Report error Found 43 Enz. Inhib. hit(s) with all data for entry = 3194
Affinity DataIC50: 100nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 270nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 760nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 960nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+3nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 2.02E+3nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 2.72E+3nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 3.92E+3nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 5.68E+3nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 5.76E+3nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 6.20E+3nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+3nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 8.80E+3nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+3nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+4nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.82E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.85E+4nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 2.41E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 2.68E+4nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 2.93E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 3.15E+4nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 5.41E+4nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 9.13E+4nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 9.84E+4nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 9.85E+4nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+5nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+5nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+5nMAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMpH: 7.5 T: 2°CAssay Description:The activity of QR2 under steady-state conditions was evaluated on SpectraMax Plus 384 UV/vis spectrophotometer by monitoring the increase in absorba...More data for this Ligand-Target Pair
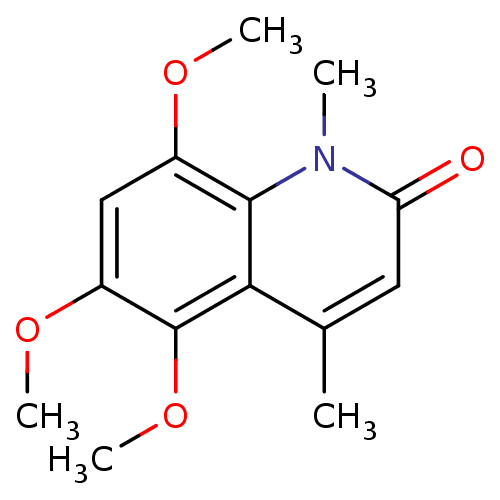
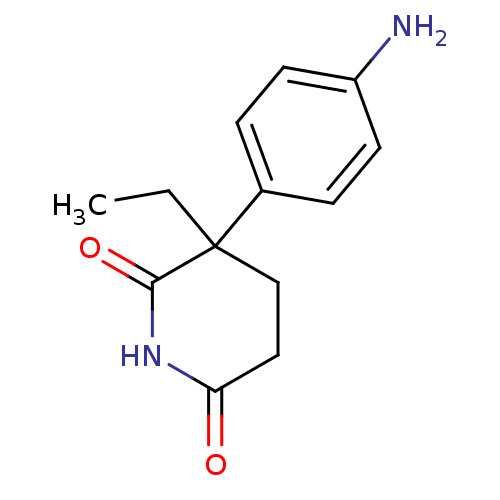

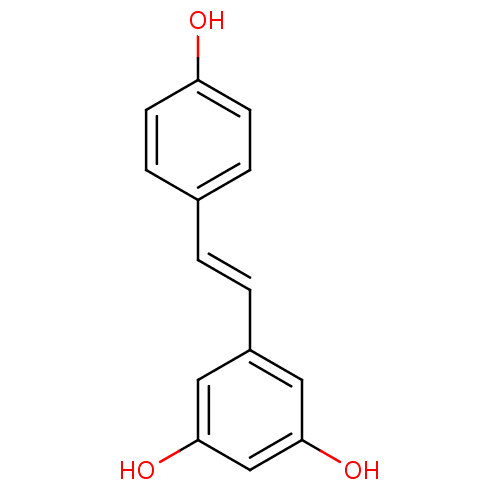
 3D Structure (crystal)
3D Structure (crystal)