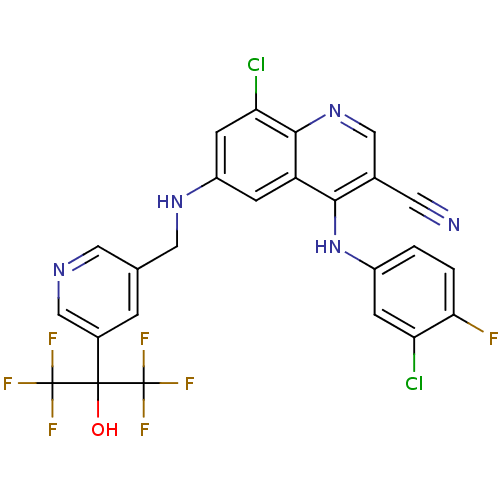
Report error Found 57 Enz. Inhib. hit(s) with all data for entry = 2552
Affinity DataIC50: 2nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:EGFR kinase activity was assayed using poly(Glu4-Tyr) as a substrate. The phosphorylation of tyrosine residue was detected by an ELISA. IC50 calculat...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 98nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 156nMAssay Description:EGFR kinase activity was assayed using poly(Glu4-Tyr) as a substrate. The phosphorylation of tyrosine residue was detected by an ELISA. IC50 calculat...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 280nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:EGFR kinase activity was assayed using poly(Glu4-Tyr) as a substrate. The phosphorylation of tyrosine residue was detected by an ELISA. IC50 calculat...More data for this Ligand-Target Pair
Affinity DataIC50: 590nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 830nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 910nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:EGFR kinase activity was assayed using poly(Glu4-Tyr) as a substrate. The phosphorylation of tyrosine residue was detected by an ELISA. IC50 calculat...More data for this Ligand-Target Pair
Affinity DataIC50: 2.08E+3nMAssay Description:EGFR kinase activity was assayed using poly(Glu4-Tyr) as a substrate. The phosphorylation of tyrosine residue was detected by an ELISA. IC50 calculat...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+3nMAssay Description:EGFR kinase activity was assayed using poly(Glu4-Tyr) as a substrate. The phosphorylation of tyrosine residue was detected by an ELISA. IC50 calculat...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:EGFR kinase activity was assayed using poly(Glu4-Tyr) as a substrate. The phosphorylation of tyrosine residue was detected by an ELISA. IC50 calculat...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMpH: 7.2 T: 2°CAssay Description:Tpl2/Cot activity was directly assayed using GST-MEK1 as a substrate. The phosphorylation on serine residues was detected by an ELISA. IC50 calculati...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:EGFR kinase activity was assayed using poly(Glu4-Tyr) as a substrate. The phosphorylation of tyrosine residue was detected by an ELISA. IC50 calculat...More data for this Ligand-Target Pair