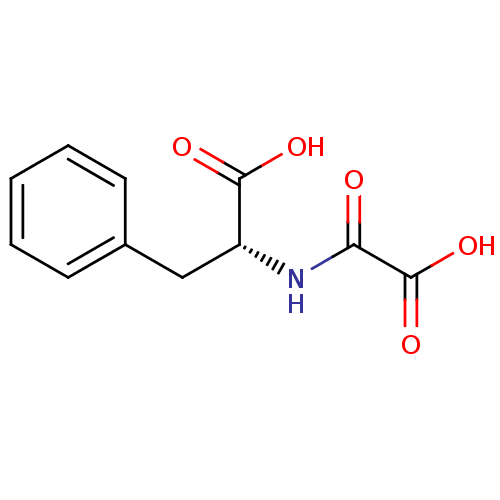
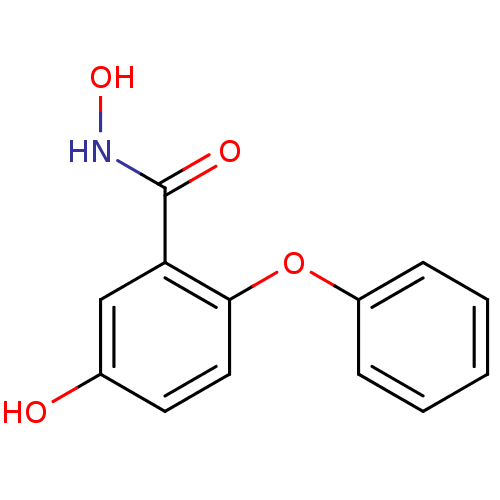
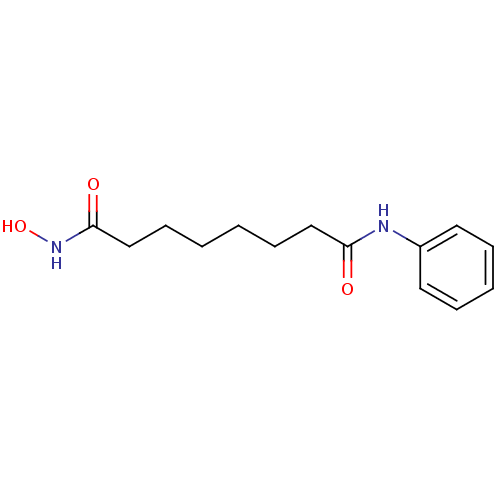
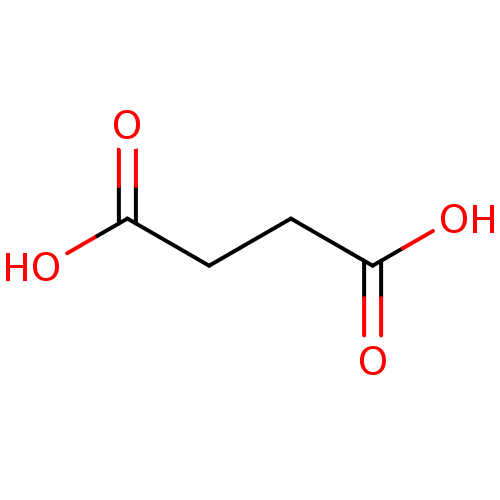
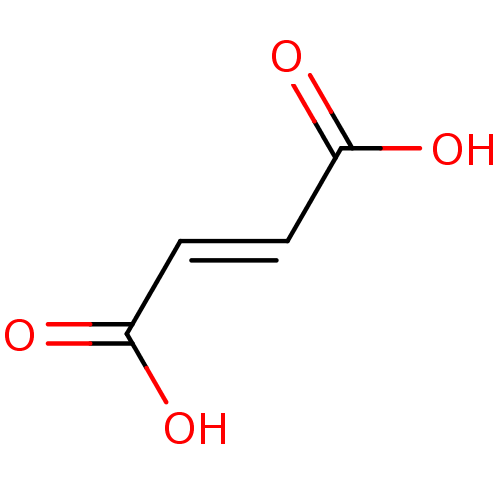
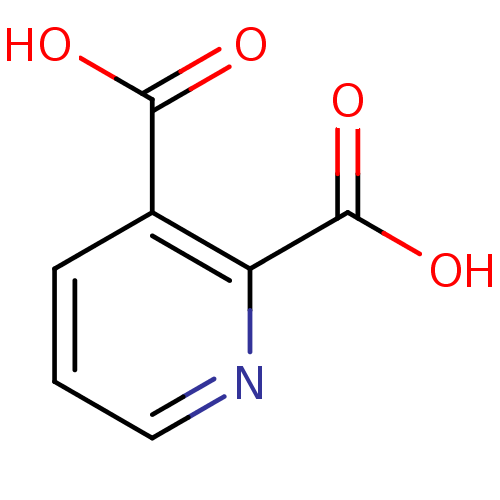
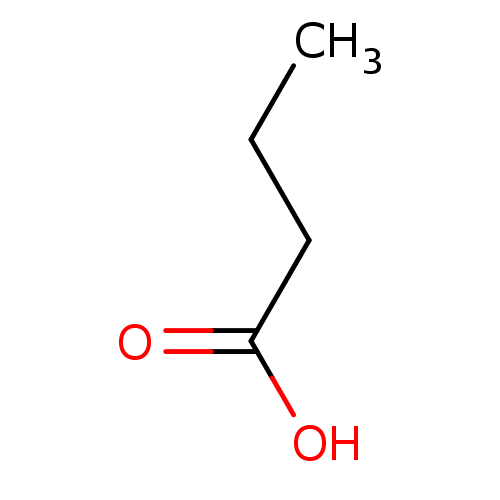
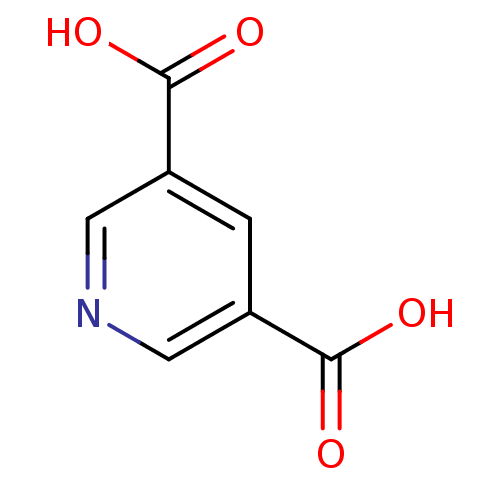
Report error Found 19 Enz. Inhib. hit(s) with all data for entry = 2925
Affinity DataIC50: 1.40E+3nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+3nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMpH: 7.5 T: 2°CAssay Description:For compounds which were shown to inhibit FDH, a MALDI-TOF-MS inhibition assay was employed. After incubation, the diluted assay mixture was then mix...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80E+4nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+5nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+5nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+5nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+5nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+5nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+5nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 7.10E+5nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+6nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+6nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+7nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+7nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+7nMpH: 7.5 T: 2°CAssay Description:A coupled-assay for JMJD2E activity employing formaldehyde dehydrogenase (FDH) from Pseudomonas putida was developed. Formaldehyde release by demethy...More data for this Ligand-Target Pair
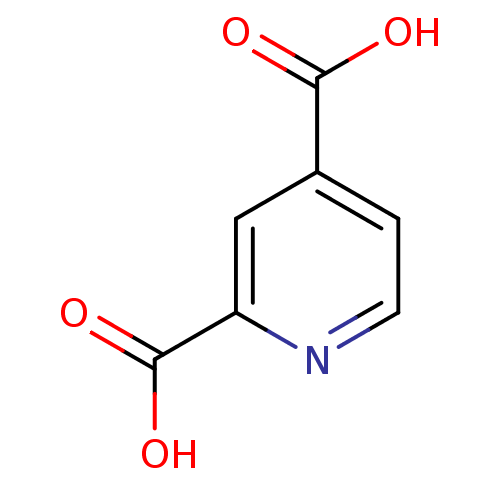
 3D Structure (crystal)
3D Structure (crystal)