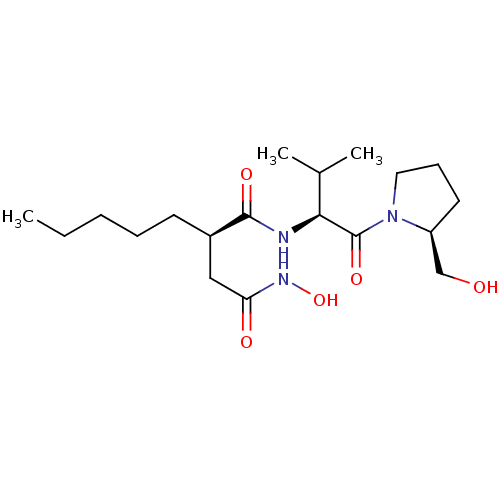
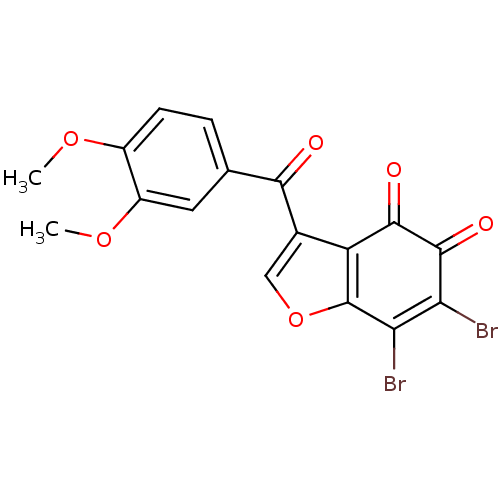
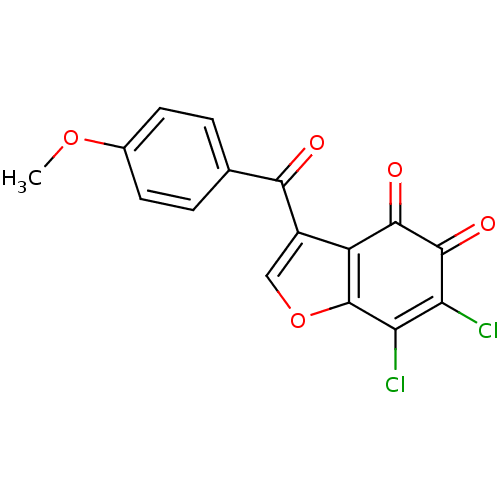
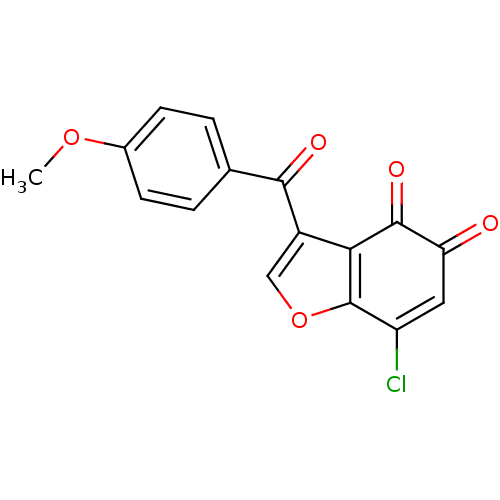
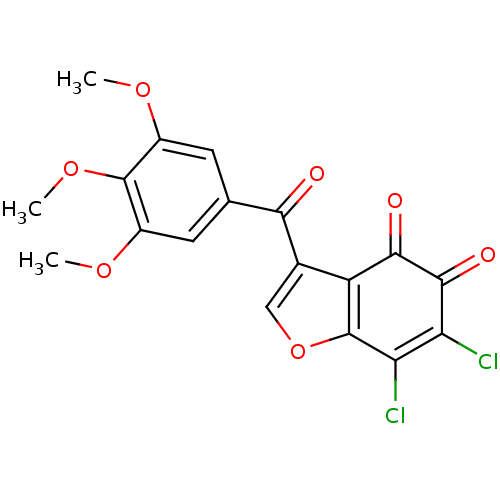
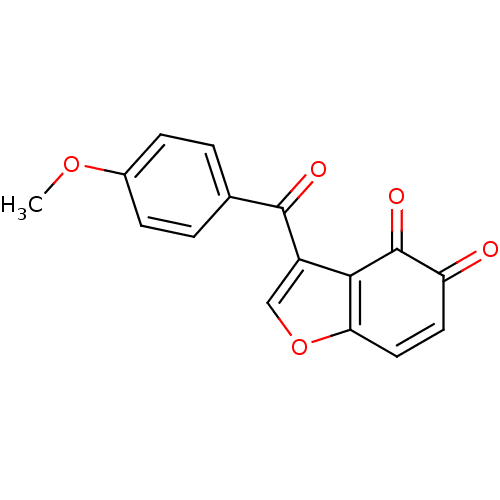
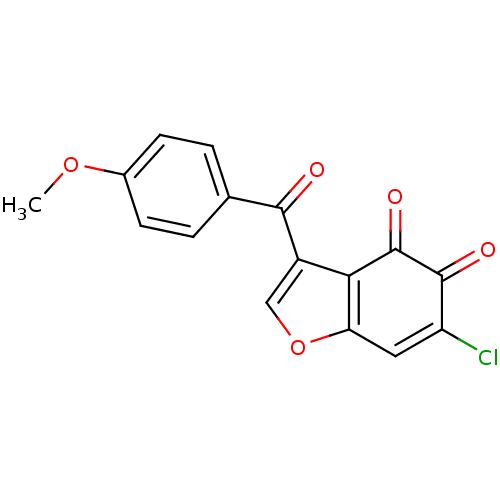
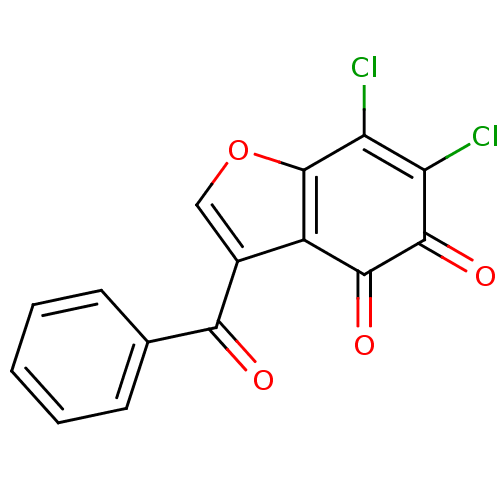
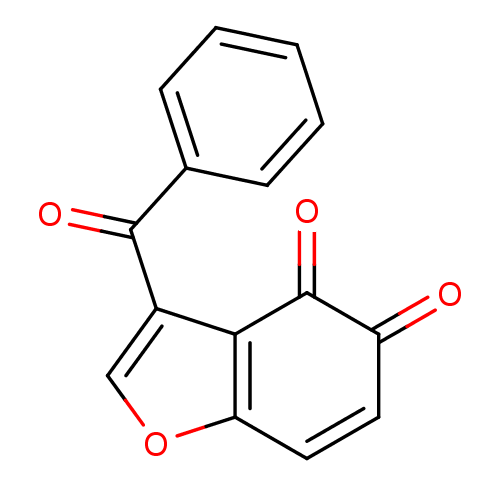
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 50033652
Affinity DataIC50: 140nMAssay Description:Inhibition of Escherichia coli peptide deformylase using fMAHA as substrate after 1 hr by FLUO assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 2.70E+3nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 5.20E+3nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 6.10E+3nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 4.50E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 5.90E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
TargetPeptide deformylase, mitochondrial(Human)
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Memorial Sloan-Kettering Cancer Center
Curated by ChEMBL
Affinity DataIC50: 6.50E+4nMAssay Description:Inhibition of human peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.70E+4nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli peptide deformylase after 1 hr by fluorescence polarization based competition assayMore data for this Ligand-Target Pair