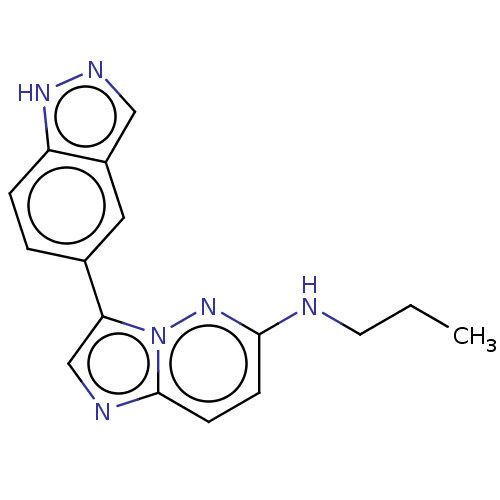
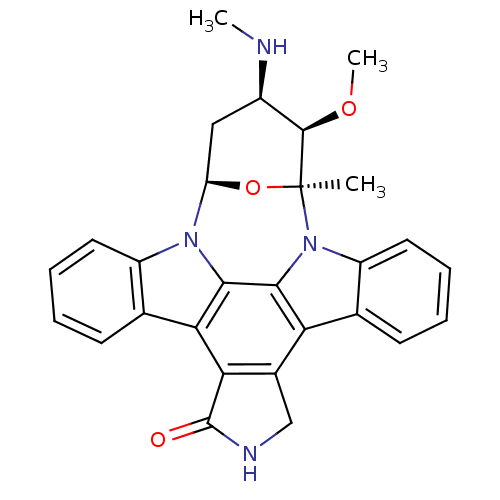
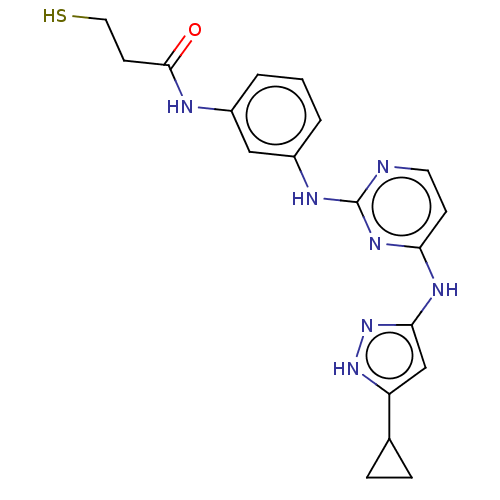
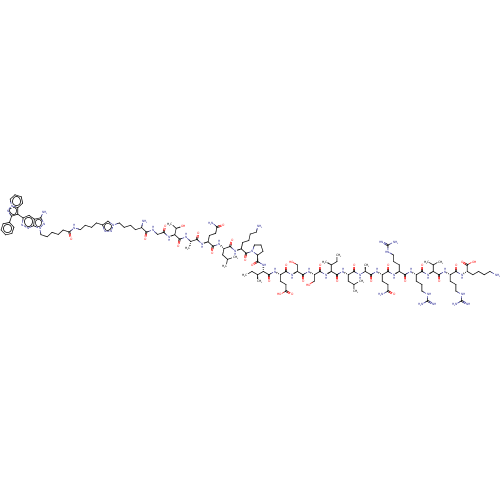
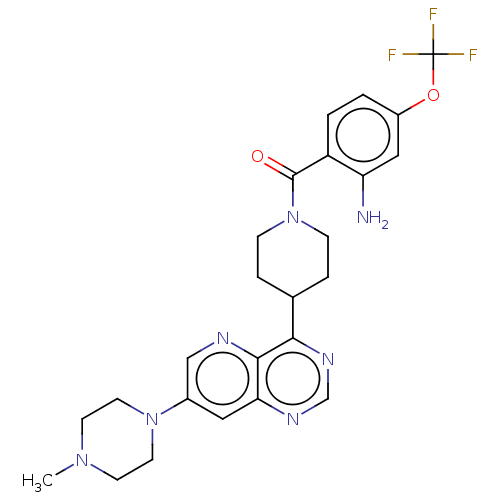
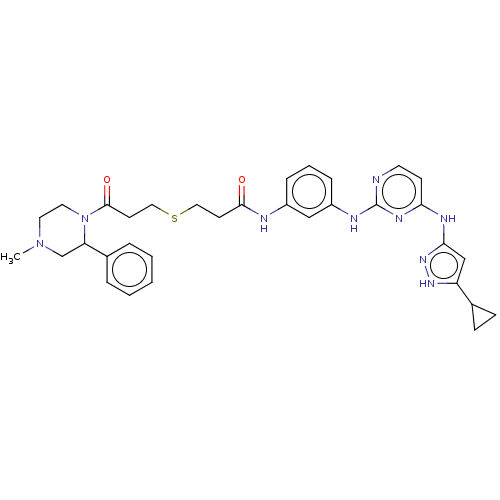
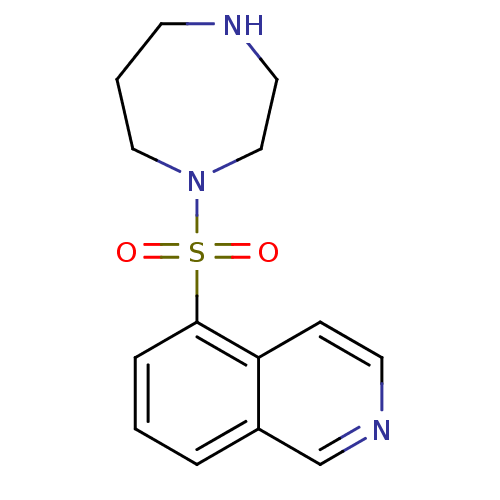
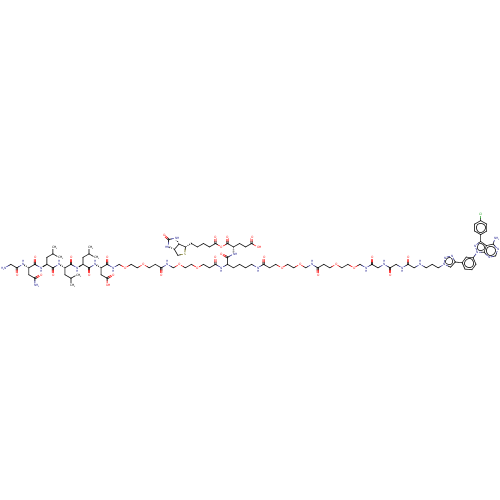
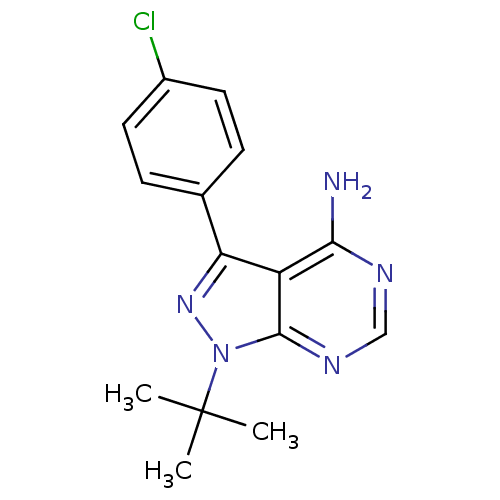
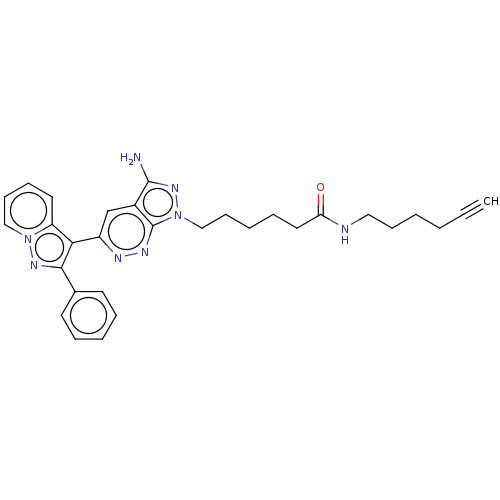
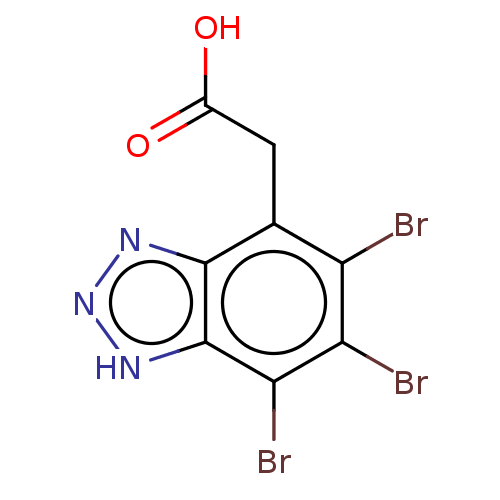
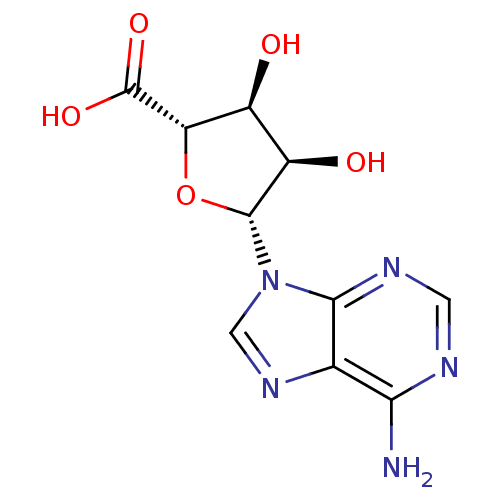
Report error Found 35 Enz. Inhib. hit(s) with all data for entry = 50015607
Affinity DataKd: 0.0130nMAssay Description:Binding affinity to human wild-type PKAc beta assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataKd: 0.0190nMAssay Description:Binding affinity to haspin (unknown origin) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataKd: 0.0190nMAssay Description:Binding affinity to a TEV-cleavable N-terminal His6-tagged human recombinant haspin (470 to 798 residues) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataKd: 0.0420nMAssay Description:Displacement of mAb(D38C6)-BTN from human recombinant PKAc alpha incubated for 15 mins by microplate reader assayMore data for this Ligand-Target Pair
Affinity DataKd: 0.100nMAssay Description:Binding affinity to PKA (unknown origin) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataKd: 0.280nMAssay Description:Binding affinity to chicken 6-His tagged c-Src expressed in Escherichia coli Bl21 (DE3) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataKd: 0.420nMAssay Description:Binding affinity to N-terminal his6-tagged human recombinant Haspin (470 to 798 residues) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of human recombinant CK2alphaMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of mTOR (unknown origin) by HTRF assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of haspin (unknown origin) by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:Inhibition of PKA (unknown origin)More data for this Ligand-Target Pair
Affinity DataKd: 4nMAssay Description:Binding affinity to a TEV-cleavable N-terminal His6-tagged human recombinant haspin (470 to 798 residues) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of PKA (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Irreversible inhibition of wild type c-Src (unknown origin) assessed as inhibition constant by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of human recombinant ERK2 expressed in Escherichia coli DH5alpha incubated for 1 hrMore data for this Ligand-Target Pair
Affinity DataKd: 30nMAssay Description:Inhibition of ROCK2 (unknown origin) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataIC50: 35nMAssay Description:Activation of ERK5 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:Inhibition of CK2alpha in human MIA PaCa-2 cells incubated for 5 hrs by western blot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 90nMAssay Description:Irreversible inhibition of wild type c-Src (unknown origin) assessed as inhibition constant by fluorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Activation of ERK5 in MEF cellsMore data for this Ligand-Target Pair
Affinity DataKd: 150nMAssay Description:Binding affinity to N-terminal his6-tagged human recombinant Haspin (470 to 798 residues) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataIC50: 158nMAssay Description:Inhibition of ROCK2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 159nMAssay Description:Inhibition of PKA (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of c-Src (unknown origin)More data for this Ligand-Target Pair
Affinity DataKd: 170nMAssay Description:Binding affinity to N-terminal his6-tagged human recombinant Haspin (470 to 798 residues) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataKd: 250nMAssay Description:Inhibition of EPHA3 (unknown origin) assessed as dissociation constant by ELISA-like binding assayMore data for this Ligand-Target Pair
Affinity DataKd: 296nMAssay Description:Binding affinity to chicken 6-His tagged c-Src expressed in Escherichia coli Bl21 (DE3) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataIC50: 670nMAssay Description:Inhibition of human CK2More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of human recombinant ERK2 expressed in Escherichia coli DH5alpha incubated for 1 hrMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of c-Src (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4.58E+3nMAssay Description:Inhibition of PKA (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of human CK2More data for this Ligand-Target Pair
Affinity DataKd: >1.50E+4nMAssay Description:Binding affinity to a TEV-cleavable N-terminal His6-tagged human recombinant haspin (470 to 798 residues) assessed as dissociation constantMore data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+4nMAssay Description:Inhibition of PKA (unknown origin)More data for this Ligand-Target Pair
Affinity DataKd: 2.00E+5nMAssay Description:Inhibition of EPHA3 (unknown origin) assessed as dissociation constant by ELISA-like binding assayMore data for this Ligand-Target Pair
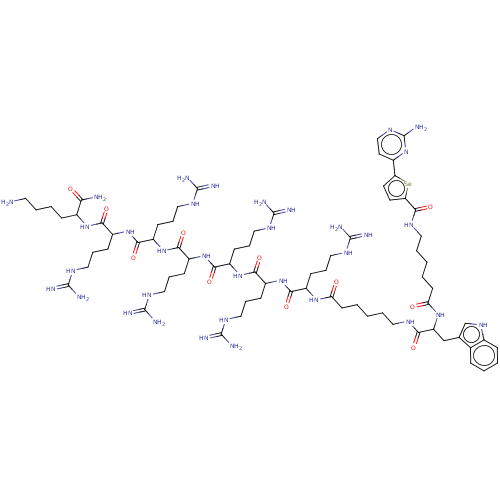

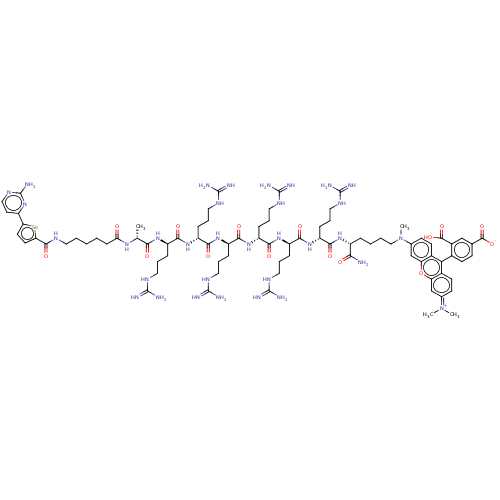
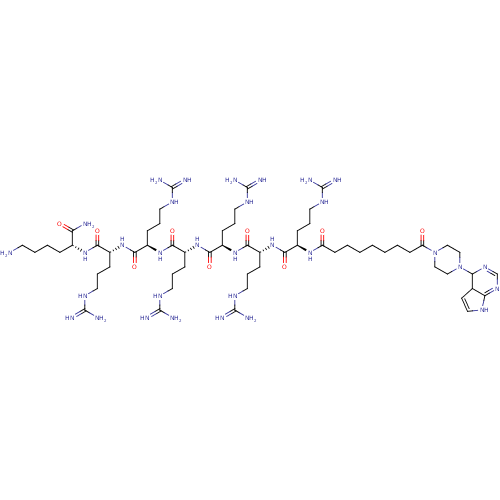
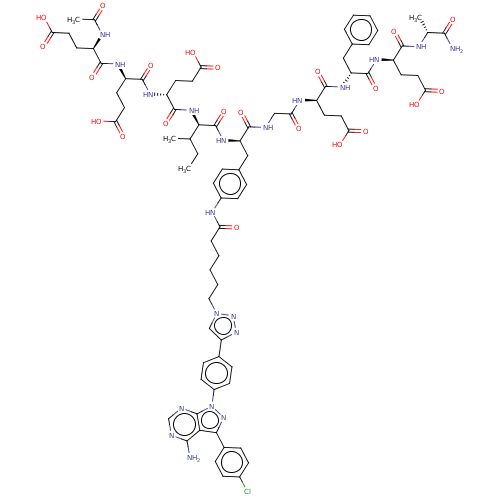
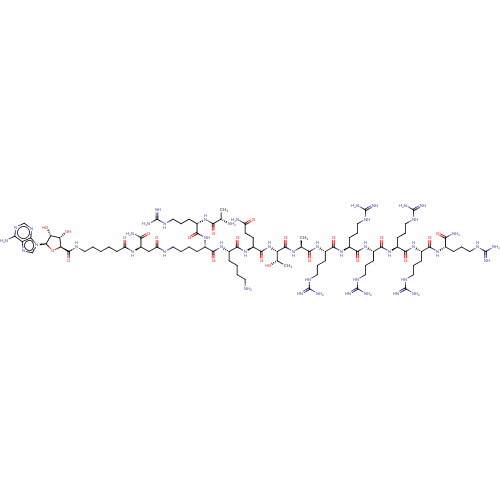
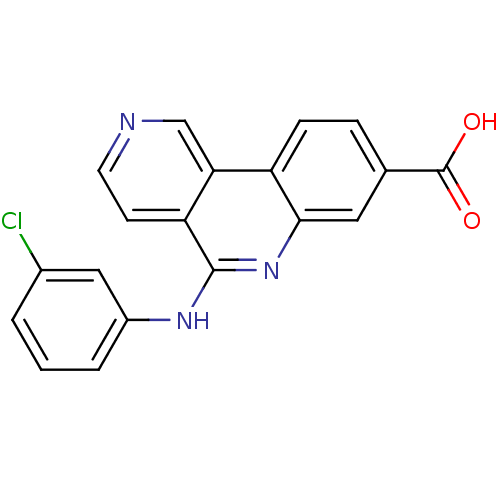
 3D Structure (crystal)
3D Structure (crystal)