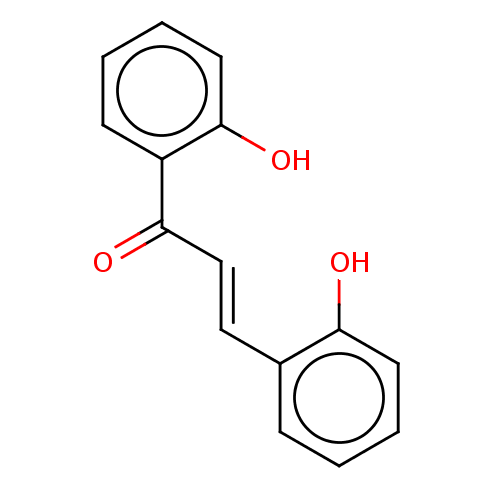
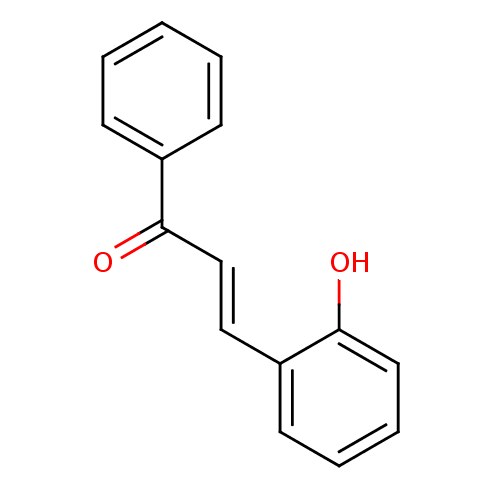
Report error Found 23 Enz. Inhib. hit(s) with all data for entry = 8162
Affinity DataIC50: 49nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 270nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 620nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 1.71E+3nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 1.94E+3nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 2.08E+3nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 2.37E+3nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 2.45E+3nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 2.63E+3nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 8.03E+3nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 1.14E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 1.18E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 1.56E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 1.94E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 2.18E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 3.04E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 4.18E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 5.81E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+4nMpH: 4.5 T: 2°CAssay Description:The assay based on fluorescenceresonance energy transfer was carried out with BACE1 enzyme at pH 4.5 with a substrate, H-Lys(DABSYL)-SEVNLDAEFR-Gin-(...More data for this Ligand-Target Pair