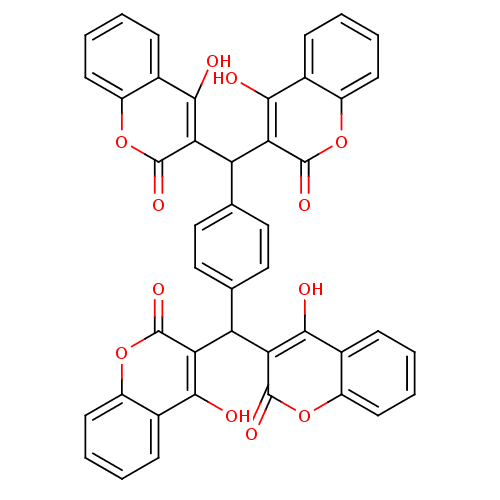
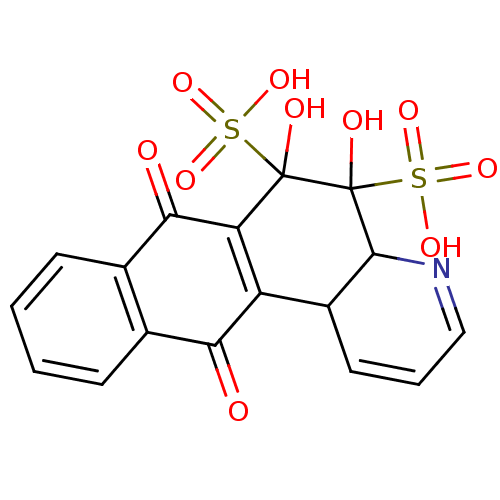
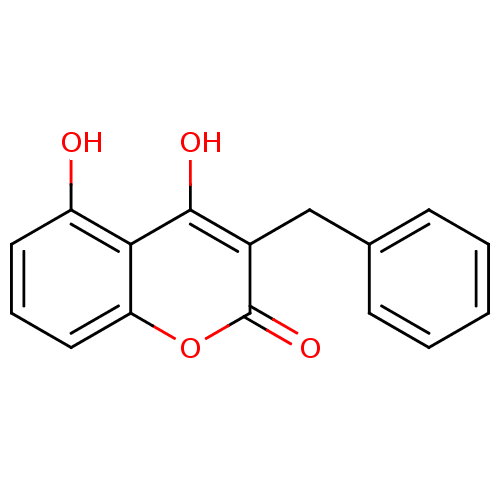
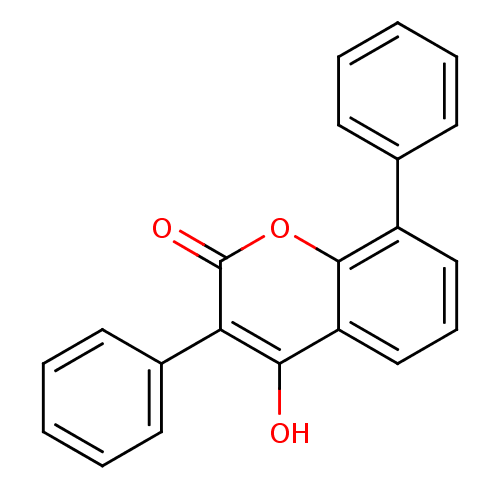
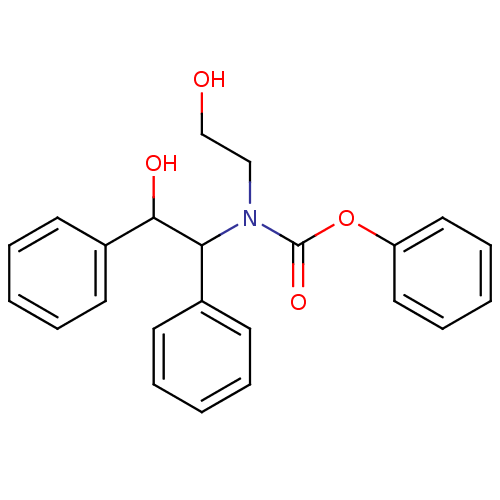
Report error Found 18 Enz. Inhib. hit(s) with all data for entry = 320
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 320nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 750nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 1.70E+3nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 2.00E+3nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 6.00E+3nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 1.30E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 1.60E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 1.70E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 2.10E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 2.30E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 2.40E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 2.60E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 2.70E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 3.00E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 3.10E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 3.20E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 3.50E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
National Institutes of Health
National Institutes of Health
Affinity DataIC50: 4.20E+4nMpH: 6.5 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Protease products were analyzed o...More data for this Ligand-Target Pair