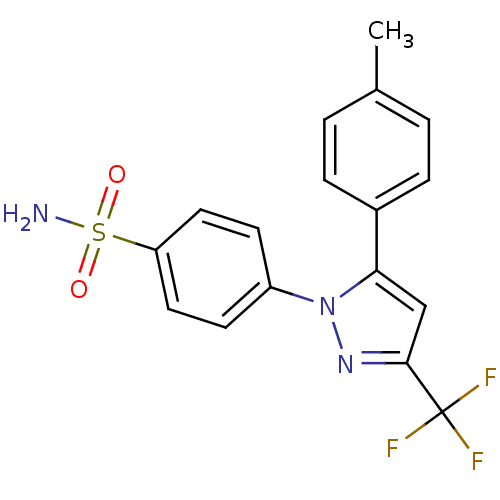
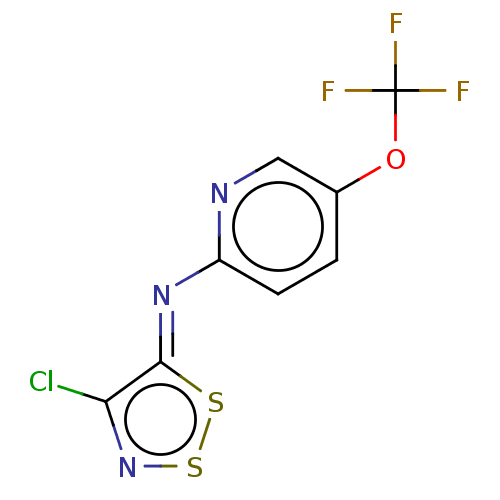
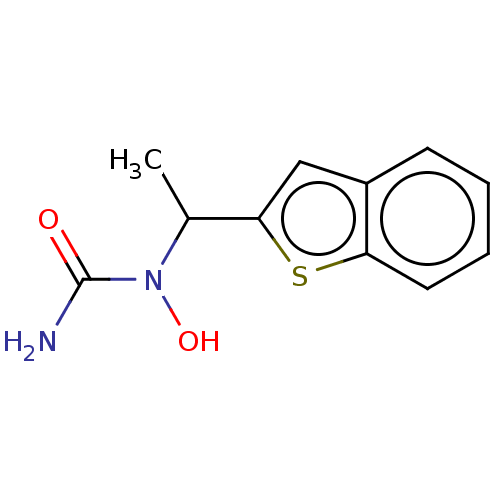
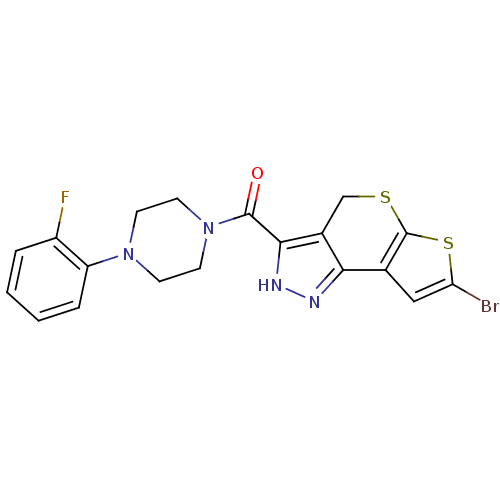
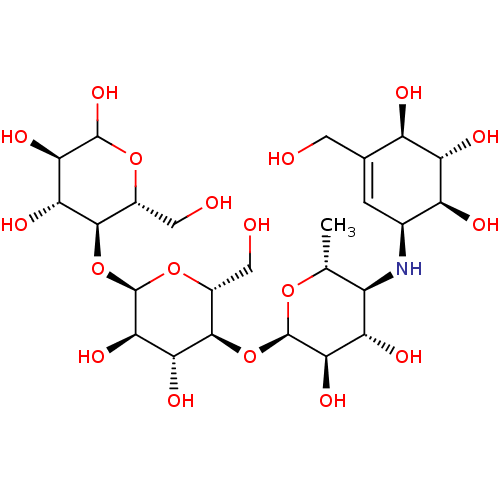
Report error Found 9 Enz. Inhib. hit(s) with all data for entry = 50009408
Target72 kDa type IV collagenase(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataIC50: 620nMAssay Description:Inhibition of MMP2 catalytic domain (unknown origin) using Ac-PLG-[2-mercapto-4-methylpentanoyl]-LG-OC2H5 as substrate preincubated for 45 to 120 min...More data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 2(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataIC50: 840nMAssay Description:Inhibition of COX2 (unknown origin) by EIAMore data for this Ligand-Target Pair
TargetProstaglandin G/H synthase 2(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataIC50: 880nMAssay Description:Inhibition of COX2 (unknown origin) by EIAMore data for this Ligand-Target Pair
TargetLarge neutral amino acids transporter small subunit 1(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataIC50: 980nMAssay Description:Inhibition of recombinant human LAT1More data for this Ligand-Target Pair
TargetProbable maltase-glucoamylase 2(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataIC50: 2.85E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl alpha-d-glucopyranoside as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
TargetPolyunsaturated fatty acid 5-lipoxygenase(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of 5-LOX (unknown origin)More data for this Ligand-Target Pair
TargetPolyunsaturated fatty acid 5-lipoxygenase(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataIC50: 5.70E+3nMAssay Description:Inhibition of 5-LOX (unknown origin)More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor gamma(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataEC50: 5.90E+3nMAssay Description:Inhibition of PPARgamma (unknown origin)More data for this Ligand-Target Pair
TargetProbable maltase-glucoamylase 2(Human)
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Maharaja Ranjit Singh Punjab Technical University
Curated by ChEMBL
Affinity DataIC50: 8.17E+5nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl alpha-d-glucopyranoside as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair