Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Tyrosine-protein kinase ABL1 [201-500,T315N]
Ligand
BDBM6572
Substrate
BDBM13531
Meas. Tech.
Kinase Assay and Binding Constant Measurement
Kd
300±n/a nM
Citation
 Carter, TA; Wodicka, LM; Shah, NP; Velasco, AM; Fabian, MA; Treiber, DK; Milanov, ZV; Atteridge, CE; Biggs, WH; Edeen, PT; Floyd, M; Ford, JM; Grotzfeld, RM; Herrgard, S; Insko, DE; Mehta, SA; Patel, HK; Pao, W; Sawyers, CL; Varmus, H; Zarrinkar, PP; Lockhart, DJ Inhibition of drug-resistant mutants of ABL, KIT, and EGF receptor kinases. Proc Natl Acad Sci U S A 102:11011-6 (2005) [PubMed] Article
Carter, TA; Wodicka, LM; Shah, NP; Velasco, AM; Fabian, MA; Treiber, DK; Milanov, ZV; Atteridge, CE; Biggs, WH; Edeen, PT; Floyd, M; Ford, JM; Grotzfeld, RM; Herrgard, S; Insko, DE; Mehta, SA; Patel, HK; Pao, W; Sawyers, CL; Varmus, H; Zarrinkar, PP; Lockhart, DJ Inhibition of drug-resistant mutants of ABL, KIT, and EGF receptor kinases. Proc Natl Acad Sci U S A 102:11011-6 (2005) [PubMed] Article More Info.:
Target
Name:
Tyrosine-protein kinase ABL1 [201-500,T315N]
Synonyms:
ABL | ABL Mutant (T315N) | ABL1 | ABL1_HUMAN | JTK7 | Proto-oncogene tyrosine-protein kinase ABL1 mutant T315N | c-ABL | p150
Type:
Enzyme
Mol. Mass.:
34608.76
Organism:
Homo sapiens (Human)
Description:
P00519[201-500,T315N]
Residue:
300
Sequence:
HHSTVADGLITTLHYPAPKRNKPTVYGVSPNYDKWEMERTDITMKHKLGGGQYGEVYEGVWKKYSLTVAVKTLKEDTMEVEEFLKEAAVMKEIKHPNLVQLLGVCTREPPFYIINEFMTYGNLLDYLRECNRQEVNAVVLLYMATQISSAMEYLEKKNFIHRDLAARNCLVGENHLVKVADFGLSRLMTGDTYTAHAGAKFPIKWTAPESLAYNKFSIKSDVWAFGVLLWEIATYGMSPYPGIDLSQVYELLEKDYRMERPEGCPEKVYELMRACWQWNPSDRPSFAEIHQAFETMFQES
Inhibitor
Name:
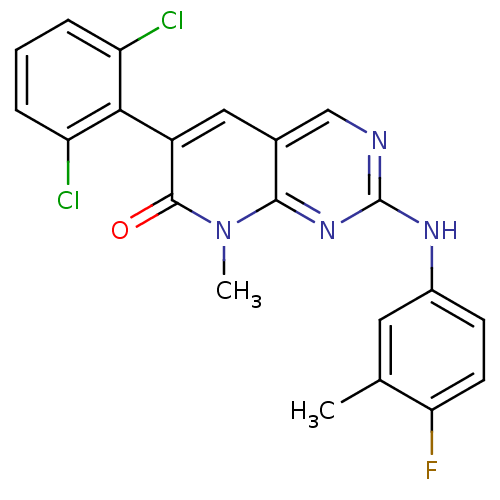
BDBM6572
Synonyms:
6-(2,6-dichlorophenyl)-2-[(4-fluoro-3-methylphenyl)amino]-8-methyl-7H,8H-pyrido[2,3-d]pyrimidin-7-one | 6-(2,6-dichlorophenyl)-2-[(4-fluoro-3-methylphenyl)amino]-8-methylpyrido[2,3-d]pyrimidin-7(8H)-one | PD-180970 | PD180970
Type:
Small organic molecule
Emp. Form.:
C21H15Cl2FN4O
Mol. Mass.:
429.274
SMILES:
Cc1cc(Nc2ncc3cc(-c4c(Cl)cccc4Cl)c(=O)n(C)c3n2)ccc1F |(-2.83,-3.52,;-4.17,-2.76,;-4.17,-1.22,;-5.51,-.46,;-5.51,1.08,;-4.18,1.86,;-4.18,3.4,;-2.84,4.17,;-1.51,3.4,;-.18,4.17,;1.16,3.4,;2.49,4.17,;2.49,5.71,;1.16,6.48,;3.83,6.48,;5.16,5.71,;5.16,4.17,;3.83,3.39,;3.83,1.85,;1.16,1.86,;2.49,1.09,;-.18,1.09,;-.18,-.45,;-1.51,1.86,;-2.84,1.09,;-6.84,-1.23,;-6.83,-2.77,;-5.5,-3.54,;-5.49,-5.08,)|
Substrate
Name:
BDBM13531
Synonyms:
4-(4-Fluorophenyl)-2-(4-hydroxyphenyl)-5-(4-pyridyl)-1H-imidazole | 4-[4-(4-Fluorophenyl)-5-(4-pyridinyl)-1H-imidazol-2-yl]phenol | 4-[4-(4-fluorophenyl)-5-(4-pyridyl)-4-imidazolin-2-ylidene]cyclohexa-2,5-dien-1-one | 4-[5-(4-fluorophenyl)-4-(pyridin-4-yl)-1H-imidazol-2-yl]phenol | SB-202190 | SB202190 | biotinylated SB202190
Type:
Small organic molecule
Emp. Form.:
C20H14FN3O
Mol. Mass.:
331.3431
SMILES:
Oc1ccc(cc1)-c1nc(c([nH]1)-c1ccc(F)cc1)-c1ccncc1