Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Glutamate receptor ionotropic, NMDA 1
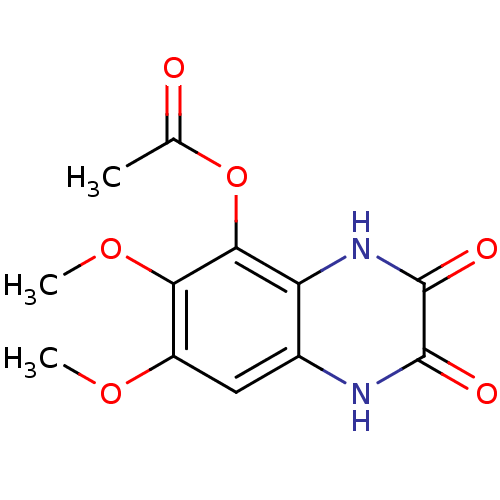
Ligand
BDBM50056603
Substrate
n/a
Meas. Tech.
ChEMBL_140856 (CHEMBL752487)
IC50
53000±n/a nM
Citation
 Cai, SX; Kher, SM; Zhou, ZL; Ilyin, V; Espitia, SA; Tran, M; Hawkinson, JE; Woodward, RM; Weber, E; Keana, JF Structure-activity relationships of alkyl- and alkoxy-substituted 1,4-dihydroquinoxaline-2,3-diones: potent and systemically active antagonists for the glycine site of the NMDA receptor. J Med Chem 40:730-8 (1997) [PubMed] Article
Cai, SX; Kher, SM; Zhou, ZL; Ilyin, V; Espitia, SA; Tran, M; Hawkinson, JE; Woodward, RM; Weber, E; Keana, JF Structure-activity relationships of alkyl- and alkoxy-substituted 1,4-dihydroquinoxaline-2,3-diones: potent and systemically active antagonists for the glycine site of the NMDA receptor. J Med Chem 40:730-8 (1997) [PubMed] Article More Info.:
Target
Name:
Glutamate receptor ionotropic, NMDA 1
Synonyms:
Glutamate (NMDA) receptor subunit zeta 1 | Glutamate [NMDA] receptor subunit zeta-1 | Glutamate-NMDA-Channel | Glutamate-NMDA-MK801 | Glutamate-NMDA-Polyamine | Grin1 | NMDA | NMDZ1_RAT | Nmdar1 | phencyclidine
Type:
Enzyme Catalytic Domain
Mol. Mass.:
105533.40
Organism:
RAT
Description:
P35439
Residue:
938
Sequence:
MSTMHLLTFALLFSCSFARAACDPKIVNIGAVLSTRKHEQMFREAVNQANKRHGSWKIQLNATSVTHKPNAIQMALSVCEDLISSQVYAILVSHPPTPNDHFTPTPVSYTAGFYRIPVLGLTTRMSIYSDKSIHLSFLRTVPPYSHQSSVWFEMMRVYNWNHIILLVSDDHEGRAAQKRLETLLEERESKAEKVLQFDPGTKNVTALLMEARELEARVIILSASEDDAATVYRAAAMLNMTGSGYVWLVGEREISGNALRYAPDGIIGLQLINGKNESAHISDAVGVVAQAVHELLEKENITDPPRGCVGNTNIWKTGPLFKRVLMSSKYADGVTGRVEFNEDGDRKFANYSIMNLQNRKLVQVGIYNGTHVIPNDRKIIWPGGETEKPRGYQMSTRLKIVTIHQEPFVYVKPTMSDGTCKEEFTVNGDPVKKVICTGPNDTSPGSPRHTVPQCCYGFCIDLLIKLARTMNFTYEVHLVADGKFGTQERVNNSNKKEWNGMMGELLSGQADMIVAPLTINNERAQYIEFSKPFKYQGLTILVKKEIPRSTLDSFMQPFQSTLWLLVGLSVHVVAVMLYLLDRFSPFGRFKVNSEEEEEDALTLSSAMWFSWGVLLNSGIGEGAPRSFSARILGMVWAGFAMIIVASYTANLAAFLVLDRPEERITGINDPRLRNPSDKFIYATVKQSSVDIYFRRQVELSTMYRHMEKHNYESAAEAIQAVRDNKLHAFIWDSAVLEFEASQKCDLVTTGELFFRSGFGIGMRKDSPWKQNVSLSILKSHENGFMEDLDKTWVRYQECDSRSNAPATLTFENMAGVFMLVAGGIVAGIFLIFIEIAYKRHKDARRKQMQLAFAAVNVWRKNLQDRKSGRAEPDPKKKATFRAITSTLASSFKRRRSSKDTSTGGGRGALQNQKDTVLPRRAIEREEGQLQLCSRHRES