Report error Found 21 Enz. Inhib. hit(s) with all data for entry = 50019642
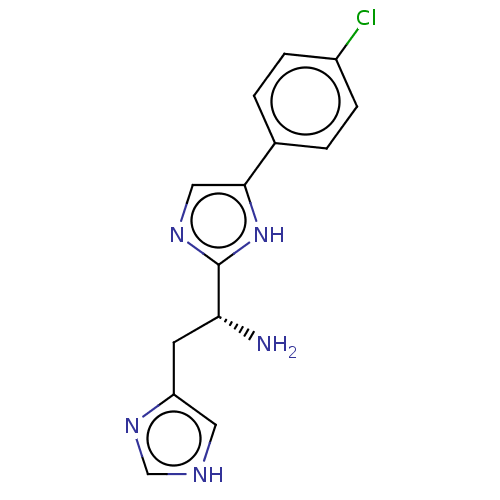
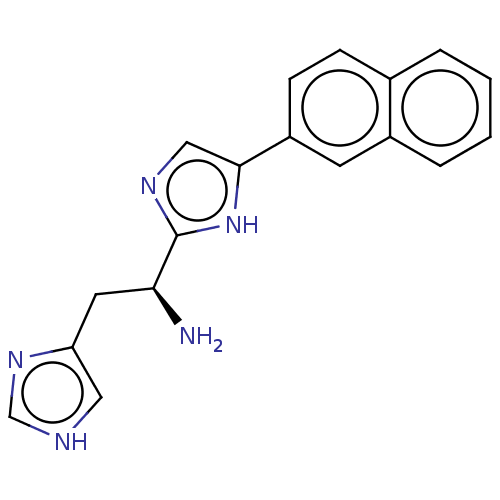
Affinity DataEC50: 2.00E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
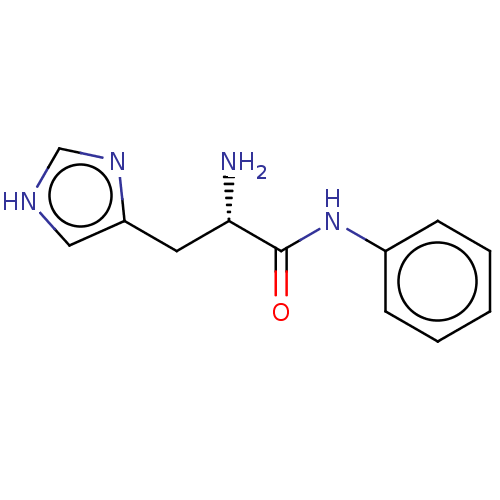
Ligand InfoSimilars
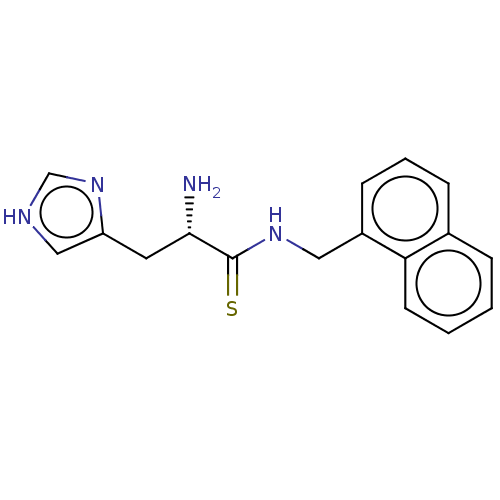
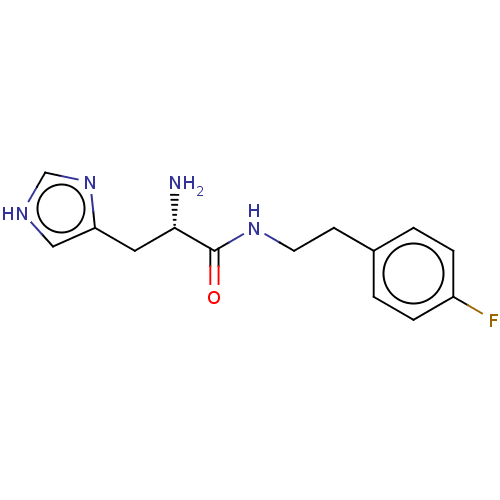
Affinity DataEC50: 2.90E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
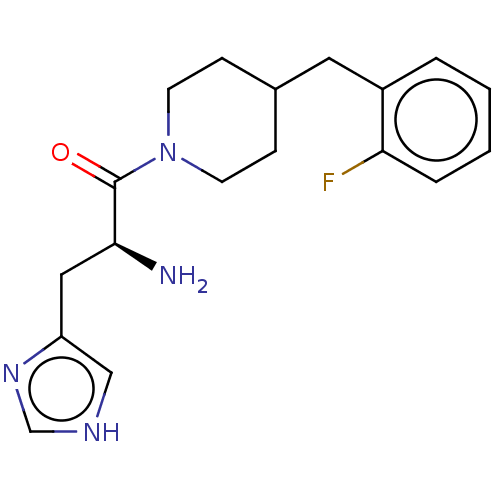
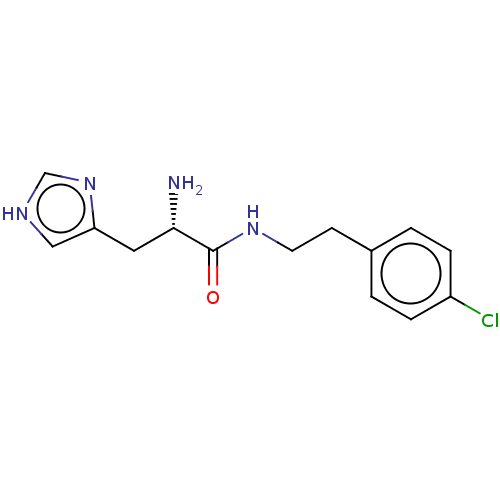
Affinity DataEC50: 3.40E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
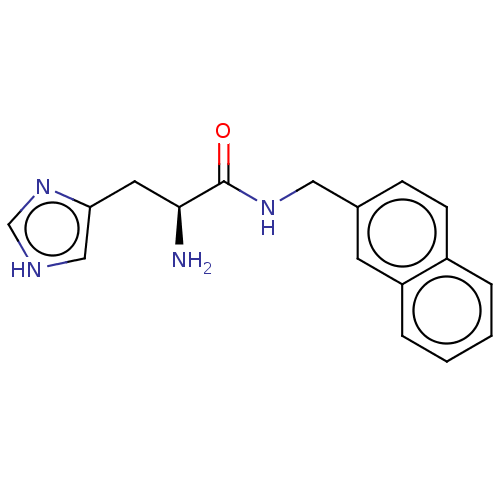
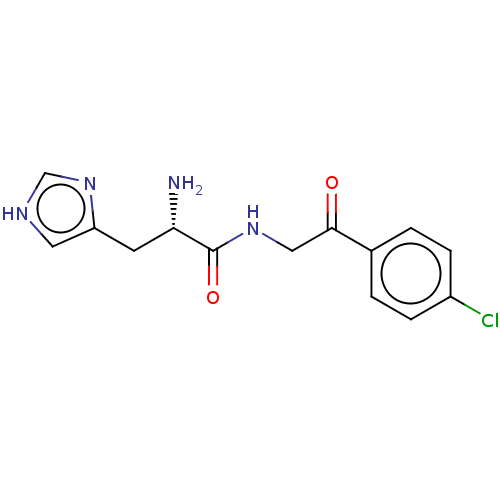
Affinity DataEC50: 3.60E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 3.80E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 4.10E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 4.30E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 4.40E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Affinity DataEC50: 5.20E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 5.40E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 5.40E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 5.50E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 6.30E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 6.80E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 7.00E+3nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 1.16E+4nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Affinity DataEC50: 2.50E+4nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 3.26E+4nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 3.37E+4nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataEC50: 8.55E+4nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair
Affinity DataEC50: 2.52E+5nMAssay Description:Activation of Nln (unknown origin) using Mca-Pro-Leu-Gly-Pro-D-Lys(DNP)- OH as substrate preincubated for 10 mins followed by substrate addition meas...More data for this Ligand-Target Pair