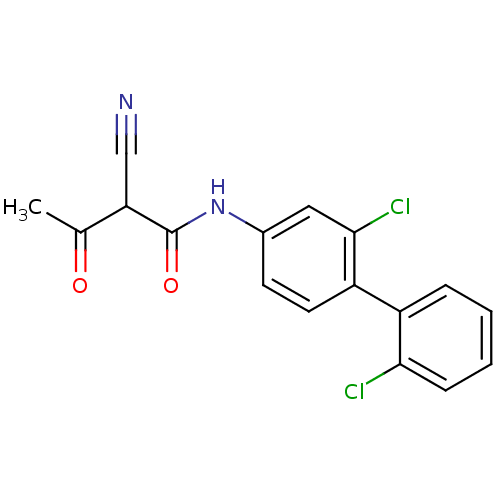
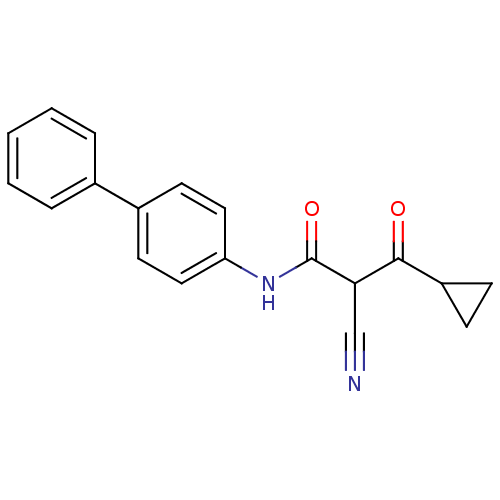
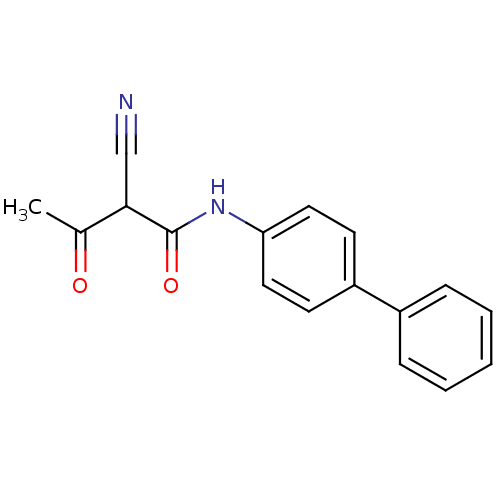
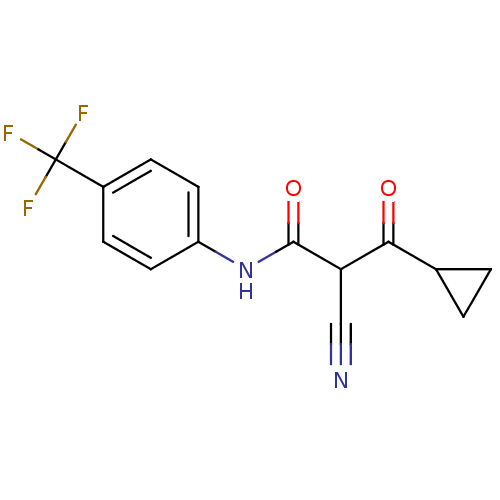
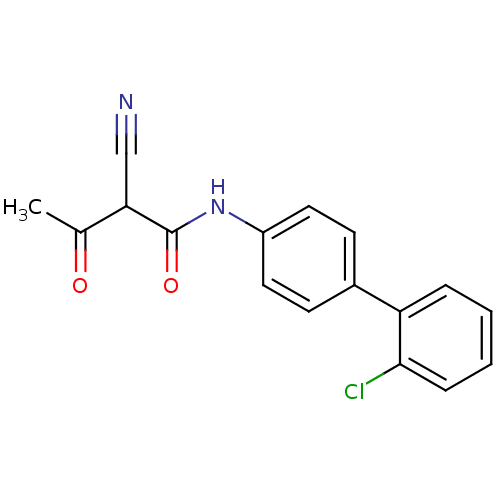
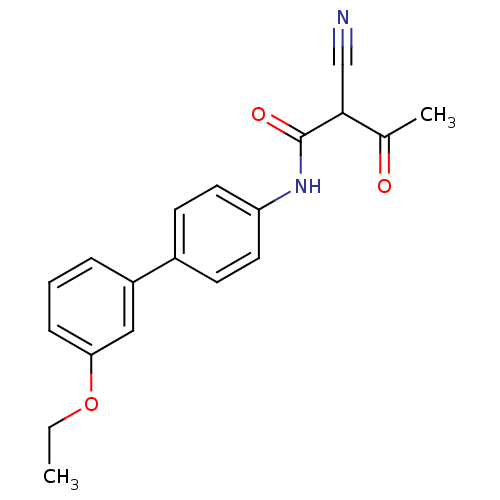
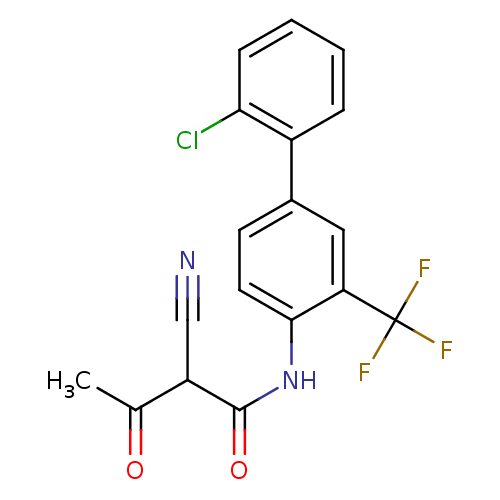
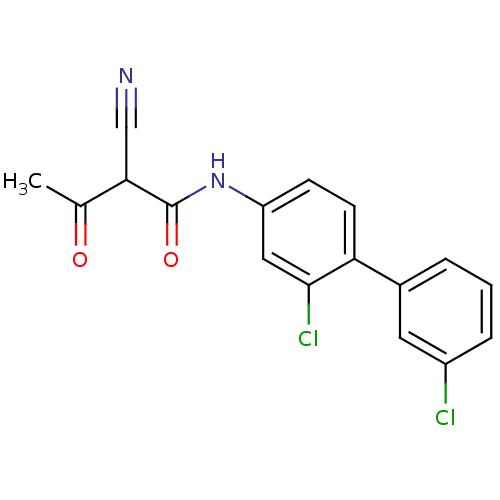
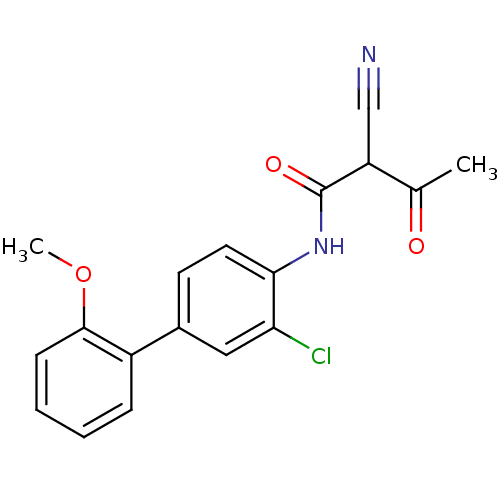
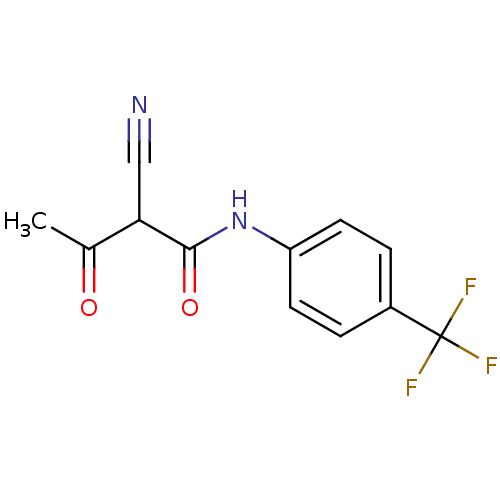
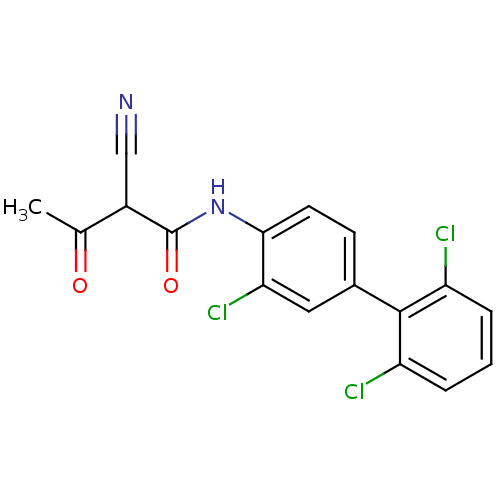
Report error Found 34 Enz. Inhib. hit(s) with all data for entry = 3177
Affinity DataKi: 2.70nM IC50: 22nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 7nM IC50: 53nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 11nM IC50: 90nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 14nM IC50: 117nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 16nM IC50: 130nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 16nM IC50: 130nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 21nM IC50: 170nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 22nM IC50: 180nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 23nM IC50: 190nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 25nM IC50: 200nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 25nM IC50: 200nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 32nM IC50: 261nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 39nM IC50: 320nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 40nM IC50: 330nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 190nM IC50: 1.52E+3nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 480nM ΔG°: -36.1kJ/mole IC50: 4.00E+3nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 730nM ΔG°: -35.0kJ/mole IC50: 6.20E+3nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 1.00E+3nM ΔG°: -34.2kJ/mole IC50: 8.60E+3nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 1.20E+3nM ΔG°: -33.8kJ/mole IC50: 9.90E+3nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 1.20E+3nM ΔG°: -33.8kJ/mole IC50: 1.02E+4nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nM IC50: 1.65E+4nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 3.00E+3nM ΔG°: -31.5kJ/mole IC50: 2.57E+4nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 3.20E+3nM ΔG°: -31.4kJ/mole IC50: 2.72E+4nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 5.60E+3nM ΔG°: -30.0kJ/mole IC50: 4.74E+4nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 5.80E+3nM ΔG°: -29.9kJ/mole IC50: 4.91E+4nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 6.00E+3nM ΔG°: -29.8kJ/mole IC50: 5.11E+4nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 6.80E+3nM ΔG°: -29.5kJ/mole IC50: 5.72E+4nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
Affinity DataKi: 1.07E+4nM IC50: 8.66E+4nMAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 1.10E+4nM ΔG°: -28.3kJ/mole IC50: 9.25E+4nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 2.05E+4nM ΔG°: -26.8kJ/mole IC50: 1.65E+5nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 2.15E+4nM ΔG°: -26.6kJ/mole IC50: 1.82E+5nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 2.20E+4nM ΔG°: -26.6kJ/mole IC50: 1.90E+5nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 2.60E+4nM ΔG°: -26.2kJ/mole IC50: 2.24E+5nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair
TargetDihydroorotate dehydrogenase (quinone), mitochondrial(Plasmodium falciparum (isolate 3D7))
University of Leeds
University of Leeds
Affinity DataKi: 2.80E+4nM ΔG°: -26.0kJ/mole IC50: 2.37E+5nMpH: 8.0 T: 2°CAssay Description:The assays were carried out by using a colorimetric DCIP method, which uses the colorimetric reagent 2, 6-dichlorophenolindophenol as the final elect...More data for this Ligand-Target Pair