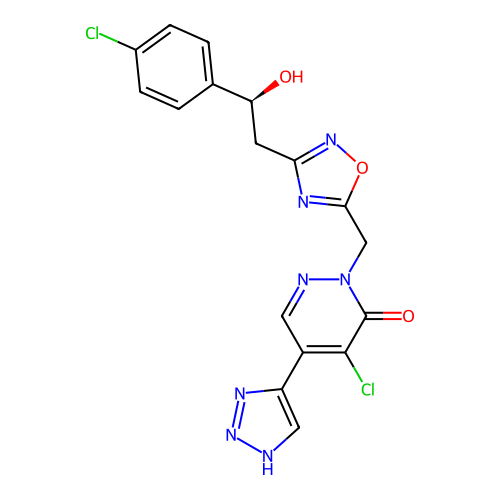
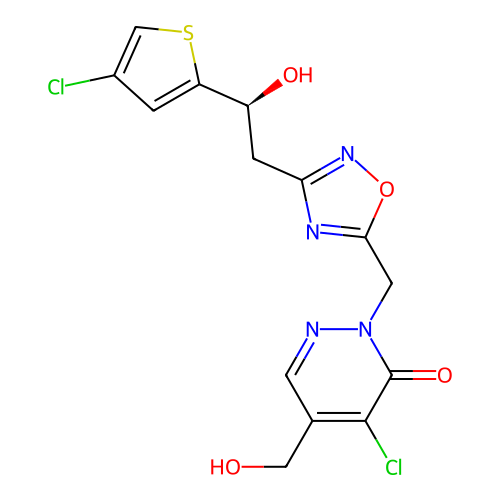
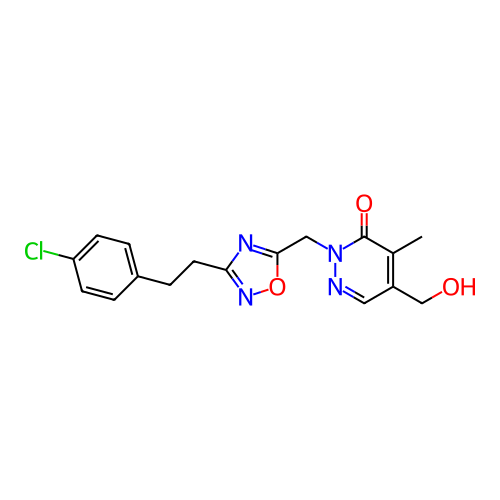
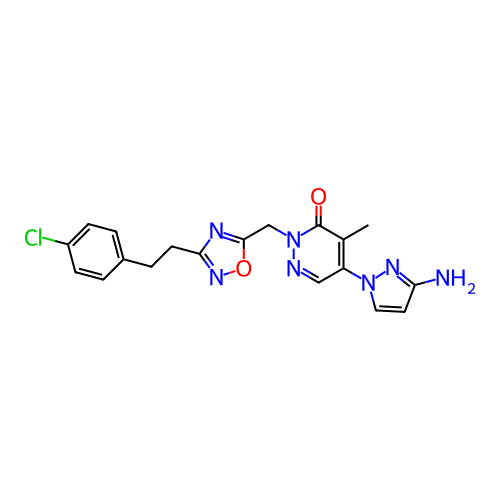
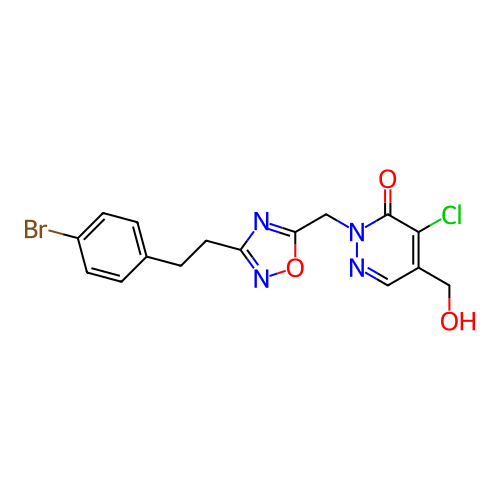
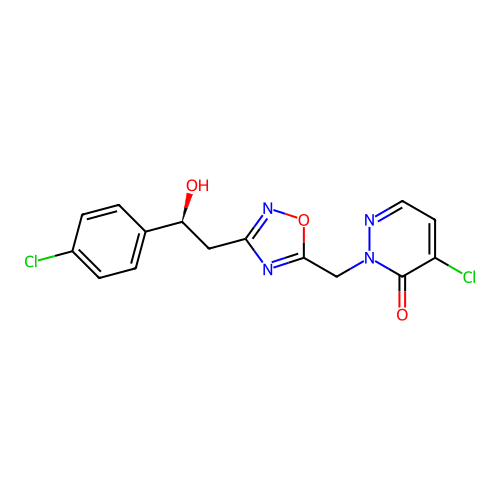
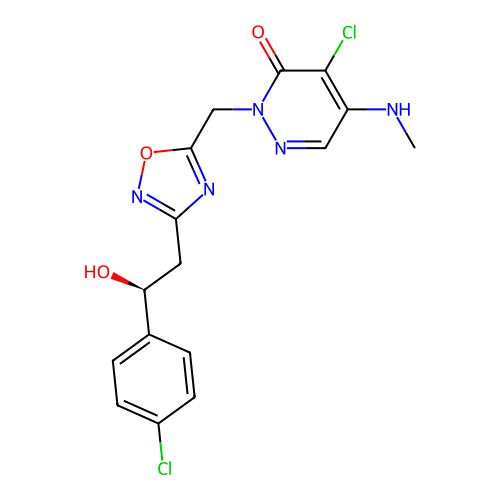
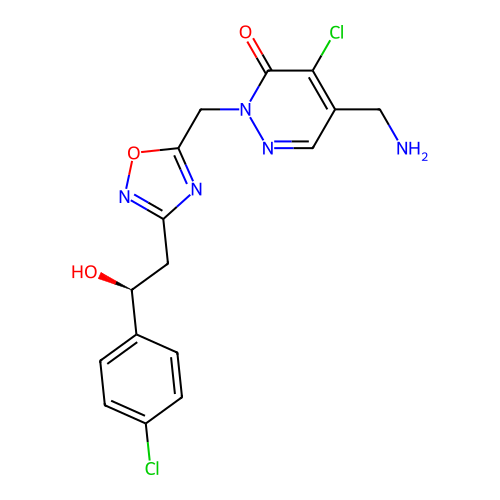
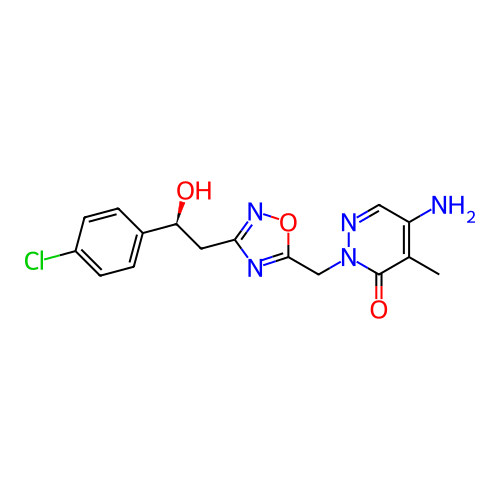
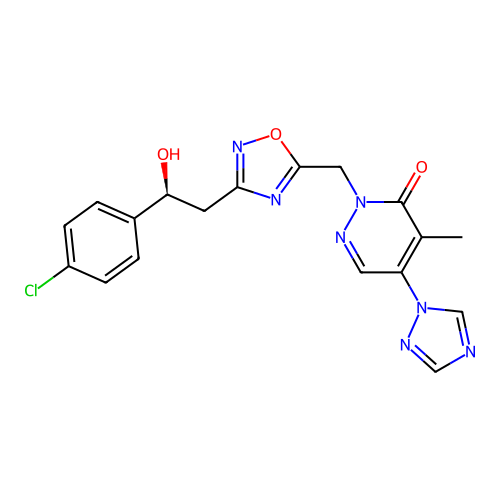
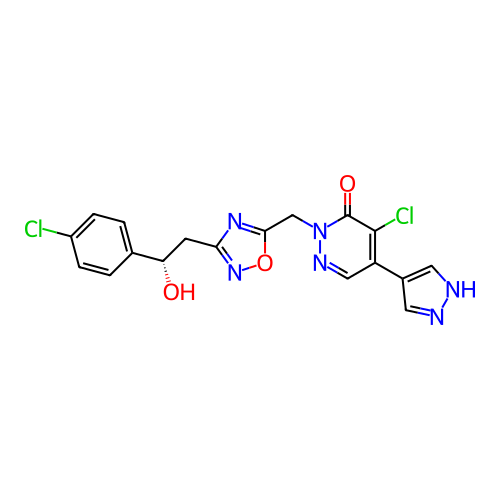
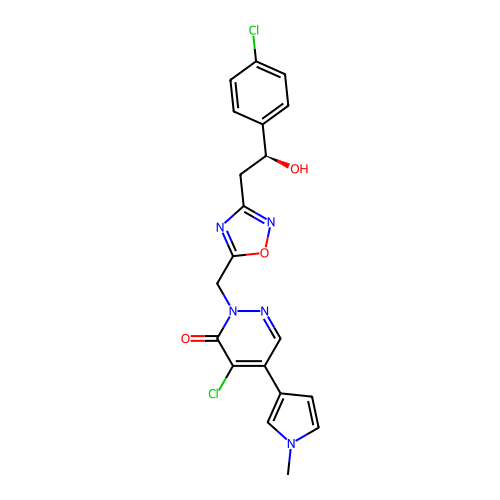
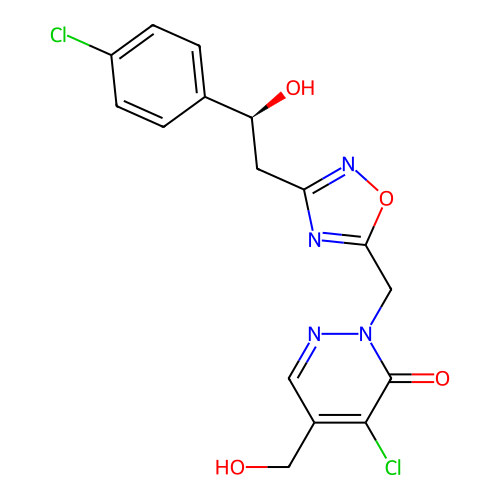
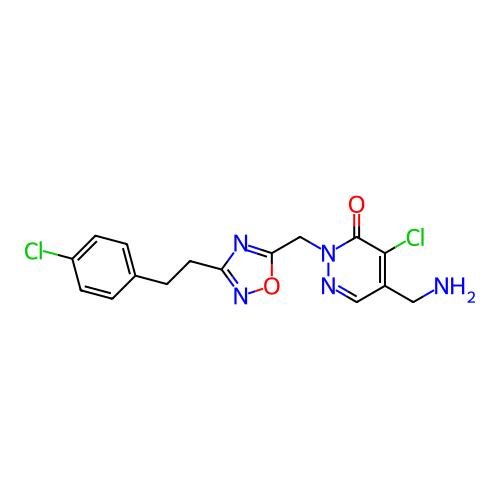
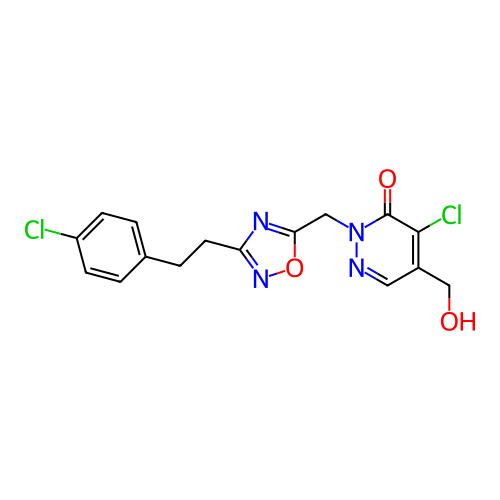
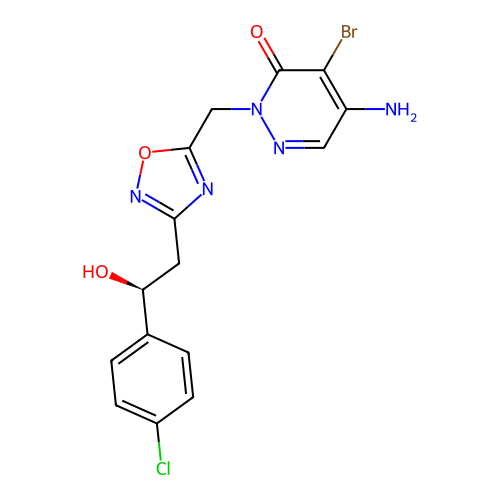
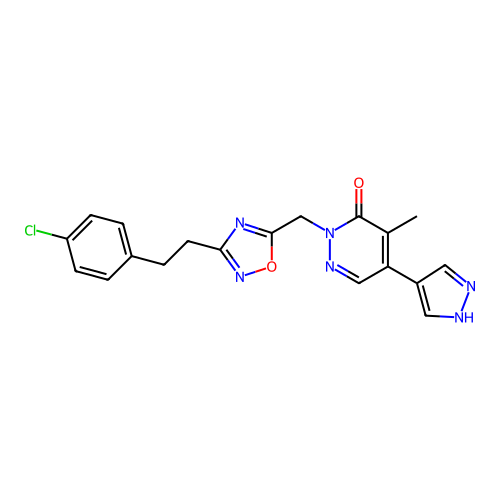
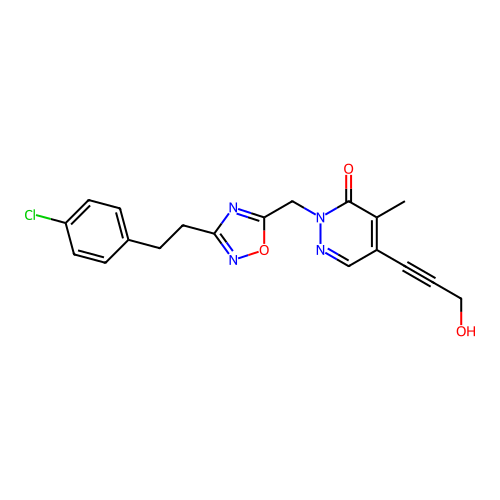
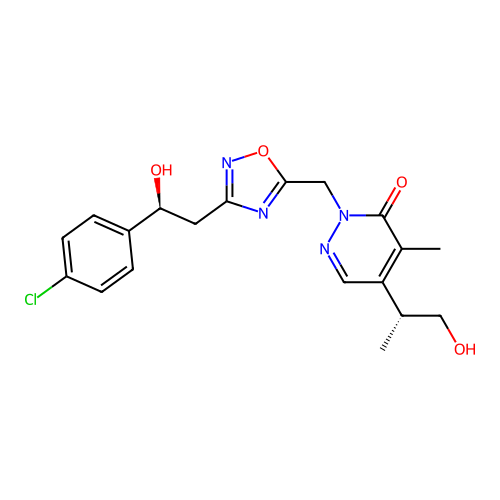
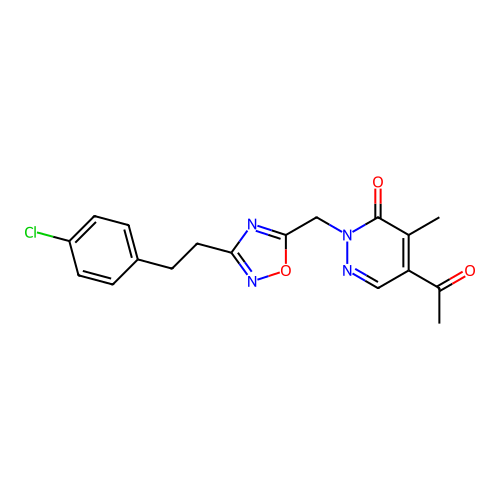
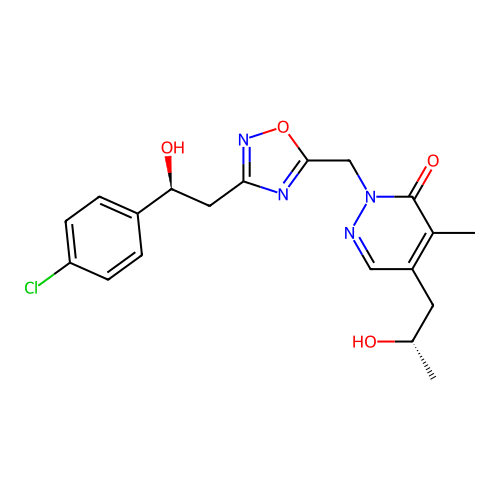
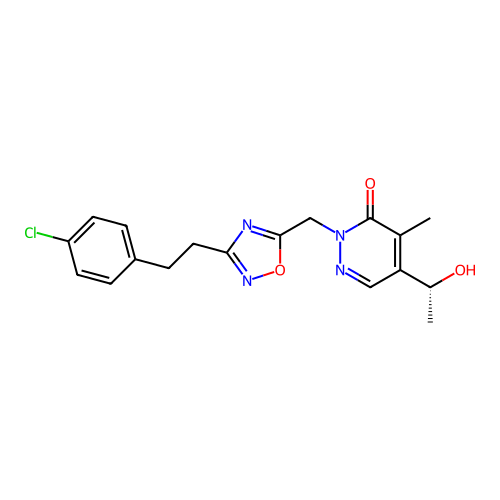
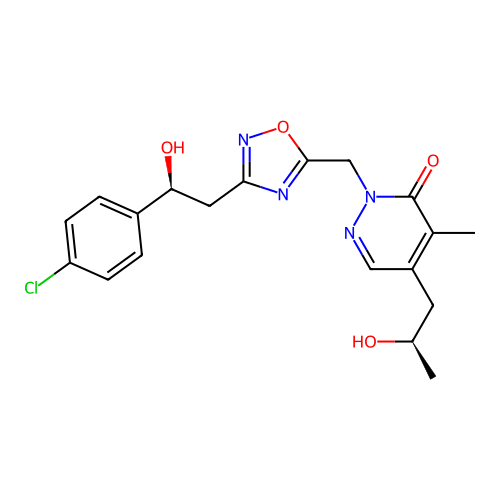
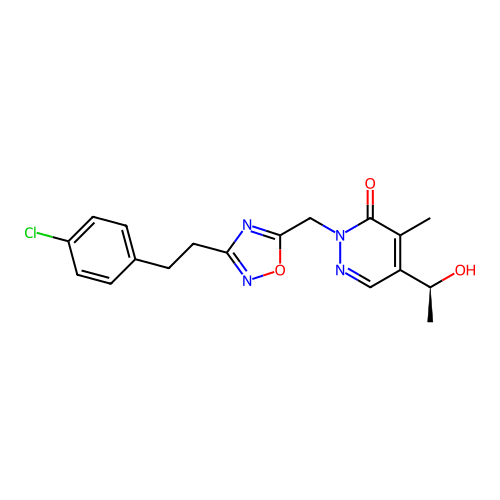
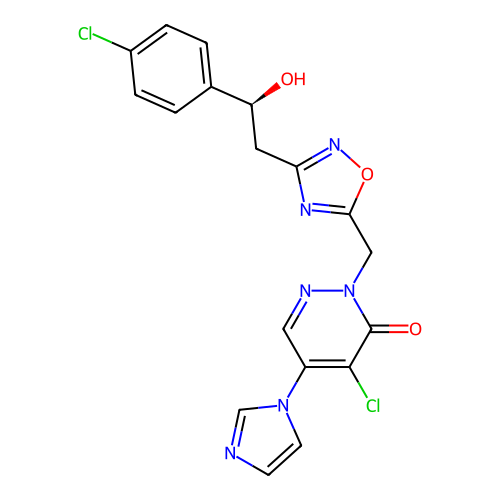
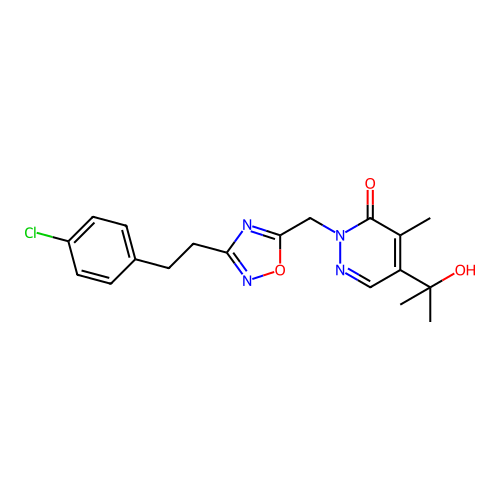
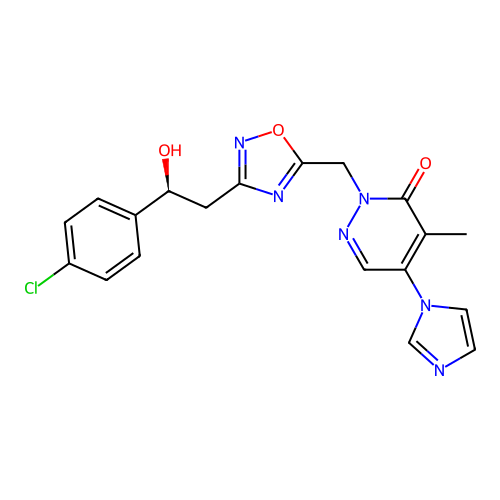
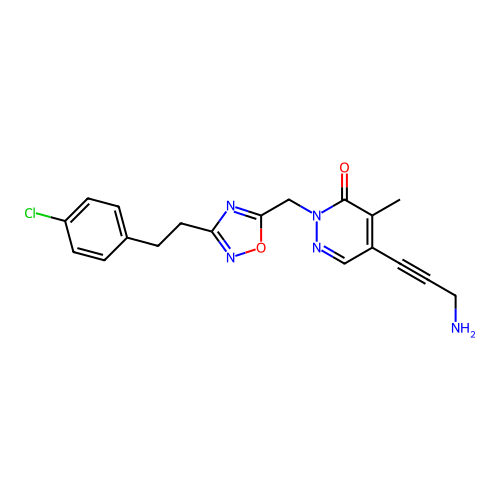
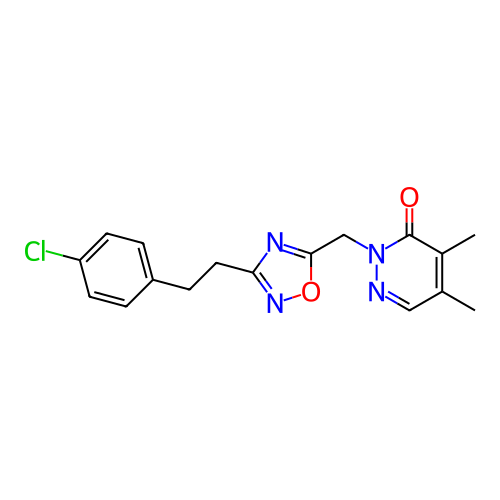
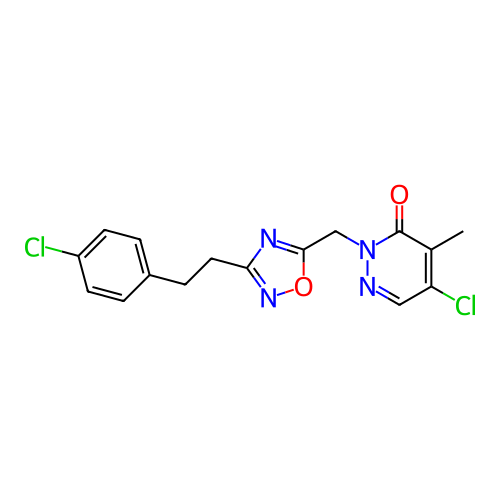
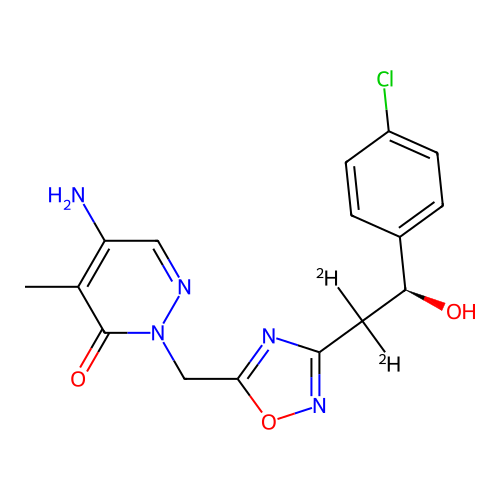
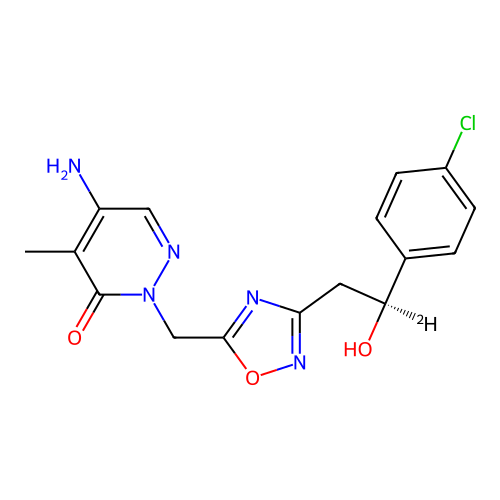
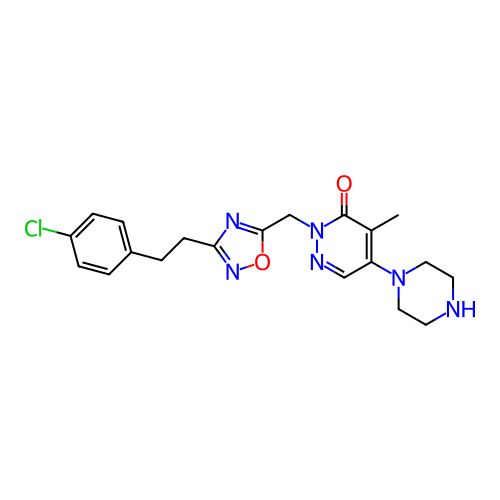
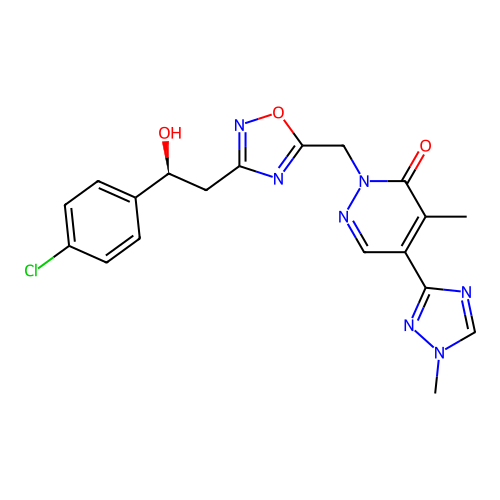
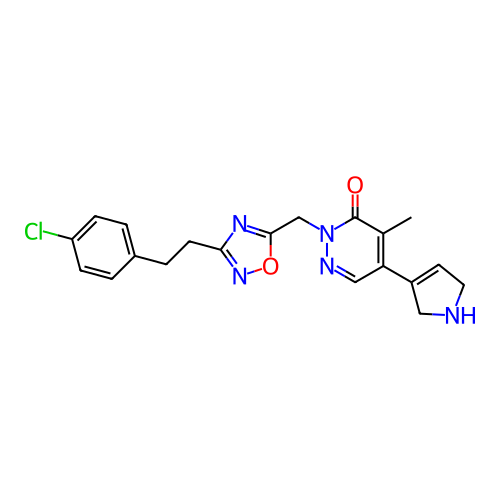
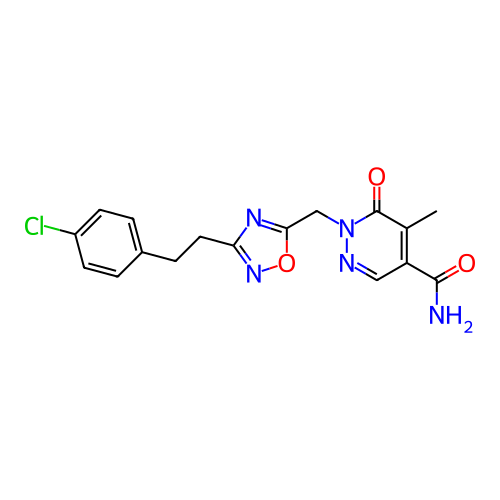
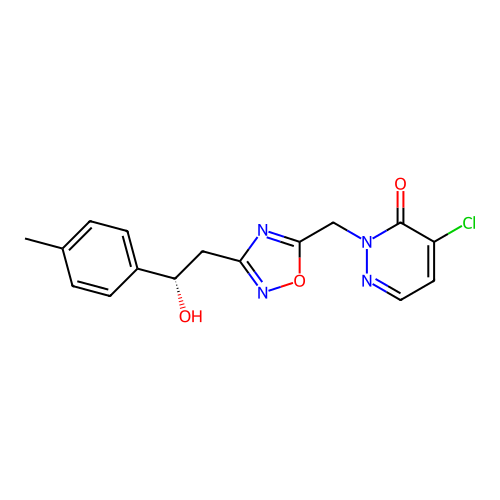
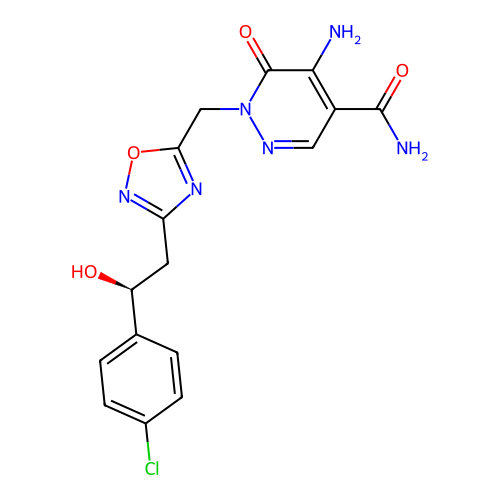
Report error Found 164 Enz. Inhib. hit(s) with all data for entry = 12916
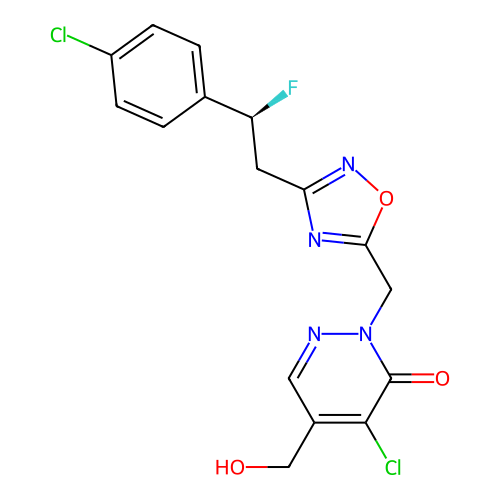
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 100nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily A member 1(Human)
D. E. Shaw Research
US Patent
D. E. Shaw Research
US Patent
Affinity DataIC50: 300nMAssay Description:The cells were bathed in an extracellular solution containing 80 mM NaCl, 60 mM NMDG, 4 mM KCl, 2 mM CaCl2), 6 mM MgCl2, 5 mM Glucose, 10 mM HEPES, 3...More data for this Ligand-Target Pair