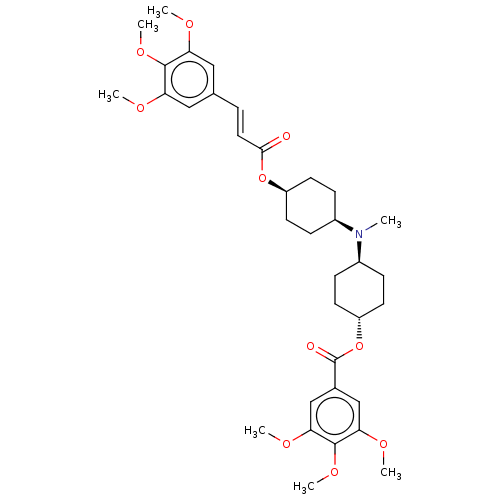
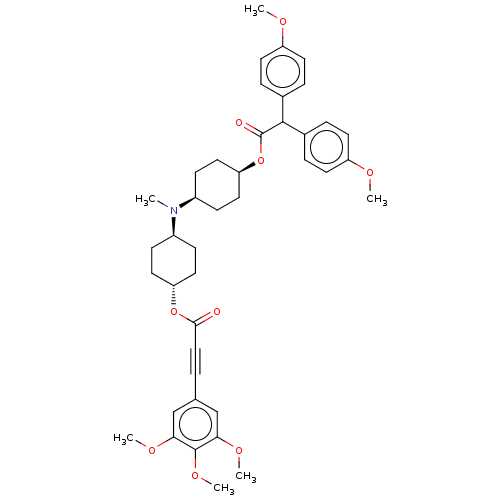
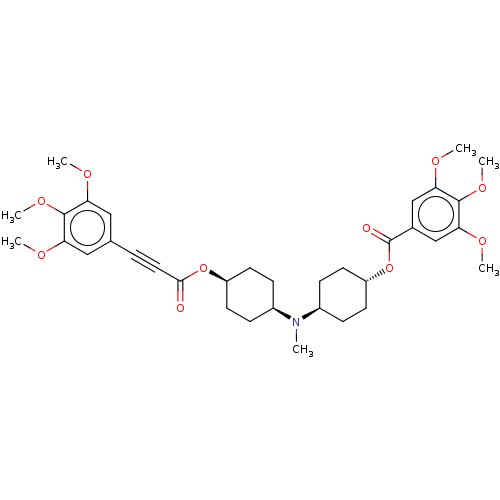
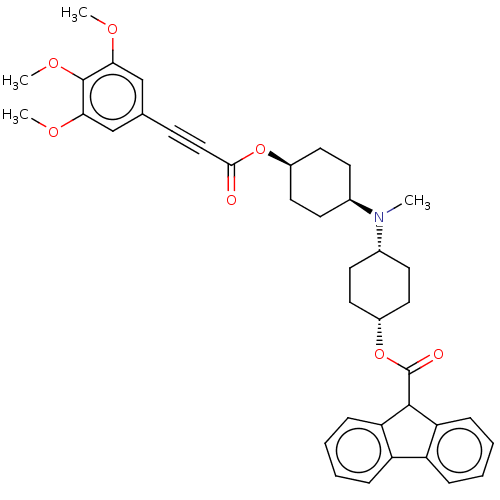
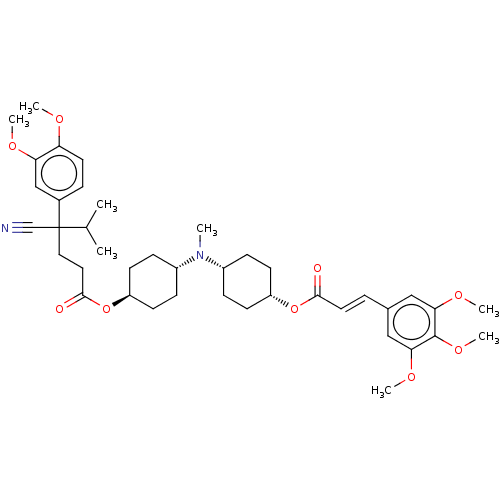
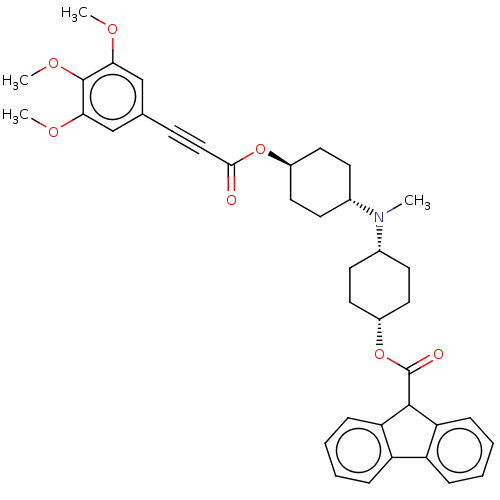
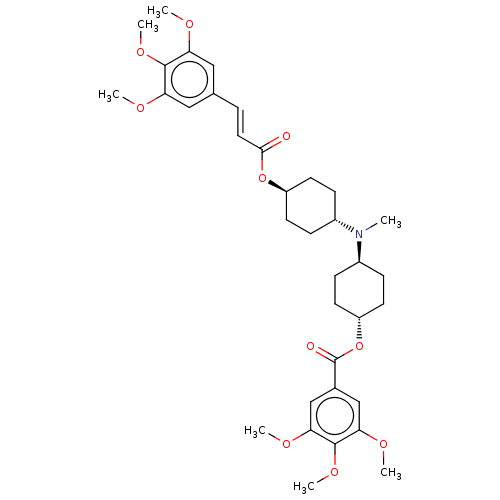
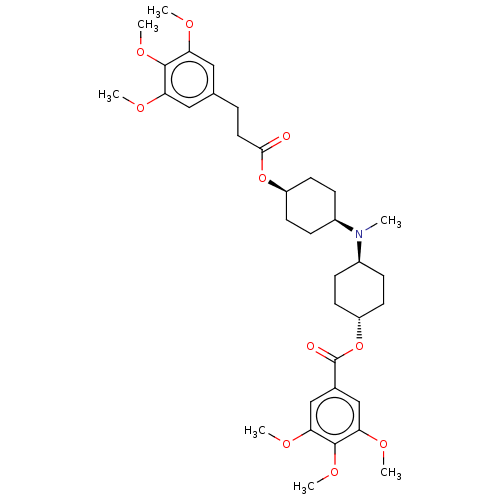
Report error Found 30 Enz. Inhib. hit(s) with all data for entry = 50005243
Affinity DataIC50: 12nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 75nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 92nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 610nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 630nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of P-glycoprotein in human doxorubicin-resistant K562 cells assessed as half maximal increase of pirarubicin accumulation by spectrofluoro...More data for this Ligand-Target Pair