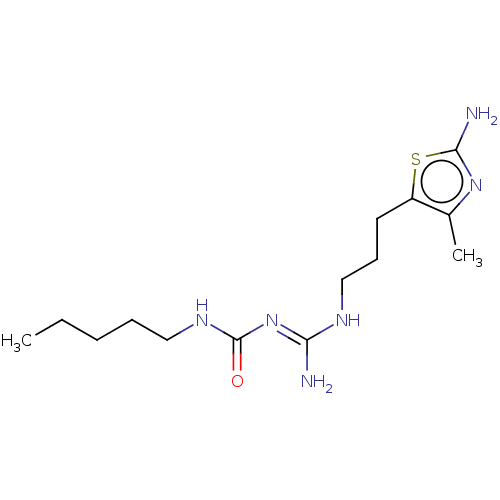
Report error Found 614 Enz. Inhib. hit(s) with all data for entry = 50017400
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 14...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0149nMAssay Description:Displacement of [3H]N-methylspiperone from human D2 long receptor stably expressed in HEK293T cells co-expressing luciferase and CEK at high concentr...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0258nMAssay Description:Displacement of [3H]N-methylspiperone from human D3 receptor stably expressed in HEK293T cells co-expressing luciferase and CEK incubated for 140 min...More data for this Ligand-Target Pair
Affinity DataKd: 0.0520nMAssay Description:Displacement of [3H]N-methylscopolamine from human muscarinic acetylcholine M4 receptor stably expressed in CHO cells by radioligand competition bind...More data for this Ligand-Target Pair
Affinity DataKd: 0.0520nMAssay Description:Displacement of [3H]N-methylscopolamine from human muscarinic acetylcholine M4 receptor stably expressed in CHO cells by radioligand competition bind...More data for this Ligand-Target Pair
Affinity DataKd: 0.0520nMAssay Description:Displacement of [3H]N-methylscopolamine from human muscarinic acetylcholine M4 receptor stably expressed in CHO cells by radioligand competition bind...More data for this Ligand-Target Pair
Affinity DataKd: 0.0520nMAssay Description:Displacement of [3H]N-methylscopolamine from human muscarinic acetylcholine M4 receptor stably expressed in CHO cells by radioligand competition bind...More data for this Ligand-Target Pair
Affinity DataKd: 0.0700nMAssay Description:Displacement of [3H]CGP12177 from human beta-2 adrenergic receptor stably expressed in HEK293T cell membrane incubated for 2 hrs by scintillation cou...More data for this Ligand-Target Pair
Affinity DataKd: 0.0700nMAssay Description:Displacement of [3H]CGP12177 from human beta-2 adrenergic receptor stably expressed in HEK293T cell membrane incubated for 2 hrs by scintillation cou...More data for this Ligand-Target Pair
Affinity DataKd: 0.0700nMAssay Description:Displacement of [3H]CGP12177 from human beta-2 adrenergic receptor stably expressed in HEK293T cell membrane incubated for 2 hrs by scintillation cou...More data for this Ligand-Target Pair
Affinity DataKd: 0.0700nMAssay Description:Displacement of [3H]CGP12177 from human beta-2 adrenergic receptor stably expressed in HEK293T cell membrane incubated for 2 hrs by scintillation cou...More data for this Ligand-Target Pair
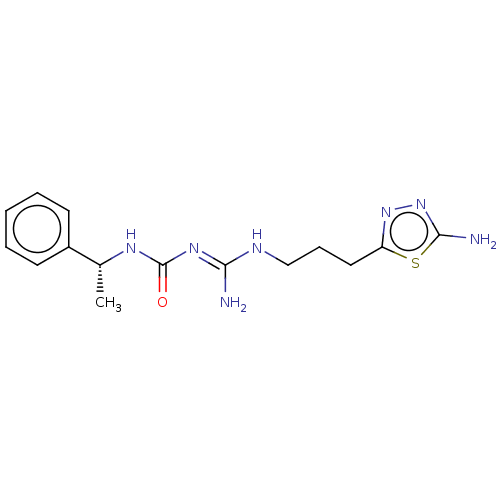
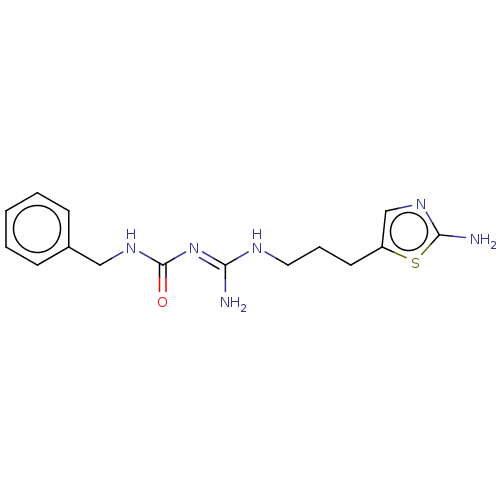

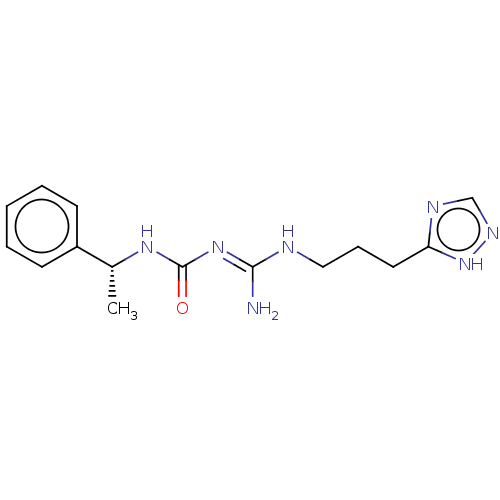




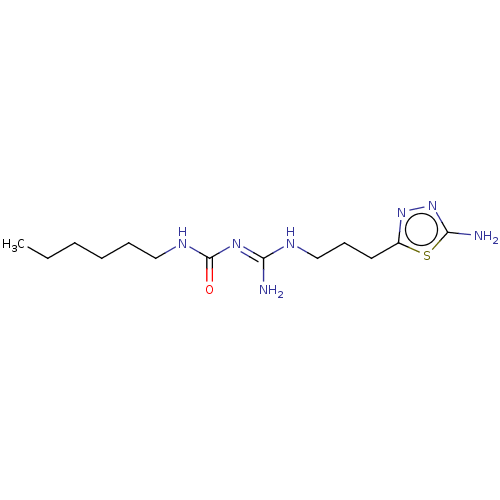
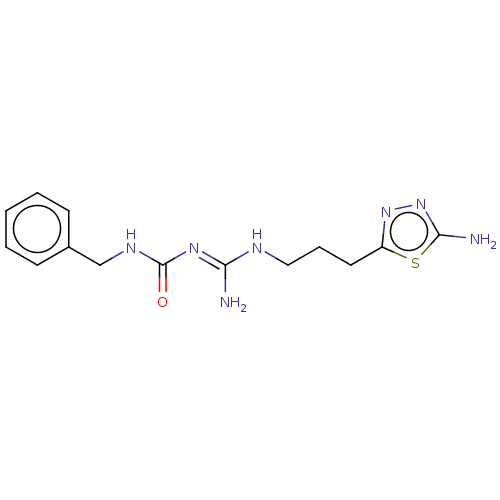
 3D Structure (crystal)
3D Structure (crystal)